Abstract
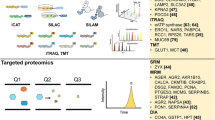
Proteomics has enabled researchers to evaluate global protein changes in a relatively rapid and comprehensive manner. Applications of these technologies in lung research include biomarker and drug discovery, elucidating disease mechanisms, and quantitative clinical assays. Two common workflows exist for quantitative proteomics studies that are aimed at determining differences in protein levels: label-free and labeling methods. Here we describe specific techniques involved in both quantitative workflows; these include extensive sample preparation methods for several lung-specific sample types. Methods are also included for mass spectrometry-based sample analysis and data analysis. While the focus is on quantitative, clinical proteomics, these strategies are appropriate for a wide array of sample types and applications.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Paone G, Leone V, Conti V, De Marchis L, Ialleni E, Graziani C, Salducci M, Ramaccia M, Munafo G (2016) Blood and sputum biomarkers in COPD and asthma: a review. Eur Rev Med Pharmacol Sci 20(4):698–708
Priyadharshini VS, Teran LM (2016) Personalized medicine in respiratory disease: role of proteomics. Adv Protein Chem Struct Biol 102:115–146. https://doi.org/10.1016/bs.apcsb.2015.11.008
Terracciano R, Pelaia G, Preiano M, Savino R (2015) Asthma and COPD proteomics: current approaches and future directions. Proteomics Clin Appl 9(1–2):203–220. https://doi.org/10.1002/prca.201400099
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422(6928):198–207. https://doi.org/10.1038/nature01511
Tao WA, Aebersold R (2003) Advances in quantitative proteomics via stable isotope tagging and mass spectrometry. Curr Opin Biotechnol 14(1):110–118
Spraggins JM, Rizzo DG, Moore JL, Noto MJ, Skaar EP, Caprioli RM (2016) Next-generation technologies for spatial proteomics: integrating ultra-high speed MALDI-TOF and high mass resolution MALDI FTICR imaging mass spectrometry for protein analysis. Proteomics 16(11–12):1678–1689. https://doi.org/10.1002/pmic.201600003
Quon BS, Dai DL, Hollander Z, Ng RT, Tebbutt SJ, Man SF, Wilcox PG, Sin DD (2016) Discovery of novel plasma protein biomarkers to predict imminent cystic fibrosis pulmonary exacerbations using multiple reaction monitoring mass spectrometry. Thorax 71(3):216–222. https://doi.org/10.1136/thoraxjnl-2014-206710
Mrozinski P, Zolotarjova N, Chen H (2008) Human serum and plasma protein depletion—novel high-capacity affinity column for the removal of the “Top 14”abundant proteins. Agilent Technologies Application Note, Santa Clara, CA
Chahrour O, Cobice D, Malone J (2015) Stable isotope labelling methods in mass spectrometry-based quantitative proteomics. J Pharm Biomed Anal 113:2–20. https://doi.org/10.1016/j.jpba.2015.04.013
Chen X, Wei S, Ji Y, Guo X, Yang F (2015) Quantitative proteomics using SILAC: principles, applications, and developments. Proteomics 15(18):3175–3192. https://doi.org/10.1002/pmic.201500108
Ong SE, Mann M (2006) A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat Protoc 1(6):2650–2660. https://doi.org/10.1038/nprot.2006.427
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Reisdorph, N.A., Michel, C., Fritz, K., Reisdorph, R. (2018). Application of Proteomics in Lung Research. In: Alper, S., Janssen, W. (eds) Lung Innate Immunity and Inflammation. Methods in Molecular Biology, vol 1809. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8570-8_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8570-8_16
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8569-2
Online ISBN: 978-1-4939-8570-8
eBook Packages: Springer Protocols