Abstract
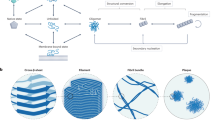
The formation of amyloid fibrils is a central phenomenon in the progressive pathology of many neurodegenerative diseases, as well as in the fabrication of functional materials. Several different molecular processes acting in concert are responsible for the formation of amyloid fibrils from monomeric protein in solution. Here, we describe a method to determine which microscopic processes drive the overall formation of fibrils by using chemical kinetics in combination with systematic experimental datasets analysed in a global manner. We outline general concepts for obtaining suitable kinetic data and detail the key stages of data analysis, from quality control to the verification of a specific mechanism of aggregation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Knowles TPJ, Vendruscolo M, Dobson CM (2015) The physical basis of protein misfolding disorders. Phys Today 68:36
Chiti F, Dobson CM (2006) Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 75:333–366
Dobson CM (2003) Protein folding and misfolding. Nature 426:884–890
Aguzzi A, O'Connor T (2010) Protein aggregation diseases: pathogenicity and therapeutic perspectives. Nat Rev Drug Discov 9:237–248
Hardy J, Selkoe DJ (2002) The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 297:353–356
Maji SK, Perrin MH, Sawaya MR et al (2009) Functional amyloids as natural storage of peptide hormones in pituitary secretory granules. Science 325(5938):328–332
Shewmaker F, McGlinchey RP, Wickner RB (2011) Structural insights into functional and pathological amyloid. J Biol Chem 286(19):16533–16540
Ferrone FA, Hofrichter J, Eaton WA (1985) Kinetics of sickle hemoglobin polymerization. II. A double nucleation mechanism. J Mol Biol 183:611–631
Oosawa F, Asakura S (1975) Thermodynamics of the polymerization of protein. Academic, New York
Knowles TPJ, Waudby CA, Devlin GL et al (2009) An analytical solution to the kinetics of breakable filament assembly. Science 326:1533–1537
Thomas CTM, Andela Š, Johnny H, et al (2018) Chemical kinetics for kridging molecular mechanisms and macroscopic measurements of amyloid fibril formation. Annu Rev Phy Chem 69:1
Meisl G, Yang X, Hellstrand E et al (2014) Differences in nucleation behavior underlie the contrasting aggregation kinetics of the Aβ40 and Aβ42 peptides. Proc Natl Acad Sci U S A 111:9384–9389
Cohen SIA, Linse S, Luheshi LM et al (2013) Proliferation of amyloid-beta42 aggregates occurs through a secondary nucleation mechanism. Proc Natl Acad Sci U S A 110:9758–9763
Walsh DM, Thulin E, Minogue AM et al (2009) A facile method for expression and purification of the Alzheimer's disease-associated amyloid beta-peptide. FEBS J 276:1266–1281
Meisl G, Kirkegaard JB, Arosio P et al (2016) Molecular mechanisms of protein aggregation from global fitting of kinetic models. Nat Protoc 11(2):252–272
Cohen SIA, Vendruscolo M, Dobson CM et al (2011) Nucleated polymerization with secondary pathways. i. time evolution of the principal moments. J Chem Phys 135:065105
Cohen SIA, Vendruscolo M, Dobson CM et al (2011) Nucleated polymerization with secondary pathways. ii. determination of self-consistent solutions to growth processes described by non-linear master equations. J Chem Phys 135:065106
Cohen SIA, Vendruscolo M, Dobson CM et al (2012) From macroscopic measurements to microscopic mechanisms of protein aggregation. J Mol Biol 421:160–171
Gaspar R, Meisl G, Buell AK et al (2017) Secondary nucleation of monomers on fibril surface dominates α-synuclein aggregation and provides autocatalytic amyloid amplification. Q Rev Biophys 50:e6
Michaels TCT, Yde P, Willis JC et al (2015) The length distribution of frangible biofilaments. J Chem Phys 143:164901
Arosio P, Vendruscolo M, Dobson CM et al (2014) Chemical kinetics for drug discovery to combat protein aggregation diseases. Trends Pharmacol Sci 35:127–135
Abelein A, Graslund A, Danielsson J (2015) Zinc as chaperone-mimicking agent for retardation of amyloid β peptide fibril formation. Proc Natl Acad Sci U S A 112:5407–5412
Cohen SIA, Arosio P, Presto J et al (2015) The molecular chaperone brichos breaks the catalytic cycle that generates toxic Ab oligomers. Nat Struct Mol Biol 22:207–213
Cukalevski R, Yang X, Meisl G et al (2015) The A-beta 40 and A-beta 42 peptides self-assemble into separate homomolecular fibrils in binary mixtures but cross-react during primary nucleation. Chem Sci 6:4215
Michaels TCT, Dear A, Kirkegaard JB et al (2016) Fluctuations in the kinetics of linear protein self-assembly. Phys Rev Lett 116:258103
Finder VH, Vodopivec I, Nitsch RM et al (2010) The recombinant amyloid-beta peptide Abeta1-42 aggregates faster and is more neurotoxic than synthetic Abeta1-42. J Mol Biol 396:9–18
Wales DJ, Doye JPK (1997) Global optimization by basin-hopping and the lowest energy structures of lennard-jones clusters containing up to 110 atoms. J Phys Chem A 101:5111–5116
Michaels TCT, Cohen SIA, Vendruscolo M et al (2016) Hamiltonian dynamics of protein filament formation. Phys Rev Lett 116:038101
Meisl G, Yang X, Frohm B et al (2016) Quantitative analysis of intrinsic and extrinsic factors in the aggregation mechanism of Alzheimer-associated Aβ-peptide. Sci Rep 6:18728
Meisl G, Rajah L, Cohen SIA, et al (2017) Scaling behaviour and rate-determining steps in filamentous self-assembly. Chem Sci 8:7087–7097
Acknowledgments
This work was supported by Sidney Sussex College Cambridge (G.M.), the Swiss National Science Foundation (T.C.T.M.), Peterhouse College, Cambridge (T.C.T.M.) the ERC (T.P.J.K., S.L.), Swedish research council (S.L.), the BBSRC (T.P.J.K.), and the Newman Foundation (T.P.J.K.). We thank the members of the Knowles and Linse research groups for their input on experiences with this method, in particular Alexander Dear, Erik Hellstrand, Risto Cukalevski, Xiaoting Yang, Kalyani Sanagavarapu, Tanja Weiffert, Celine Galvagnion, Ricardo Gaspar, Ryan Limbocker, Catherine Xu and Patrick Flagmeier.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Meisl, G., Michaels, T.C.T., Linse, S., Knowles, T.P.J. (2018). Kinetic Analysis of Amyloid Formation. In: Sigurdsson, E., Calero, M., Gasset, M. (eds) Amyloid Proteins. Methods in Molecular Biology, vol 1779. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7816-8_12
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7816-8_12
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7815-1
Online ISBN: 978-1-4939-7816-8
eBook Packages: Springer Protocols