Abstract
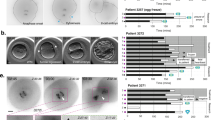
Chromothripsis is a phenomenon observed in cancer cells, wherein a single or few chromosome(s) exhibit vast genomic rearrangements. Recent studies elucidated a striking series of events in which defective segregation of chromosomes causes their incorporation into micronuclei, where they are subject to extensive DNA damage prior to re-joining the main mass of chromosomes in a subsequent cell cycle, which provide an appealing mechanism for the etiology of chromothripsis. Micronuclei are well known to be common in human preimplantation embryos. We recently showed that, unlike in cancer cells, in mouse preimplantation embryos the micronuclei are maintained during multiple cell generations and apparently fail to re-join the main set of chromosomes. This unexpected finding could safeguard the early embryonic genome from chromothripsis. Here, we describe an approach that combines live and immunofluorescence imaging methods that was pivotal in that study to reveal the lack of a functional kinetochore in chromosomes from mouse embryo micronuclei.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kloosterman WP, Hoogstraat M, Paling O et al (2011) Chromothripsis is a common mechanism driving genomic rearrangements in primary and metastatic colorectal cancer. Genome Biol 12:R103. https://doi.org/10.1186/gb-2011-12-10-r103
Stephens PJ, Greenman CD, Fu B et al (2011) Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144:27–40
Adey A, Burton JN, Kitzman JO et al (2013) The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line. Nature 500:207–211
Forment JV, Kaidi A, Jackson SP (2012) Chromothripsis and cancer: causes and consequences of chromosome shattering. Nat Rev Cancer 12:663–670
Storchova Z, Kloosterman WP (2016) The genomic characteristics and cellular origin of chromothripsis. Curr Opin Cell Biol 40:106–113
Thompson SL, Compton DA (2011) Chromosome missegregation in human cells arises through specific types of kinetochore–microtubule attachment errors. Proc Natl Acad Sci 108:17974–17978
Cimini D, Howell B, Maddox P et al (2001) Merotelic kinetochore orientation is a major mechanism of aneuploidy in mitotic mammalian tissue cells. J Cell Biol 153:517–528
Crasta K, Ganem NJ, Dagher R et al (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482:53–58
Hatch EM, Fischer AH, Deerinck T et al (2013) Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154:47–60
Ly P, Teitz LS, Kim DH et al (2017) Selective Y centromere inactivation triggers chromosome shattering in micronuclei and repair by non-homologous end joining. Nat Cell Biol 19:68–75
Zhang CZ, Spektor A, Cornils H et al (2015) Chromothripsis from DNA damage in micronuclei. Nature 522:179–184
Fragouli E, Alfarawati S, Spath K et al (2013) The origin and impact of embryonic aneuploidy. Hum Genet 132:1001–1013
Kort DH, Chia G, Treff NR et al (2016) Human embryos commonly form abnormal nuclei during development: a mechanism of DNA damage, embryonic aneuploidy, and developmental arrest. Hum Reprod 31:312–323
Vázquez-Diez C, Yamagata K, Trivedi S et al (2016) Micronucleus formation causes perpetual unilateral chromosome inheritance in mouse embryos. Proc Natl Acad Sci U S A 113:626–631
Rieder CL, Khodjakov A (2003) Mitosis through the microscope: advances in seeing inside live dividing cells. Science 300:91–96
Hinchcliffe EH, Day CA, Karanjeet KB et al (2016) Chromosome missegregation during anaphase triggers p53 cell cycle arrest through histone H3.3 Ser31 phosphorylation. Nat Cell Biol 18:668–675
Hornick JE, Mader CC, Tribble EK et al (2011) Amphiastral mitotic spindle assembly in vertebrate cells lacking centrosomes. Curr Biol 21:598–605
Gregan J, Polakova S, Zhang L et al (2011) Merotelic kinetochore attachment: causes and effects. Trends Cell Biol 21:374–381
Nakagawa S, FitzHarris G (2016) Quantitative microinjection of morpholino antisense oligonucleotides into mouse oocytes to examine gene function in meiosis-I. Methods Mol Biol 1457:217–230
Nagy A (2003) Manipulating the mouse embryo: a laboratory manual, 3rd edn. Cold Spring Harbor, New York
Erbach GT, Lawitts JA, Papaioannou VE et al (1994) Differential growth of the mouse preimplantation embryo in chemically defined media. Biol Reprod 50:1027–1033
Gardner DK, Leese HJ (1986) Non-invasive measurement of nutrient uptake by single cultured pre-implantation mouse embryos. Hum Reprod 1:25–27
Acknowledgments
Work in GF’s lab is supported by Fondation JL Lévesque, NSERC, and CIHR.
We thank Gaudeline Rémillard-Labrosse and Angus Macaulay for critical reading and discussion of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Vázquez-Diez, C., FitzHarris, G. (2018). Correlative Live Imaging and Immunofluorescence for Analysis of Chromosome Segregation in Mouse Preimplantation Embryos. In: Pellestor, F. (eds) Chromothripsis. Methods in Molecular Biology, vol 1769. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7780-2_20
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7780-2_20
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7779-6
Online ISBN: 978-1-4939-7780-2
eBook Packages: Springer Protocols