Abstract
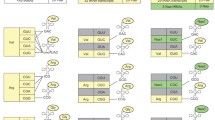
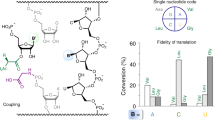
In ribosomal polypeptide synthesis, the 61 sense codons redundantly code for the 20 proteinogenic amino acids. The genetic code contains eight family codon boxes consisting of synonymous codons that redundantly code for the same amino acid. Here, we describe the protocol of a recently published method to artificially divide such family codon boxes and encode multiple nonproteinogenic amino acids in addition to the 20 proteinogenic ones in a reprogrammed genetic code. To achieve this, an in vitro translation system reconstituted with 32 in vitro transcribed tRNASNN’s (S = C or G; N = U, C, A or G) was first developed, where the 32 tRNA transcripts can be charged with 20 proteinogenic amino acids by aminoacyl-tRNA synthetases in situ and orthogonally decode the corresponding 31 NNS sense codons as well as the AUG initiation codon. When some redundant tRNAGNN’s are replaced with tRNAGNN’s precharged with nonproteinogenic amino acids by means of flexizymes, the nonproteinogenic and proteinogenic aminoacyl-tRNAs can decode the NNC and NNG codons in the same family codon box independently. In this protocol, we describe expression of model peptides, including a macrocyclic peptide containing three kinds of N-methyl-amino acids reassigned to the vacant codons generated by the method of artificial division of codon boxes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Murakami H, Ohta A, Ashigai H, Suga H (2006) A highly flexible tRNA acylation method for non-natural polypeptide synthesis. Nat Methods 3(5):357–359
Goto Y, Ohta A, Sako Y, Yamagishi Y, Murakami H, Suga H (2008) Reprogramming the translation initiation for the synthesis of physiologically stable cyclic peptides. ACS Chem Biol 3(2):120–129
Goto Y, Murakami H, Suga H (2008) Initiating translation with D-amino acids. RNA 14(7):1390–1398
Kawakami T, Murakami H, Suga H (2008) Messenger RNA-programmed incorporation of multiple N-methyl-amino acids into linear and cyclic peptides. Chem Biol 15(1):32–42
Xiao H, Murakami H, Suga H, Ferre-D'Amare AR (2008) Structural basis of specific tRNA aminoacylation by a small in vitro selected ribozyme. Nature 454(7202):358–361
Ohta A, Murakami H, Higashimura E, Suga H (2007) Synthesis of polyester by means of genetic code reprogramming. Chem Biol 14(12):1315–1322
Goto Y, Katoh T, Suga H (2011) Flexizymes for genetic code reprogramming. Nat Protoc 6(6):779–790
Forster AC, Tan Z, Nalam MN, Lin H, Qu H, Cornish VW, Blacklow SC (2003) Programming peptidomimetic syntheses by translating genetic codes designed de novo. Proc Natl Acad Sci U S A 100(11):6353–6357
Josephson K, Hartman MC, Szostak JW (2005) Ribosomal synthesis of unnatural peptides. J Am Chem Soc 127(33):11727–11735
Yamagishi Y, Shoji I, Miyagawa S, Kawakami T, Katoh T, Goto Y, Suga H (2011) Natural product-like macrocyclic N-methyl-peptide inhibitors against a ubiquitin ligase uncovered from a ribosome-expressed de novo library. Chem Biol 18(12):1562–1570
Nemoto N, Miyamoto-Sato E, Husimi Y, Yanagawa H (1997) In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett 414(2):405–408
Roberts RW, Szostak JW (1997) RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc Natl Acad Sci U S A 94(23):12297–12302
Terasaka N, Suga H (2014) Flexizymes-facilitated genetic code reprogramming leading to the discovery of drug-like peptides. Chem Lett 43(1):11–19
Tanaka Y, Hipolito CJ, Maturana AD, Ito K, Kuroda T, Higuchi T, Katoh T, Kato HE, Hattori M, Kumazaki K, Tsukazaki T, Ishitani R, Suga H, Nureki O (2013) Structural basis for the drug extrusion mechanism by a MATE multidrug transporter. Nature 496(7444):247–251
Hayashi Y, Morimoto J, Suga H (2012) In vitro selection of anti-Akt2 thioether-macrocyclic peptides leading to isoform-selective inhibitors. ACS Chem Biol 7(3):607–613
Morimoto J, Hayashi Y, Suga H (2012) Discovery of macrocyclic peptides armed with a mechanism-based warhead: isoform-selective inhibition of human deacetylase SIRT2. Angew Chem Int Ed Engl 51(14):3423–3427
Yamagata K, Goto Y, Nishimasu H, Morimoto J, Ishitani R, Dohmae N, Takeda N, Nagai R, Komuro I, Suga H, Nureki O (2014) Structural basis for potent inhibition of SIRT2 deacetylase by a macrocyclic peptide inducing dynamic structural change. Structure 22(2):345–352
Shimizu Y, Inoue A, Tomari Y, Suzuki T, Yokogawa T, Nishikawa K, Ueda T (2001) Cell-free translation reconstituted with purified components. Nat Biotechnol 19(8):751–755
Komine Y, Adachi T, Inokuchi H, Ozeki H (1990) Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J Mol Biol 212(4):579–598
FVt M, Ramakrishnan V (2004) Structure of a purine-purine wobble base pair in the decoding center of the ribosome. Nat Struct Mol Biol 11(12):1251–1252
Gustilo EM, Vendeix FA, Agris PF (2008) tRNA's modifications bring order to gene expression. Curr Opin Microbiol 11(2):134–140. https://doi.org/10.1016/j.mib.2008.02.003
Crick FH (1966) Codon—anticodon pairing: the wobble hypothesis. J Mol Biol 19(2):548–555
Iwane Y, Hitomi A, Murakami H, Katoh T, Goto Y, Suga H (2016) Expanding the amino acid repertoire of ribosomal polypeptide synthesis via the artificial division of codon boxes. Nat Chem 8(4):317–325
Milligan JF, Groebe DR, Witherell GW, Uhlenbeck OC (1987) Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res 15(21):8783–8798
Schagger H, von Jagow G (1987) Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem 166(2):368–379
Schagger H (2006) Tricine-SDS-PAGE. Nat Protoc 1(1):16–22
Christian EL, McPheeters DS, Harris ME (1998) Identification of individual nucleotides in the bacterial ribonuclease P ribozyme adjacent to the pre-tRNA cleavage site by short-range photo-cross-linking. Biochemistry 37(50):17618–17628
Guo X, Campbell FE, Sun L, Christian EL, Anderson VE, Harris ME (2006) RNA-dependent folding and stabilization of C5 protein during assembly of the E. coli RNase P holoenzyme. J Mol Biol 360(1):190–203
Rappsilber J, Ishihama Y, Mann M (2003) Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal Chem 75(3):663–670
Tamura K, Himeno H, Asahara H, Hasegawa T, Shimizu M (1992) In vitro study of E.coli tRNAArg and tRNALys identity elements. Nucleic Acids Res 20(9):2335–2339
Sylvers LA, Rogers KC, Shimizu M, Ohtsuka E, Soll D (1993) A 2-thiouridine derivative in tRNAGlu is a positive determinant for aminoacylation by Escherichia coli glutamyl-tRNA synthetase. Biochemistry 32(15):3836–3841
Nureki O, Niimi T, Muramatsu T, Kanno H, Kohno T, Florentz C, Giege R, Yokoyama S (1994) Molecular recognition of the identity-determinant set of isoleucine transfer RNA from Escherichia coli. J Mol Biol 236(3):710–724
Acknowledgment
This research was supported by the Japan Science and Technology Agency (JST) Core Research for Evolutional Science and Technology (CREST) of Molecular Technologies to H.S., Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Young Scientists (B) to Y.G. (22750145), and Grants-in-Aid for JSPS Fellows to Y.I. (26-9576).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Iwane, Y., Katoh, T., Goto, Y., Suga, H. (2018). Artificial Division of Codon Boxes for Expansion of the Amino Acid Repertoire of Ribosomal Polypeptide Synthesis. In: Lemke, E. (eds) Noncanonical Amino Acids. Methods in Molecular Biology, vol 1728. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7574-7_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7574-7_2
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7573-0
Online ISBN: 978-1-4939-7574-7
eBook Packages: Springer Protocols