Abstract
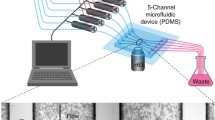
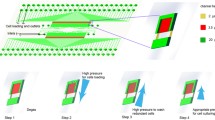
Time-lapse fluorescence imaging of yeast cells allows the study of multiple fluorescent targets in single cells, but is often hampered by the tedious cultivation using agar pads or glass bottom wells. Here, we describe the fabrication and operation of a microfluidic device for long-term imaging of yeast cells under constant or changing media conditions. The device allows acquisition of high quality images as cells are fixed in a two-dimensional imaging plane. Four yeast strains can be analyzed simultaneously over several days while up to four different media can be flushed through the chip. The microfluidic device does not rely on specialized equipment for its operation. To illustrate the use of the chip in DNA damage research, we show how common readouts for DNA damage or genomic instability behave upon induction with genotoxic chemicals (MMS, HU) or induction of a single double-strand break using induced CRISPR-Cas9 expression.
Similar content being viewed by others
References
Lisby M, Rothstein R, Mortensen UH (2001) Rad52 forms DNA repair and recombination centers during S phase. Proc Natl Acad Sci USA 98(15):8276–8282. doi:10.1073/pnas.121006298
Lisby M, Mortensen UH, Rothstein R (2003) Colocalization of multiple DNA double-strand breaks at a single Rad52 repair centre. Nat Cell Biol 5(6):572–577. doi:10.1038/ncb997
Lisby M, Barlow JH, Burgess RC, Rothstein R (2004) Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118(6):699–713. doi:10.1016/j.cell. 2004.08.015
Silva S, Gallina I, Eckert-Boulet N, Lisby M (2012) Live cell microscopy of DNA damage response in Saccharomyces cerevisiae. Methods Mol Biol 920:433. doi:10.1006/cbir.1999.0447. arXiv:1011.1669v3
Tkach JM, Yimit A, Lee AY, Riffle M, Costanzo M, Jaschob D, Hendry JA, Ou J, Moffat J, Boone C, Davis TN, Nislow C, Brown GW (2012) Dissecting {DNA} damage response pathways by analysing protein localization and abundance changes during {DNA} replication stress. Nat Cell Biol 14(9):966–976. doi:10.1038/ncb2549
Gasch AP, Huang M, Metzner S, Botstein D, Elledge SJ, Brown PO (2001) Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol Biol Cell 12(10):2987–3003. doi:10.1091/mbc.12.10.2987
Birrell GW, Brown JA, Wu HI, Giaever G, Chu AM, Davis RW, Brown JM (2002) Transcriptional response of Saccharomyces cerevisiae to DNA-damaging agents does not identify the genes that protect against these agents. Proc Natl Acad Sci USA 99(13):8778–8783. doi:10.1073/ pnas.132275199
Johnson SA, You Z, Hunter T (2007) Monitoring ATM kinase activity in living cells. DNA Repair 6(9):1277–1284. doi:10.1016/j.dnarep.2007.02. 025
Burgess A, Lorca T, Castro A (2012) Quantitative live imaging of endogenous DNA replication in mammalian cells. PLoS One 7(9). doi:10. 1371/journal.pone.0045726
Bush A, Chernomoretz A, Yu R, Gordon A, Colman-Lerner A (2012) Using cell-ID 1.4 with R for microscope-based cytometry. Curr Protoc Mol Biol (Suppl 100):1–27. doi:10.1002/0471142727.mb1418s100
Duncombe TA, Tentori AM, Herr AE (2015) Microfluidics: reframing biological enquiry. Nat Rev Mol Cell Biol 16(9):554–567. doi:10.1038/ nrm4041
Dénervaud N, Becker J, Delgado-Gonzalo R, Damay P, Rajkumar AS, Unser M, Shore D, Naef F, Maerkl SJ (2013) A chemostat array enables the spatio-temporal analysis of the yeast proteome. Proc Natl Acad Sci USA 110(39):15842–15847. doi:10.1073/pnas.1308265110
Frey O, Rudolf F, Schmidt GW, Hierlemann A (2015) Versatile, simple-to-use microfluidic cell-culturing chip for long-term, high-resolution, time-lapse imaging. Anal Chem 87(8):4144–4151. doi:10.1021/ac504611t
Ferry MS, Razinkov IA, Hasty J (2011) Microfluidics for synthetic biology: from design to execution. In: Methods in enzymology, vol 497. Elsevier, Amsterdam, pp 295–372
Lang M, Rudolf F, Stelling J (2012) Use of YouScope to implement systematic microscopy protocols. In: Current protocols in molecular biology, vol 14. Wiley, New York, pp 1–23. doi:10.1002/0471142727. mb1421s98
Mayer C, Dimopoulos S, Rudolf F, Stelling J (2013) Using CellX to quantify intracellular events. In: Current protocols in molecular biology, vol 14, pp 1–20. doi:10.1002/0471142727.mb1422s101
Acknowledgements
This work was supported by the FP7 grant no 28995 ITN ISOLATE and the Ambizione grant 142440 of the Swiss National Science Foundation to Olivier Frey. We thank Andreas Hierlemann and Jörg Stelling for their support. We acknowledge Mjriam Bächler and Andreas Cuny for testing the microfluidic device and Jennifer Ewald and Tania Roberts for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Schmidt, G.W., Frey, O., Rudolf, F. (2018). The CellClamper: A Convenient Microfluidic Device for Time-Lapse Imaging of Yeast. In: Muzi-Falconi, M., Brown, G. (eds) Genome Instability. Methods in Molecular Biology, vol 1672. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7306-4_36
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7306-4_36
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7305-7
Online ISBN: 978-1-4939-7306-4
eBook Packages: Springer Protocols