Abstract
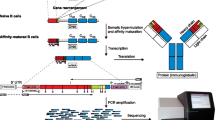
Methods for tracking B-cell repertoires and clonal history in normal and malignant B-cells based on immunoglobulin variable region (IGV) gene analysis have developed rapidly with the advent of massive parallel next-generation sequencing (mpNGS) protocols. mpNGS permits a depth of analysis of IGV genes not hitherto feasible, and presents challenges of bioinformatics analysis, which can be readily met by current pipelines. This strategy offers a potential resolution of B-cell usage at a depth that may capture fully the natural state, in a given biological setting. Conventional methods based on RT-PCR amplification and Sanger sequencing are also available where mpNGS is not accessible. Each method offers distinct advantages. Conventional methods for IGV gene sequencing are readily adaptable to most laboratories and provide an ease of analysis to capture salient features of B-cell use. This chapter describes two methods in detail for analysis of IGV genes, mpNGS and conventional RT-PCR with Sanger sequencing.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Stevenson F, Sahota S, Zhu D, Ottensmeier C, Chapman C, Oscier D, Hamblin T (1998) Insight into the origin and clonal history of B-cell tumors as revealed by analysis of immunoglobulin variable region genes. Immunol Rev 162:247–259
Georgiou G, Ippolito GC, Beausang J, Busse CE, Wardemann H, Quake SR (2014) The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat Biotechnol 32(2):158–168. doi:10.1038/nbt.2782
Tonegawa S (1983) Somatic generation of antibody diversity. Nature 302(5909):575–581
Cook GP, Tomlinson IM (1995) The human immunoglobulin VH repertoire. Immunol Today 16(5):237–242. doi:10.1016/0167-5699(95)80166-9
MacLennan IC (1994) Germinal centers. Annu Rev Immunol 12:117–139. doi:10.1146/annurev.iy.12.040194.001001
Boyd SD, Gaeta BA, Jackson KJ, Fire AZ, Marshall EL, Merker JD, Maniar JM, Zhang LN, Sahaf B, Jones CD, Simen BB, Hanczaruk B, Nguyen KD, Nadeau KC, Egholm M, Miklos DB, Zehnder JL, Collins AM (2010) Individual variation in the germline Ig gene repertoire inferred from variable region gene rearrangements. J Immunol 184(12):6986–6992. doi:10.4049/jimmunol.1000445
Glanville J, Kuo TC, von Budingen HC, Guey L, Berka J, Sundar PD, Huerta G, Mehta GR, Oksenberg JR, Hauser SL, Cox DR, Rajpal A, Pons J (2011) Naive antibody gene-segment frequencies are heritable and unaltered by chronic lymphocyte ablation. Proc Natl Acad Sci U S A 108(50):20066–20071. doi:10.1073/pnas.1107498108
Briney BS, Willis JR, McKinney BA, Crowe JE Jr (2012) High-throughput antibody sequencing reveals genetic evidence of global regulation of the naive and memory repertoires that extends across individuals. Genes Immun 13(6):469–473. doi:10.1038/gene.2012.20
Benichou J, Glanville J, Prak ET, Azran R, Kuo TC, Pons J, Desmarais C, Tsaban L, Louzoun Y (2013) The restricted DH gene reading frame usage in the expressed human antibody repertoire is selected based upon its amino acid content. J Immunol 190(11):5567–5577. doi:10.4049/jimmunol.1201929
Weston-Bell N, Townsend M, Di Genova G, Forconi F, Sahota SS (2009) Defining origins of malignant B cells: a new circulating normal human IgM(+)D(+) B-cell subset lacking CD27 expression and displaying somatically mutated IGHV genes as a relevant memory population. Leukemia 23(11):2075–2080. doi:10.1038/leu.2009.178
Cowan G, Weston-Bell NJ, Bryant D, Seckinger A, Hose D, Zojer N, Sahota SS (2015) Massive parallel IGHV gene sequencing reveals a germinal center pathway in origins of human multiple myeloma. Oncotarget 6(15):13229–13240. doi:10.18632/oncotarget.3644
Larimore K, McCormick MW, Robins HS, Greenberg PD (2012) Shaping of human germline IgH repertoires revealed by deep sequencing. J Immunol 189(6):3221–3230. doi:10.4049/jimmunol.1201303
Shugay M, Britanova OV, Merzlyak EM et al (2014) Towards error-free profiling of immune repertoires. Nat Meth 11:653–655. doi:10.1038/nmeth.2960
Weston-Bell NJ, Tapper W, Gibson J, Bryant D, Moreno Y, John M, Ennis S, Kluin-Nelemans HC, Collins AR, Sahota SS (2016) Exome sequencing in classic hairy cell leukaemia reveals widespread variation in acquired somatic mutations between individual tumours apart from the signature BRAF V(600)E lesion. PLoS One 11(2):e0149162. doi:10.1371/journal.pone.0149162
Corkum CP, Ings DP, Burgess C, Karwowska S, Kroll W, Michalak TI (2015) Immune cell subsets and their gene expression profiles from human PBMC isolated by Vacutainer Cell Preparation Tube (CPT) and standard density gradient. BMC Immunol 16:48. doi:10.1186/s12865-015-0113-0
Autissier P, Soulas C, Burdo TH, Williams KC (2010) Evaluation of a 12-color flow cytometry panel to study lymphocyte, monocyte, and dendritic cell subsets in humans. Cytometry A 77(5):410–419. doi:10.1002/cyto.a.20859
Terstappen LW, Johnsen S, Segers-Nolten IM, Loken MR (1990) Identification and characterization of plasma cells in normal human bone marrow by high-resolution flow cytometry. Blood 76(9):1739–1747
Clark P, Normansell DE, Innes DJ, Hess CE (1986) Lymphocyte subsets in normal bone marrow. Blood 67(6):1600–1606
Acknowledgement
This work was funded by Bloodwise (UK).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Weston-Bell, N.J., Cowan, G., Sahota, S.S. (2017). Tracking B-Cell Repertoires and Clonal Histories in Normal and Malignant Lymphocytes. In: Calado, D. (eds) Germinal Centers. Methods in Molecular Biology, vol 1623. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7095-7_21
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7095-7_21
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7094-0
Online ISBN: 978-1-4939-7095-7
eBook Packages: Springer Protocols