Abstract
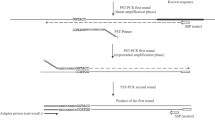
A method is described that uses arbitrarily primed PCR followed by many cycles of amplification under stringent conditions and selection by computational means to obtain a set of sequence tags that can be used for the comparison of metagenomes. Relative to unselective shot-gun sequencing, the results are small data sets that can be csompared electronically or plotted as scattergrams that are simple to interpret. The method can be used to compare groups of samples of any size to build in-house databases from which, for example, the provenance of trace soil samples may be inferred. The method also allows for selection of primers with locus-specificity and an example is given in which a South Australian sequence-related to a Portuguese thermophile (Rubrobacter radiotolerans) is extracted and tested on a set of soils.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities. Annu Rev Genet 38:525–552. doi:10.1146/annurev.genet.38.072902.091216
Wooley JC, Godzik A, Friedberg I (2010) A primer on metagenomics. PLoS Comput Biol 6(2):e1000667. doi:10.1371/journal.pcbi.1000667
Sharpton TJ (2014) An introduction to the analysis of shotgun metagenomics data. Front Plant Sci 16:209. doi:10.3389/fpls.2014.00209
Oulas A, Pavloud C, Polymenakou P, Pavlopoulos GS, Papanikolaou N, Kotoulas G, Arvanitidis C, Iliopoulos I (2015) Metagenomics: tools and insights for analyzing next-generation sequencing data derived from biodiversity studies. Bioinform Biol Insights 9:75–88. doi:10.4137/BBI.S12462
Khodakova AS, Smith RJ, Burgoyne L, Abarno D, Linacre A (2014) Random whole metagenomic sequencing for forensic discrimination of soils. PLoS One 9:e104996. doi:10.1371/journal.pone.0104996
Waters JM, Eariss G, Yeadon PJ, Kirkbride KP, Burgoyne LA, Catcheside DEA (2012) Arbitrary single primer amplification of trace DNA substrates yields sequence content profiles that are discriminatory and reproducible. Electrophoresis 33:492–498. doi:10.1002/elps.201100359
Khodakova AS, Burgoyne L, Abarno D, Linacre A (2013) Forensic analysis of soils using single arbitrarily primed amplification and high throughput sequencing. Forensic Sci Int Genet 4:e39–e40. doi:10.1016/j.fsigss.2013.10.019
Burgoyne L, Koh L, Catcheside D (2015) A study of one soil, its relatives and contaminants by arbitrary primed PCR with 50mer based analysis. Forensic Sci Int Genet 5:e503–e505. doi:10.1016/j.fsigss.2015.09.199
Egas C, Barroso C, Froufe HJC, Pacheco J, Albuquerque L, da Costa MS (2014) Complete genome sequence of the radiation-resistant bacterium Rubrobacter radiotolerans RSPS-4. Stand Genomic Sci 9:1062–1075. doi:10.4056/sigs.5661021
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Burgoyne, L., Koh, L.Y., Catcheside, D. (2017). Arbitrarily Primed PCR for Comparison of Meta Genomes and Extracting Useful Loci from Them. In: Domingues, L. (eds) PCR. Methods in Molecular Biology, vol 1620. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-7060-5_18
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7060-5_18
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-7059-9
Online ISBN: 978-1-4939-7060-5
eBook Packages: Springer Protocols