Abstract
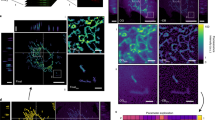
Mitochondria are shaped by opposing fission (division) and fusion events. Mounting evidence indicates that mitochondrial shape influences numerous aspects of mitochondrial function, including ATP production, Ca2+ buffering, and quality control. Despite the recognized importance of mitochondrial dynamics, the literature is rife with subjective, categorical estimates of mitochondrial morphology, preventing reliable comparison of results between groups. This chapter describes stringent, but easily implemented methods for quantification of mitochondrial shape changes using the open-source software package ImageJ. While we provide examples for analysis of epifluorescence images of cultured primary neurons, these methods are easily generalized to other cell types and imaging techniques.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Popov V, Medvedev NI, Davies HA, Stewart MG (2005) Mitochondria form a filamentous reticular network in hippocampal dendrites but are present as discrete bodies in axons: a three-dimensional ultrastructural study. J Comp Neurol 492(1):50–65. doi:10.1002/cne.20682
De Stefani D, Rizzuto R, Pozzan T (2016) Enjoy the trip: calcium in mitochondria back and forth. Annu Rev Biochem. doi:10.1146/annurev-biochem-060614-034216
Bertholet AM, Delerue T, Millet AM, Moulis MF, David C, Daloyau M, Arnaune-Pelloquin L, Davezac N, Mils V, Miquel MC, Rojo M, Belenguer P (2016) Mitochondrial fusion/fission dynamics in neurodegeneration and neuronal plasticity. Neurobiol Dis 90:3–19. doi:10.1016/j.nbd.2015.10.011
Alexander C, Votruba M, Pesch UE, Thiselton DL, Mayer S, Moore A, Rodriguez M, Kellner U, Leo-Kottler B, Auburger G, Bhattacharya SS, Wissinger B (2000) OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat Genet 26(2):211–215
Zuchner S, Mersiyanova IV, Muglia M, Bissar-Tadmouri N, Rochelle J, Dadali EL, Zappia M, Nelis E, Patitucci A, Senderek J, Parman Y, Evgrafov O, Jonghe PD, Takahashi Y, Tsuji S, Pericak-Vance MA, Quattrone A, Battaloglu E, Polyakov AV, Timmerman V, Schroder JM, Vance JM (2004) Mutations in the mitochondrial GTPase mitofusin 2 cause Charcot-Marie-Tooth neuropathy type 2A. Nat Genet 36(5):449–451
Waterham HR, Koster J, van Roermund CW, Mooyer PA, Wanders RJ, Leonard JV (2007) A lethal defect of mitochondrial and peroxisomal fission. N Engl J Med 356(17):1736–1741
Sheffer R, Douiev L, Edvardson S, Shaag A, Tamimi K, Soiferman D, Meiner V, Saada A (2016) Postnatal microcephaly and pain insensitivity due to a de novo heterozygous DNM1L mutation causing impaired mitochondrial fission and function. Am J Med Genet A. doi:10.1002/ajmg.a.37624
Koch J, Feichtinger RG, Freisinger P, Pies M, Schrodl F, Iuso A, Sperl W, Mayr JA, Prokisch H, Haack TB (2016) Disturbed mitochondrial and peroxisomal dynamics due to loss of MFF causes Leigh-like encephalopathy, optic atrophy and peripheral neuropathy. J Med Genet 53(4):270–278. doi:10.1136/jmedgenet-2015-103500
Shamseldin HE, Alshammari M, Al-Sheddi T, Salih MA, Alkhalidi H, Kentab A, Repetto GM, Hashem M, Alkuraya FS (2012) Genomic analysis of mitochondrial diseases in a consanguineous population reveals novel candidate disease genes. J Med Genet 49(4):234–241. doi:10.1136/jmedgenet-2012-100836
Fahrner JA, Liu R, Perry MS, Klein J, Chan DC (2016) A novel de novo dominant negative mutation in DNM1L impairs mitochondrial fission and presents as childhood epileptic encephalopathy. Am J Med Genet A. doi:10.1002/ajmg.a.37721
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675
Lim IA, Merrill MA, Chen Y, Hell JW (2003) Disruption of the NMDA receptor-PSD-95 interaction in hippocampal neurons with no obvious physiological short-term effect. Neuropharmacology 45(6):738–754
Sternberger SR (1983) Biomedical image processing. IEEE Comput 18:22–34
Frangi AF, Niessen WJ, Vincken KL, Viergever MA (1998) Multiscale vessel enhancement filtering. In: Wells WM, Colchester A, Delp SL (eds) Medical image computing and computer-assisted intervention, Lecture notes in computer sciences, vol 1496. Springer, Berlin, pp 130–137
Sato Y, Nakajima S, Shiraga N, Atsumi H, Yoshida S, Koller T, Gerig G, Kikinis R (1998) Three-dimensional multi-scale line filter for segmentation and visualization of curvilinear structures in medical images. Med Image Anal 2(2):143–168
Acknowledgments
This work is currently supported by NIH grants NS056244 and NS087908 to S.S. We thank past and present members of the laboratory for providing critical feedback for development of the methods described in this chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendix—Morphometry Macro
Appendix—Morphometry Macro
var ch = 0; // channel to be analyzed for RGB images/** Measure mitochondrial morphology in the current selection* Ctrl+Shift+O closes current and opens next image*/macro “Morphometry [F7]” {title = getTitle();morphometry(title, false); // not batch mode}/** Batch-apply a set of “named” ROIs to analyze images with that file name*/macro “Batch Morphometry [F8]” {dir = getDirectory(“Select an image directory”);while (roiManager(“Count”) == 0)waitForUser(“Please open named ROIs into ROI manager”);prevName = imgName = “”;n = roiManager(“Count”);for (i = 0; i < n; ++i) { // loop through the ROI Manager tableprevName = imgName;imgName = call(“ij.plugin.frame.RoiManager.getName”, i);if (isOpen(imgName)) { // named image is openselectWindow(imgName);} else { // done with current image, close and open nextif (isOpen(prevName)) {selectWindow(prevName);close();}open(dir + imgName);}roiManager(“Select”, i);morphometry(imgName, true); // batch mode}}function morphometry(title, batchMode) {while (ch < 1 || ch > 3) { /* RGB channel not yet selected, initialize; reinstall macro to change channel */ch = getNumber(“Analyze RGB channel(1-3):”, 1);run(“Set Measurements...”, “decimal=5 area perimeter fit”);print(“image\t n\t area2\t area-weighted ff\t form factor\t aspect ratio\t length”); /* header for results table */}if (bitDepth == 24) // RGB imagerun(“Make Composite”);if (isOpen(“Binary”)) {selectWindow(“Binary”);close();} // close previous working imageif (isOpen(“Skeleton”)) {selectWindow(“Skeleton”);close();} // close previous working imageselectWindow(title);if (selectionType() == -1) // no selectionrun(“Select All”);if (!batchMode) {roiManager(“Add”); // save selection to ROI Manager for batch processinglast = roiManager(“Count”) - 1;roiManager(“Select”, last);roiManager(“Rename”, title);/* roiManager(“Save”, File.directory + “named_ROIs.zip”); */ /* un-comment to save ROIs automatically */}// copy selection to new window and clear outsidesetSlice(ch); // ignored if grayscalerun(“Duplicate...”, “title=Binary”);run(“Make Inverse”);if (selectionType != -1) { // outside of ROI is selectedrun(“Duplicate...”, “ “); // make a mask of the backgroundrun(“Convert to Mask”);run(“Create Selection”);run(“Make Inverse”);roiManager(“Add”);close();n = roiManager(“Count”);roiManager(“Select”, n - 1);getRawStatistics(_area, backG); // mean is backgroundsetColor(backG);run(“Restore Selection”); // fill outside of selection with backgroundfill();run(“Gaussian Blur...”, “radius=64”); // smooth abrupt background transitionroiManager(“Delete”); /* delete masking selection (ROI manager has cell selections) */}run(“Select None”);// subtract background and thresholdrun(“Subtract Background...”, “rolling=50”); /* non-destructive filter even if already applied */run(“Make Binary”);// also try other threshold methods included with Fiji, e.g.: run(“Auto Threshold”, “method=Li white”);// create Results table of metrics, one line/particlerun(“Analyze Particles...”, “size=9-Infinity circularity=0.00-1.00 show=Masks pixel clear”);awff = ff = ar = sum_a = a2 = len = 0;for (i = 0; i < nResults; i++) { // for every particle in tablea = getResult(“Area”, i);p = getResult(“Perim.”, i);ar += getResult(“Major”, i) / getResult(“Minor”, i); /* aspect ratio = length / width */sum_a += a;a2 += a * a; // area2 = a2 / (sum_a * sum_a)awff += b = (p * p) / (4 * 3.14159265358979); // awff = ff * (a / sum_area)ff += b / a; // ff = p^2 / (4 * pi * a)}nParticles = nResults;// skeletonize to get lengthselectWindow(“Mask of Binary”); /* created by Analyze Particles .., excludes noise (< 9 pixels) */rename(“Skeleton”);run(“Skeletonize”);run(“Analyze Particles...”, “size=0-Infinity show=Nothing pixel clear”);for (i = 0; i < nResults; i++)len += getResult(“Area”, i);// average and outputa2 /= sum_a * sum_a;awff /= sum_a;ff /= nParticles;ar /= nParticles;len /= nResults;print(title + “\t “ + nParticles + “\t “ + a2 + “\t “ + awff + “\t “ + ff + “\t “ + ar + “\t “ + len);selectWindow(title);
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Merrill, R.A., Flippo, K.H., Strack, S. (2017). Measuring Mitochondrial Shape with ImageJ. In: Strack, S., Usachev, Y. (eds) Techniques to Investigate Mitochondrial Function in Neurons. Neuromethods, vol 123. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6890-9_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6890-9_2
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6888-6
Online ISBN: 978-1-4939-6890-9
eBook Packages: Springer Protocols