Abstract
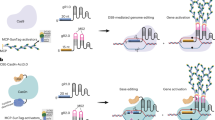
Multiplex CRISPR-Cas9 nuclease mediated genome editing is an efficient method for disrupting gene function in plants. Use of CRISPR-Cas9 has escalated rapidly in recent years and is expected to become routine practice in molecular biology and related fields of research. Due to the relatively novel and widespread adoption of this technology, first-time users may not have regular access to experienced guidance or technical support from peers or mentors. Here, we offer guidance and technical support in the form of a detailed and tested protocol for simultaneous targeting of three separate loci on the TRANSPARENT TESTA 4 (TT4) gene in Arabidopsis thaliana using multiplex CRISPR-Cas9. Although we target multiple loci on a single gene in Arabidopsis, the same approach can be used to target multiple genes or alleles in other plant species as well. We recommend the use of a molecular toolkit to streamline the process and make recommendations for this type of approach. The protocol starts with an overview of the reagents and covers designing of gRNAs and assembly of components into a final T-DNA delivery molecule through Golden Gate cloning and Multisite Gateway LR recombination.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. doi:10.1126/science.1225829
Belhaj K, Chaparro-Garcia A, Kamoun S, Patron NJ, Nekrasov V (2015) Editing plant genomes with CRISPR/Cas9. Curr Opin Biotechnol 32:76–84
Baltes NJ, Voytas DF (2015) Enabling plant synthetic biology through genome engineering. Trends Biotechnol 33(2):120–131
Deltcheva E, Chylinski K, Sharma C, Gonzales K, Chao Y, Pirzada Z, Eckert M, Vogel J, Charpentier E (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471:602–609
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826. doi:10.1126/science.1232033
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823. doi:10.1126/science.1231143
Li J-F, Norville J, Aach J, McCormack M, Zhang D, Bush J, Chruch G, Sheen J (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691
Carroll D (2011) Genome engineering with zinc-finger nucleases. Genetics 188(4):773–782. doi:10.1534/genetics.111.131433
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186(2):757–761. doi:10.1534/genetics.110.120717
Weber J, Ollinger R, Mathias F (2015) CRISPR/Cas9 somatic multiplex-mutagenesis for high-throughput functional cancer genomics in mice. Proc Natl Acad Sci U S A 112(45):13982–13987
Zhou H, Liu B, Weeks D, Spalding M, Yang B (2014) Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res. doi:10.1093/nar/gku806
Ran A, Hsu P, Lin C-Y, Gootenberg J, Konermann S, Trevino A, Scott D, Inoue A, Matoba S, Zhang Y, Zhang F (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
Lowder L, Zhang D, Baltes N, Paul J III, Tang X, Zheng X, Voytas D, Hsieh T-F, Zhang Y, Qi Y (2015) A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol 169:971–985
Curtis MD, Grossniklaus U (2003) A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 133(2):462–469. doi:10.1104/pp.103.027979
Graham D, Root D (2015) Resources for the design of CRISPR gene editing experiments. Genome Biol 16:260
Shirley B, Kubasek W, Storz G, Bruggemann E, Koornneef M, Ausubel F, Goodman H (1995) Analysis of Arabidopsis mutants deficient in flavonoid biosynthesis. Plant J 8(5):659–671
Sun W, Meng X, Liang L, Jiang W, Huang Y, He J, Hu H, Almqvist J, Gao X, Wang L (2015) Molecular and biochemical analysis of chalcone synthase from Freesia hybrid in flavonoid biosynthetic pathway. PLoS One. doi:10.1371/journal.pone.0119054
Zhang F, Maeder M, Unger-Wallace E, Hoshaw J, Reyon D, Christian M, Li X, Pierick C, Dobbs D, Peterson T, Joung K, Voytas D (2010) High frequency targeted mutagenesis in Arabidopsis thaliana using zing finger nucleases. Proc Natl Acad Sci U S A 107(26):12028–12033
Qi Y, Zhang Y, Zhang F, Baller J, Cleland S, Ryu Y, Starker C, Voytas D (2013) Increasing frequencies of site-specific mutagenesis and gene targeting in Arabidopsis by manipulating DNA repair pathways. Genome Res 23:547–554
Feng Z, Mao Y, Zhang B, Wei P, Yang D-L, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L, Liu X, Zhu J-K (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc Natl Acad Sci U S A 111(12):4632–4637
Doench J, Hartenian E, Graham D, Tothova Z, Hegde M, Smith I, Sullender M, Ebert B, Xavier R, Root D (2014) Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32:1262–1267
Wang T, Wei J, Sabatini D, Lander E (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343:80–84
Singh R, Kuscu C, Quinlan A, Qi Y, Adli M (2015) Cas9-chromatin binding information enables more accurate CRISPR off-target prediction. Nucleic Acids Res 43(18). doi:10.1093/nar/gkv575
Tsai S, Zheng Z, Nguyen N, Liebers M, Topkar V, Thapar V, Wyvekens N, Khayter C, Iafrate J, Le L, Aryee M, Joung K (2015) GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat Biotechnol 33(2):187–197
Vaucheret H (2006) Post-transcriptional small RNA pathways in plants: mechanisms and regulations. Genes Dev 20:759–771
De Wilde C, Van Houdt H, De Buck S, Angenon G, De Jaeger G (2000) Depicker A. Plants as bioreactors for protein production: avoiding the problem of transgene silencing 43:347–359
Nishimasu H, Ran A, Hsu P, Konermann S, Shehata S, Dohmae N, Ishitani R (2014) Crystal structure of Cas9. Cell 156:935–949
Acknowledgments
This work is supported by startup funds from East Carolina University and a Collaborative Funding Grant from North Carolina Biotechnology Center and Syngenta (2016-CFG-8003) to Y.Q.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Lowder, L., Malzahn, A., Qi, Y. (2017). Rapid Construction of Multiplexed CRISPR-Cas9 Systems for Plant Genome Editing. In: Shan, L., He, P. (eds) Plant Pattern Recognition Receptors. Methods in Molecular Biology, vol 1578. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6859-6_25
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6859-6_25
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6858-9
Online ISBN: 978-1-4939-6859-6
eBook Packages: Springer Protocols