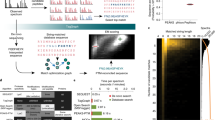
Abstract
In the past decade, proteomics and mass spectrometry have taken tremendous strides forward, particularly in the life sciences, spurred on by rapid advances in technology resulting in generation and conglomeration of vast amounts of data. Though this has led to tremendous advancements in biology, the interpretation of the data poses serious challenges for many practitioners due to the immense size and complexity of the data. Furthermore, the lack of annotation means that a potential gold mine of relevant biological information may be hiding within this data. We present here a simple and intuitive workflow for the research community to investigate and mine this data, not only to extract relevant data but also to segregate usable, quality data to develop hypotheses for investigation and validation. We apply an MS evidence workflow for verifying peptides of proteins from one’s own data as well as publicly available databases. We then integrate a suite of freely available bioinformatics analysis and annotation software tools to identify homologues and map putative functional signatures, gene ontology and biochemical pathways. We also provide an example of the functional annotation of missing proteins in human chromosome 7 data from the NeXtProt database, where no evidence is available at the proteomic, antibody, or structural levels. We give examples of protocols, tools and detailed flowcharts that can be extended or tailored to interpret and annotate the proteome of any novel organism.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Laukens K, Naulaerts S, Berghe WV (2015) Bioinformatics approaches for the functional interpretation of protein lists: from ontology term enrichment to network analysis. Proteomics 15(5-6):981–996. doi:10.1002/pmic.201400296
Kumar C, Mann M (2009) Bioinformatics analysis of mass spectrometry-based proteomics data sets. FEBS Lett 583(11):1703–1712. doi:10.1016/j.febslet.2009.03.035
Carnielli CM, Winck FV, Paes Leme AF (2015) Functional annotation and biological interpretation of proteomics data. Biochim Biophys Acta 1854(1):46–54. doi:10.1016/j.bbapap.2014.10.019
Draghici S, Khatri P, Martins RP, Ostermeier GC, Krawetz SA (2003) Global functional profiling of gene expression. Genomics 81(2):98–104. doi: 10.1016/S0888-7543(02)00021-6
Khatri P, Draghici S (2005) Ontological analysis of gene expression data: current tools, limitations, and open problems. Bioinformatics 21(18):3587–3595. doi:10.1093/bioinformatics/bti565
Goeman JJ, Buhlmann P (2007) Analyzing gene expression data in terms of gene sets: methodological issues. Bioinformatics 23(8):980–987. doi:10.1093/bioinformatics/btm051
Deutsch EW, Albar JP, Binz PA, Eisenacher M, Jones AR, Mayer G, Omenn GS, Orchard S, Vizcaino JA, Hermjakob H (2015) Development of data representation standards by the human proteome organization proteomics standards initiative. J Am Med Inform Assoc 22(3):495–506. doi:10.1093/jamia/ocv001
Haga SW, Wu HF (2014) Overview of software options for processing, analysis and interpretation of mass spectrometric proteomic data. J Mass Spectrom 49(10):959–969. doi:10.1002/jms.3414
Omenn GS, Lane L, Lundberg EK, Beavis RC, Nesvizhskii AI, Deutsch EW (2015) Metrics for the Human Proteome Project 2015: Progress on the Human Proteome and Guidelines for High-Confidence Protein Identification. J Proteome Res 14(9):3452–3460. doi:10.1021/acs.jproteome.5b00499
Islam MT, Garg G, Hancock WS, Risk BA, Baker MS, Ranganathan S (2014) Protannotator: a semiautomated pipeline for chromosome-wise functional annotation of the "missing" human proteome. J Proteome Res 13(1):76–83. doi:10.1021/pr400794x
Ranganathan S, Khan JM, Garg G, Baker MS (2013) Functional annotation of the human chromosome 7 "missing" proteins: a bioinformatics approach. J Proteome Res 12(6):2504–2510. doi:10.1021/pr301082p
Islam MT, Mohamedali A, Garg G, Khan JM, Gorse AD, Parsons J, Marshall P, Ranganathan S, Baker MS (2013) Unlocking the puzzling biology of the black Perigord truffle Tuber melanosporum. J Proteome Res 12(12):5349–5356. doi:10.1021/pr400650c
Gaudet P, Argoud-Puy G, Cusin I, Duek P, Evalet O, Gateau A, Gleizes A, Pereira M, Zahn-Zabal M, Zwahlen C, Bairoch A, Lane L (2013) neXtProt: organizing protein knowledge in the context of human proteome projects. J Proteome Res 12(1):293–298. doi:10.1021/pr300830v
Full Chromosome Reports from neXtProt. ftp://ftp.nextprot.org/pub/current_release/chr_reports. Accessed 27 October 2016
Simplified chromosome reports from neXtProt. ftp://ftp.nextprot.org/pub/current_release/custom/hpp. Accessed 27 October 2016
UniProt Consortium (2012) Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucleic Acids Res 40(Database issue):D71–75. doi:10.1093/nar/gkr981
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28(1):235–242. doi: 10.1093/nar/28.1.235
Protein Data Bank (PDB) http://www.rcsb.org/pdb/download/download.do. Accessed 27 October 2016
Chen C, Li Z, Huang H, Suzek BE, Wu CH (2013) A fast Peptide Match service for UniProt Knowledgebase. Bioinformatics 29(21):2808-2809. doi: 10.1093/bioinformatics/btt484
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. doi:10.1016/S0022-2836(05)80360-2
NCBI BLAST ftp://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/LATEST/. Accessed 27 October 2016
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33 (Web Server issue):W116-120. doi:10.1093/nar/gki442
InterProScan. http://www.ebi.ac.uk/Tools/pfa/iprscan5/ http://www.ebi.ac.uk/interpro/search/sequence-search. Accessed 27 October 2016
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35 (Web Server issue):W182-185. doi:10.1093/nar/gkm321
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39 (Web Server issue):W316-322. doi:10.1093/nar/gkr483
Martens L, Hermjakob H, Jones P, Adamski M, Taylor C, States D, Gevaert K, Vandekerckhove J, Apweiler R (2005) PRIDE: the proteomics identifications database. Proteomics 5(13):3537–3545. doi:10.1002/pmic.200401303
Craig R, Cortens JP, Beavis RC (2004) Open source system for analyzing, validating, and storing protein identification data. J Proteome Res 3(6):1234–1242. doi:10.1021/pr049882h
Schaab C, Geiger T, Stoehr G, Cox J, Mann M (2012) Analysis of high accuracy, quantitative proteomics data in the MaxQB database. Molecular & cellular proteomics : MCP 11 (3):M111 014068. doi:10.1074/mcp.M111.014068
Wilhelm M, Schlegl J, Hahne H, Gholami AM, Lieberenz M, Savitski MM, Ziegler E, Butzmann L, Gessulat S, Marx H (2014) Mass-spectrometry-based draft of the human proteome. Nature 509(7502):582–587. doi: 10.1038/nature13319
Nesvizhskii AI, Aebersold R (2005) Interpretation of shotgun proteomic data: the protein inference problem. Molecular & cellular proteomics : MCP 4(10):1419–1440. doi:10.1074/mcp.R500012-MCP200
InterProScan Search. http://www.ebi.ac.uk/interpro/search/sequence-search. Accessed 27 October 2016
KOBAS 2.0. http://kobas.cbi.pku.edu.cn. Accessed 27 October 2016
Scrivano G GNU Wget. http://www.gnu.org/software/wget/. Accessed 27 October 2016
Stenberg D curl. http://curl.haxx.se/. Accessed 27 October 2016
Deutsch EW, Sun Z, Campbell D, Kusebauch U, Chu CS, Mendoza L, Shteynberg D, Omenn GS, Moritz RL (2015) State of the Human Proteome in 2014/2015 As Viewed through PeptideAtlas: Enhancing Accuracy and Coverage through the AtlasProphet. J Proteome Res 14(9):3461–3473. doi:10.1021/acs.jproteome.5b00500
Hulstaert N, Reisinger F, Rameseder J, Barsnes H, Vizcaino JA, Martens L (2013) Pride-asap: automatic fragment ion annotation of identified PRIDE spectra. Journal of proteomics 95:89–92. doi:10.1016/j.jprot.2013.04.011
Sadygov RG, Cociorva D, Yates JR 3rd (2004) Large-scale database searching using tandem mass spectra: looking up the answer in the back of the book. Nature methods 1(3):195–202. doi:10.1038/nmeth725
Craig R, Beavis RC (2004) TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20(9):1466–1467. doi:10.1093/bioinformatics/bth092
Protannotator. http://www.biolinfo.org/protannotator/human_Chr7.php. Accessed 27 October 2016
InterProScan Download and Requirements. https://github.com/ebi-pf-team/interproscan/wiki/HowToDownload AND https://github.com/ebi-pf-team/interproscan/wiki/InstallationRequirements. Accessed 27 October2016
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Islam, M.T., Mohamedali, A., Ahn, S.B., Nawar, I., Baker, M.S., Ranganathan, S. (2017). A Systematic Bioinformatics Approach to Identify High Quality Mass Spectrometry Data and Functionally Annotate Proteins and Proteomes. In: Keerthikumar, S., Mathivanan, S. (eds) Proteome Bioinformatics. Methods in Molecular Biology, vol 1549. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6740-7_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6740-7_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6738-4
Online ISBN: 978-1-4939-6740-7
eBook Packages: Springer Protocols