Abstract
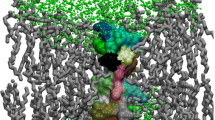
A great deal of research has been undertaken in order to discover antimicrobial peptides (AMPs) with unexploited mechanisms of action to counteract the health-threatening issues associated with bacterial resistance. The intrinsic effectiveness of AMPs is strongly influenced by their initial interactions with the bacterial cell membrane. Understanding these interactions in the atomistic details is important for the design of the less prone bacteria-resistant peptides. However, these studies always require labor-intensive and difficult steps. With this regard, modeling studies of the AMPs binding to simple lipid membrane systems, e.g., lipid bilayers, is of great advantage. In this chapter, we present an applicable step-by-step protocol to run the molecular dynamics (MD) simulation of the interaction between cyclo-RRWFWR (c-WFW) (a small cyclic AMP) and 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) lipid bilayer using the Groningen machine for chemical simulations (GROMACS) package. The protocol as described here may simply be optimized for other peptide–lipid systems of interest.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Cho W, Stahelin RV (2005) Membrane-protein interactions in cell signaling and membrane trafficking. Annu Rev Biophys Biomol Struct 34:119–151
Volinsky R, Kinnunen PK (2013) Oxidized phosphatidylcholines in membrane-level cellular signaling: from biophysics to physiology and molecular pathology. FEBS J 280:2806–2816
Peschel A, Sahl HG (2006) The co-evolution of host cationic antimicrobial peptides and microbial resistance. Nat Rev Microbiol 4:529–536
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395
Willumeit R, Kumpugdee M, Funari SS, Lohner K, Navas BP, Brandenburg K, Linser S, Andrä J (2005) Structural rearrangement of model membranes by the peptide antibiotic NK-2. Biochim Biophys Acta 1669:125–134
Pabst G, Grage SL, Danner-Pongratz S, Jing W, Ulrich AS, Watts A, Lohner K, Hickel A (2008) Membrane thickening by the antimicrobial peptide PGLa. Biophys J 95:5779–5788
Schmidt NW, Wong GC (2013) Antimicrobial peptides and induced membrane curvature: geometry, coordination chemistry, and molecular engineering. Curr Opin Solid State Mater Sci 17:151–163
McHenry AJ, Sciacca MF, Brender JR, Ramamoorthy A (2012) Does cholesterol suppress the antimicrobial peptide induced disruption of lipid raft containing membranes? Biochim Biophys Acta 1818:3019–3024
Bagheri M (2015) Cationic antimicrobial peptides (AMPs): thermodynamic characterization of peptide-lipid interactions and biological efficacy of surface-tethered peptides. ChemistryOpen 4:389–393
Arouri A, Dathe M, Blume A (2009) Peptide induced demixing in PG/PE lipid mixtures: a mechanism for the specificity of antimicrobial peptides towards bacterial membranes? Biochim Biophys Acta 1788:650–659
Kobayashi S, Chikushi A, Tougu S, Imura Y, Nishida M, Yano Y, Matsuzaki K (2004) Membrane translocation mechanism of the antimicrobial peptide buforin 2. Biochemistry 43:15610–15616
Nascimento JM, Oliveira MD, Franco OL, Migliolo L, de Melo CP, Andrade CA (2014) Elucidation of mechanisms of interaction of a multifunctional peptide Pa-MAP with lipid membranes. Biochim Biophys Acta 1838:2899–2909
Legrand B, Laurencin M, Sarkis J, Duval E, Mouret L, Hubert JF, Collen M, Vié V, Zatylny-Gaudin C, Henry J, Baudy-Floc’h M, Bondon A (2011) Structure and mechanism of action of a de novo antimicrobial detergent-like peptide. Biochim Biophys Acta 1808:106–116
Bechinger B (1999) The structure, dynamics and orientation of antimicrobial peptides in membranes by multidimensional solid-state NMR spectroscopy. Biochim Biophys Acta 1462:157–183
Nichols M, Kuljanin M, Nategholeslam M, Hoang T, Vafaei S, Tomberli B, Gray CG, DeBruin L, Jelokhani-Niaraki M (2013) Dynamic turn conformation of a short tryptophan-rich cationic antimicrobial peptide and its interaction with phospholipid membranes. J Phys Chem B 117:14697–14708
Bagheri M, Arasteh S, Haney EF, Hancock REW (2016) Tryptic stability of synthetic bactenecin derivatives is determined by the side chain length of cationic residues and the peptide conformation. J Med Chem 59:3079–3086
Nauli S, Farr S, Lee YJ, Kim HY, Faham S, Bowie JU (2007) Polymer-driven crystallization. Protein Sci 16:2542–2551
Bagheri M, Keller S, Dathe M (2011) Interaction of W-substituted analogs of cyclo-RRWFWR with bacterial lipopolysaccharides: the role of the aromatic cluster in antimicrobial activity. Antimicrob Agents Chemother 55:788–797
Scheinpflug K, Krylova O, Nikolenko H, Thurm C, Dathe M (2015) Evidence for a novel mechanism of antimicrobial action of a cyclic R-, W-rich hexapeptide. PLoS One 10:e0125056
Finger S, Kerth A, Dathe M, Blume A (2015) The efficacy of trivalent cyclic hexapeptides to induce lipid clustering in PG/PE membranes correlates with their antimicrobial activity. Biochim Biophys Acta 1848:2998–3006
Tieleman DP, Sansom MSP (2001) Molecular dynamics simulations of antimicrobial peptides: from membrane binding to trans-membrane channels. Int J Quantum Chem 83:166–179
Arasteh S, Bagheri M, Goliaei B (2014) Membrane selectivity of small cyclic antimicrobial hexapeptides studied by molecular dynamics simulations. J Pept Sci 20:S279–S280
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. Graphics 14:33–38
Allen WJ, Lemkul JA, Bevan DR (2009) GridMAT-MD: a grid-based membrane analysis tool for use with molecular dynamics. J Comput Chem 30:1952–1958
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem 25:1656–1676
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4:187–217
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc 118:11225–11236
Pearlman DA, Case DA, Caldwell JW, Ross WS, Cheatham TE III, DeBolt S, Ferguson D, Seibele G, Kollman P (1995) AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput Phys Commun 91:1–41
Gfeller D, Michielin O, Zoete V (2013) SwissSidechain: a molecular and structural database of non-natural sidechains. Nucleic Acids Res 41:D327–D332
Klauda JB, Venable RM, Freites JA, O’Connor JW, Tobias DJ, Mondragon-Ramirez C, Vorobyov I, MacKerell AD Jr, Pastor RW (2010) Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J Phys Chem B 114:7830–7843
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Cheng A, Merz KM Jr (1996) Application of the Nosé–Hoover chain algorithm to the study of protein dynamics. J Phys Chem 100:1927–1937
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126:014101
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190
Kucerka N, Tristram-Nagle S, Nagle JF (2005) Structure of fully hydrated fluid phase lipid bilayers with monounsaturated chains. J Membr Biol 208:193–202
Acknowledgments
The financial supports to MB from the Iran National Science Foundation (reference No. 94012757), the Iran’s National Elites Foundation (reference No. 15/45362-1392) and University of Tehran are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Arasteh, S., Bagheri, M. (2017). Molecular Dynamics Simulation and Analysis of the Antimicrobial Peptide–Lipid Bilayer Interactions. In: Hansen, P. (eds) Antimicrobial Peptides. Methods in Molecular Biology, vol 1548. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6737-7_8
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6737-7_8
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6735-3
Online ISBN: 978-1-4939-6737-7
eBook Packages: Springer Protocols