Abstract
Bacterial blood stream infection (BSI) potentially leads to life-threatening clinical conditions and medical emergencies such as severe sepsis, septic shock, and multi organ failure syndrome. Blood culturing is currently the gold standard for the identification of microorganisms and, although it has been automated over the decade, the process still requires 24–72 h to complete. This long turnaround time, especially for the identification of antimicrobial resistance, is driving the development of rapid molecular diagnostic methods. Rapid detection of microbial pathogens in blood related to bloodstream infections will allow the clinician to decide on or adjust the antimicrobial therapy potentially reducing the morbidity, mortality, and economic burden associated with BSI. For molecular-based methods, there is a lot to gain from an improved and straightforward method for isolation of bacteria from whole blood for downstream processing.
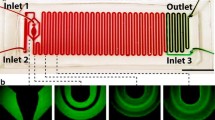
We describe a microfluidic-based sample-preparation approach that rapidly and selectively lyses all blood cells while it extracts intact bacteria for downstream analysis. Whole blood is exposed to a mild detergent, which lyses most blood cells, and then to osmotic shock using deionized water, which eliminates the remaining white blood cells. The recovered bacteria are 100 % viable, which opens up possibilities for performing drug susceptibility tests and for nucleic-acid-based molecular identification.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Levy MM, Artigas A, Phillips GS, Rhodes A, Beale R, Osborn T et al (2012) Outcomes of the surviving sepsis campaign in intensive care units in the USA and Europe: a prospective cohort study. Lancet Infect Dis 12(12):919–924
Kumar A, Roberts D, Wood KE, Light B, Parrillo JE, Sharma S et al (2006) Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit Care Med 34(6):1589–1596
Tissari P, Zumla A, Tarkka E, Mero S, Savolainen L, Vaara M et al (2010) Accurate and rapid identification of bacterial species from positive blood cultures with a DNA-based microarray platform: an observational study. Lancet 375(9710):224–230
Turenne CY, Witwicki E, Hoban DJ, Karlowsky JA, Kabani AM (2000) Rapid identification of bacteria from positive blood cultures by fluorescence-based PCR-single-strand conformation polymorphism analysis of the 16S rRNA gene. J Clin Microbiol 38(2):513–520
Wellinghausen N, Wirths B, Essig A, Wassill L (2004) Evaluation of the hyplex bloodscreen multiplex PCR-enzyme-linked immunosorbent assay system for direct identification of gram-positive cocci and gram-negative bacilli from positive blood cultures. J Clin Microbiol 42(7):3147–3152
McCann CD, Jordan JA (2014) Evaluation of MolYsisTM Complete5 DNA extraction method for detecting Staphylococcus aureus DNA from whole blood in a sepsis model using PCR/pyrosequencing. J Microbiol Methods 99:1–7
Vince A, Lepej SZ, Barsić B, Dusek D, Mitrović Z, Serventi-Seiwerth R et al (2008) LightCycler SeptiFast assay as a tool for the rapid diagnosis of sepsis in patients during antimicrobial therapy. J Med Microbiol 257(Pt 10):1306–1307
Tang Y-W, Kilic A, Yang Q, McAllister SK, Li H, Miller RS et al (2007) StaphPlex system for rapid and simultaneous identification of antibiotic resistance determinants and Panton-Valentine leukocidin detection of staphylococci from positive blood cultures. J Clin Microbiol 45(6):1867–1873
Jordan JA, Durso MB (2005) Real-time polymerase chain reaction for detecting bacterial DNA directly from blood of neonates being evaluated for sepsis. J Mol Diagn 7(5):575–581
Wellinghausen N, Kochem A-J, Disqué C, Mühl H, Gebert S, Winter J et al (2009) Diagnosis of bacteremia in whole-blood samples by use of a commercial universal 16S rRNA gene-based PCR and sequence analysis. J Clin Microbiol 47(9):2759–2765
Weller SA, Cox V, Essex-Lopresti A, Hartley MG, Parsons TM, Rachwal PA et al (2012) Evaluation of two multiplex real-time PCR screening capabilities for the detection of Bacillus anthracis, Francisella tularensis and Yersinia pestis in blood samples generated from murine infection models. J Med Microbiol 61(Pt 11):1546–1555
Kellogg JA, Manzella JP, Bankert DA (2000) Frequency of low-level bacteremia in children from birth to fifteen years of age. J Clin Microbiol 38(6):2181–2185
Henry NK, McLimans CA, Wright AJ, Thompson RL, Wilson WR, Washington JA (1983) Microbiological and clinical evaluation of the isolator lysis-centrifugation blood culture tube. J Clin Microbiol 17(5):864–869
Yagupsky P, Nolte FS (1990) Quantitative aspects of septicemia. Clin Microbiol Rev 3(3):269–279
Zelenin S, Hansson J, Ardabili S, Ramachandraiah H, Brismar H, Russom A (2015) Microfluidic-based isolation of bacteria from whole blood for sepsis diagnostics. Biotechnol Lett 37(4):825–830
Pichereau V, Hartke A, Auffray Y (2000) Starvation and osmotic stress induced multiresistances. Influence of extracellular compounds. Int J Food Microbiol 55(1–3):19–25
Corcoran BM, Stanton C, Fitzgerald G, Ross RP (2008) Life under stress: the probiotic stress response and how it may be manipulated. Curr Pharm Des 14(14):1382–1399
Ho SN (2006) Intracellular water homeostasis and the mammalian cellular osmotic stress response. J Cell Physiol 206(1):9–15
Stroock AD, Dertinger SKW, Ajdari A, Mezic I, Stone HA, Whitesides GM (2002) Chaotic mixer for microchannels. Science 295(5555):647–651
Acknowledgment
This work was supported by the European Commission (projects FP7 InTopSens and IMI RAPP-ID)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Zelenin, S., Ramachandraiah, H., Faridi, A., Russom, A. (2017). Microfluidic-Based Bacteria Isolation from Whole Blood for Diagnostics of Blood Stream Infection. In: Taly, V., Viovy, JL., Descroix, S. (eds) Microchip Diagnostics. Methods in Molecular Biology, vol 1547. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6734-6_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6734-6_14
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6732-2
Online ISBN: 978-1-4939-6734-6
eBook Packages: Springer Protocols