Abstract
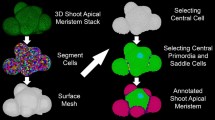
A comprehensive understanding of plant growth and development requires the integration of the spatial and temporal dynamics of gene regulatory networks with changes in cellular geometry during 3D organ growth. 3DCellAtlas is an integrative computational pipeline that semi-automatically identifies cell type and position within radially symmetric plant organs, and simultaneously quantifies 3D cell anisotropy and reporter abundance at single-cell resolution. It is a powerful tool that generates digital single-cell cellular atlases of plant organs and enables 3D cell geometry and reporter abundance (gene/protein/biosensor) from multiple samples to be integrated at single-cell resolution across whole organs. Here we describe how to use 3DCellAtlas to process and analyze radially symmetric organs, and to identify cell types and extract geometric cell data within these 3D cellular datasets. We detail how to use two statistical tools in 3DCellAtlas to compare cellular geometries, and to analyze reporter abundance at single-cell resolution.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Roeder AH et al (2011) Computational morphodynamics of plants: integrating development over space and time. Nat Rev Mol Cell Biol 12(4):265–273
Fernandez R et al (2010) Imaging plant growth in 4D: robust tissue reconstruction and lineaging at cell resolution. Nat Methods 7(7):547–553
Kierzkowski D et al (2012) Elastic domains regulate growth and organogenesis in the plant shoot apical meristem. Science 335(6072):1096–1099
Truernit E et al (2008) High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of Phloem development and structure in Arabidopsis. Plant Cell 20(6):1494–1503
Bassel GW et al (2014) Mechanical constraints imposed by 3D cellular geometry and arrangement modulate growth patterns in the Arabidopsis embryo. Proc Natl Acad Sci U S A 111(23):8685–8690
Yoshida S et al (2014) Genetic control of plant development by overriding a geometric division rule. Dev Cell 29(1):75–87
Barbier de Reuille P et al (2015) MorphoGraphX: a platform for quantifying morphogenesis in 4D. eLife 4:e05864
Shapiro BE et al (2015) Analysis of cell division patterns in the Arabidopsis shoot apical meristem. Proc Natl Acad Sci U S A 112(15):4815–4820
Montenegro-Johnson TD et al (2015) Digital single-cell analysis of plant organ development using 3DCellAtlas. Plant Cell 27(4):1018–1033
Bassel GW (2015) Accuracy in quantitative 3D image analysis. Plant Cell 27(4):950–953
Acknowledgments
G.W.B. was funded by Biotechnology and Biological Sciences Research Council (BBSRC) Grant BB/L010232/1 and a Birmingham Research Fellowship. P.S. was funded by BBSRC grant BB/J017604/1. T.D.M.J. is supported by a Royal Commission for the Exhibition of 1851 Fellowship. R.S.S. was funded by Swiss National Science Foundation Interdisciplinary Project Grant number CR32I3_143833 and the Max Planck Society.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Stamm, P., Strauss, S., Montenegro-Johnson, T.D., Smith, R., Bassel, G.W. (2017). In Silico Methods for Cell Annotation, Quantification of Gene Expression, and Cell Geometry at Single-Cell Resolution Using 3DCellAtlas. In: Kleine-Vehn, J., Sauer, M. (eds) Plant Hormones. Methods in Molecular Biology, vol 1497. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6469-7_11
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6469-7_11
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6467-3
Online ISBN: 978-1-4939-6469-7
eBook Packages: Springer Protocols