Abstract
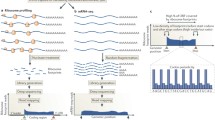
Ribosome profiling provides a genome-wide view of translation with unprecedented resolution. Application of this approach to fission and budding yeast revealed widespread regulation of translational efficiency, translation of short open reading frames on unannotated transcripts, and frequent translation of open reading frames in 5′ leader sequences. We present here a detailed protocol for the application of ribosome profiling to meiotic fission yeast cells, although the approach should be easily adapted to budding yeast.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Yanagida M (2002) The model unicellular eukaryote, Schizosaccharomyces pombe. Genome Biol 3(3):COMMENT2003
Harigaya Y, Yamamoto M (2007) Molecular mechanisms underlying the mitosis-meiosis decision. Chromosome Res 15(5):523–537
Mata J, Lyne R, Burns G, Bähler J (2002) The transcriptional program of meiosis and sporulation in fission yeast. Nat Genet 32(1):143–147
Mata J, Bähler J (2006) Global roles of Ste11p, cell type, and pheromone in the control of gene expression during early sexual differentiation in fission yeast. Proc Natl Acad Sci U S A 103(42):15517–15522
Mata J, Wilbrey A, Bähler J (2007) Transcriptional regulatory network for sexual differentiation in fission yeast. Genome Biol 8(10):R217
Amorim MJ, Cotobal C, Duncan C, Mata J (2010) Global coordination of transcriptional control and mRNA decay during cellular differentiation. Mol Syst Biol 6:380
Duncan CD, Mata J (2014) The translational landscape of fission-yeast meiosis and sporulation. Nat Struct Mol Biol 21(7):641–647. doi:10.1038/nsmb.2843
Harigaya Y, Tanaka H, Yamanaka S, Tanaka K, Watanabe Y, Tsutsumi C, Chikashige Y, Hiraoka Y, Yamashita A, Yamamoto M (2006) Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442(7098):45–50
Ingolia NT, Ghaemmaghami S, Newman JR, Weissman JS (2009) Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324(5924):218–223. doi:10.1126/science.1168978
Ingolia NT (2014) Ribosome profiling: new views of translation, from single codons to genome scale. Nat Rev Genet 15(3):205–213. doi:10.1038/nrg3645
Chew GL, Pauli A, Rinn JL, Regev A, Schier AF, Valen E (2013) Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140(13):2828–2834. doi:10.1242/dev.098343
Gerashchenko MV, Lobanov AV, Gladyshev VN (2012) Genome-wide ribosome profiling reveals complex translational regulation in response to oxidative stress. Proc Natl Acad Sci U S A 109(43):17394–17399. doi:10.1073/pnas.1120799109
Ingolia NT, Brar GA, Stern-Ginossar N, Harris MS, Talhouarne GJ, Jackson SE, Wills MR, Weissman JS (2014) Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep 8(5):1365–1379. doi:10.1016/j.celrep.2014.07.045
Brar GA, Yassour M, Friedman N, Regev A, Ingolia NT, Weissman JS (2012) High-resolution view of the yeast meiotic program revealed by ribosome profiling. Science 335(6068):552–557. doi:10.1126/science.1215110
Ingolia NT, Lareau LF, Weissman JS (2011) Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147(4):789–802. doi:10.1016/j.cell.2011.10.002
Konig J, Zarnack K, Rot G, Curk T, Kayikci M, Zupan B, Turner DJ, Luscombe NM, Ule J (2010) iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat Struct Mol Biol 17(7):909–915. doi:10.1038/nsmb.1838
Mata J (2013) Genome-wide mapping of polyadenylation sites in fission yeast reveals widespread alternative polyadenylation. RNA Biol 10(8):1407–1414. doi:10.4161/rna.25758
Iino Y, Yamamoto M (1985) Mutants of Schizosaccharomyces pombe which sporulate in the haploid state. Mol Gen Genet 198:416–421
Nurse P (1985) Mutants of the fission yeast Schizosacharomyces pombe which alter the shift between cell proliferation and sporulation. Mol Gen Genet 198:497
Moreno S, Klar A, Nurse P (1991) Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol 194:795–823
Cipak L, Zhang C, Kovacikova I, Rumpf C, Miadokova E, Shokat KM, Gregan J (2011) Generation of a set of conditional analog-sensitive alleles of essential protein kinases in the fission yeast Schizosaccharomyces pombe. Cell Cycle 10(20):3527–3532. doi:10.4161/cc.10.20.17792
Guerra-Moreno A, Alves-Rodrigues I, Hidalgo E, Ayte J (2012) Chemical genetic induction of meiosis in Schizosaccharomyces pombe. Cell Cycle 11(8):1621–1625. doi:10.4161/cc.20051
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1)
Wood V, Harris MA, McDowall MD, Rutherford K, Vaughan BW, Staines DM, Aslett M, Lock A, Bähler J, Kersey PJ, Oliver SG (2012) PomBase: a comprehensive online resource for fission yeast. Nucleic Acids Res 40(Database issue):D695–D699. doi:10.1093/nar/gkr853
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36. doi:10.1186/gb-2013-14-4-r36
Thorvaldsdottir H, Robinson JT, Mesirov JP (2013) Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14(2):178–192. doi:10.1093/bib/bbs017
Chung BY, Hardcastle TJ, Jones JD, Irigoyen N, Firth AE, Baulcombe DC, Brierley I (2015) The use of duplex-specific nuclease in ribosome profiling and a user-friendly software package for Ribo-seq data analysis. RNA 21(10):1731–1745. doi:10.1261/rna.052548.115
Gerashchenko MV, Gladyshev VN (2014) Translation inhibitors cause abnormalities in ribosome profiling experiments. Nucleic Acids Res 42(17), e134. doi:10.1093/nar/gku671
Liu B, Han Y, Qian SB (2013) Cotranslational response to proteotoxic stress by elongation pausing of ribosomes. Mol Cell 49(3):453–463. doi:10.1016/j.molcel.2012.12.001
Esposito AM, Mateyak M, He D, Lewis M, Sasikumar AN, Hutton J, Copeland PR, Kinzy TG (2010) Eukaryotic polyribosome profile analysis. J Vis Exp 40:pii:1948. doi:10.3791/1948
Ingolia NT, Brar GA, Rouskin S, McGeachy AM, Weissman JS (2012) The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat Protoc 7(8):1534–1550. doi:10.1038/nprot.2012.086
Acknowledgement
This was supported by a Biotechnology and Biological Sciences Research Council (BBSRC) project grant to Juan Mata (BB/M021483/1). We thank Maria J. Amorim for help with sucrose gradient centrifugation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Duncan, C., Mata, J. (2017). Ribosome Profiling for the Analysis of Translation During Yeast Meiosis. In: Stuart, D. (eds) Meiosis. Methods in Molecular Biology, vol 1471. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6340-9_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6340-9_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6338-6
Online ISBN: 978-1-4939-6340-9
eBook Packages: Springer Protocols