Abstract
Sensory photoreceptors underpin optogenetics by mediating the noninvasive and reversible perturbation of living cells by light with unprecedented temporal and spatial resolution. Spurred by seminal optogenetic applications of natural photoreceptors, the engineering of photoreceptors has recently garnered wide interest and has led to the construction of a broad palette of novel light-regulated actuators. Photoreceptors are modularly built of photosensors that receive light signals, and of effectors that carry out specific cellular functions. These modules have to be precisely connected to allow efficient communication, such that light stimuli are relayed from photosensor to effector. The engineering of photoreceptors benefits from a thorough understanding of the underlying signaling mechanisms. This chapter gives a brief overview of key characteristics and signal-transduction mechanisms of sensory photoreceptors. Adaptation of these concepts in photoreceptor engineering has enabled the generation of novel optogenetic tools that greatly transcend the repertoire of natural photoreceptors.
*The first two authors contributed euqally to this chapter.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
As genetically encoded, light-re gulated actuators, photoreceptors provide the basis for optogenetics, the noninvasive, reversible, and spatiotemporally precise manipulation of biological processes by light. In signal transduction as in biology in general, researchers often tackle complex natural systems by disassembling them into simpler building blocks with more tractable attributes. For signal receptors such building blocks commonly correspond to sensor modules that receive environmental stimuli as input signals and effector modules that exert specific cellular functions in response to a given stimulus. These modules distribute into several classes with recurring structural and functional motifs as well as common principles of signal transduction. The modular nature of signaling proteins often allows the recombination of sensor and effector modules to accommodate new input or output modalities, or to vary functional parameters (e.g., light sensitivity, response kinetics , or dynamic range) of the composite sensor-effector system. In this chapter, we focus on the engineering of photoreceptors for which sensor and effector are organized in distinct protein domains or proteins [1–3]; by contrast, we only brush upon receptors in which these modules are integrated into a single domain, as for example within the large group of microbial rhodopsins acting as light-gated ion channels and pumps that kick-started optogenetics, reviewed elsewhere [4–6].
Intact signal transmission between photosensor and effector modules depends on diverse and dynamic allosteric coupling mechanisms. In many rational photoreceptor engineering approaches fundamental information on the structural and functional attributes of these modules is a prerequisite for the generation of functional sensor-effector combinations. To obtain this information, photoreceptors are often decomposed into isolated modules with a reduced number of parameters, and we organize this chapter in a corresponding manner by first introducing characteristic attributes of candidate photosensor (Subheading 2) and effector modules (Subheading 3), before c ontinuing with the mechanistic principles of signal transduction (Subheading 4) that motivate the choice of the eventual design strategy (Subheading 5).
2 The Photosensor
To receive the environmental st imulus light, all photosensors harbor an organic chromophore (Subheading 2.1) with a conjugated π electron system that absorbs photons in the UV/visible range of the electromagnetic spectrum and transmits part of the absorbed energy to the protein scaffold [7, 8]. Light absorption by the dark-adapted state D initiates a so-called photocycle (Subheading 2.2), eventually leading to population of the signaling state S. This state then persists from milliseconds to many hours depending upon photosensor before it reverts to D in a thermally driven, spontaneous reaction, denoted “dark recovery”. The kinetics for the reversion to D (Subheading 2.3) significantly affect the temporal resolution of optogenetic applications (off kinetics) and might effectively limit their reversibility on biologically relevant timescales.
2.1 Chromophore
The chromophore and the surrounding photosensor scaffold determine spectral sensitivity and photochemistry, based on which photoreceptors divide into several classes (Fig. 1) [7, 8]. The chromophore is embedded in the photosensor module, which mostly consists of a single protein domain but in case of phytochrome red-light sensors comprises three separate domains, denoted “photosensory core” (PSC) [9]. In view of eventual optogenetic applications, the choice of photosensor should be guided by at least two important considerations: chromophore availability in the target tissue; and wavelength used for stimulation.
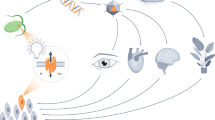
Properties of selected chromophores and sensory photoreceptors. The penetration depth of light in mammalian tissue increases strongly with increasing wavelengths with a maximum in the near infrared denoted as “near infrared window”. Blue-light se nsitive cryptochromes, BLUF, and LOV flavoproteins incorporate flavin nucleotides, and rhodopsins use retinal as chromophore, all of which are naturally present in most mammalian tissues. Plant and cyanobacterial phytochromes (Phys) as well as cyanobacteriochromes (CBCRs) require exogenous supply of their cofactor phycocyanobilin or phytochromobilin for optogenetic applications in vivo, since these reduced bilin derivatives cannot be provided by most cells. In contrast, bacterial phytochromes (BPhys) utilize the oxidized linear tetrapyrrole biliverdin, which is a direct heme degradation product, meaning exogenous chromophore addition is not necessary in most tissues tested to date. Photoreceptors are often built modularly, consisting of at least a sensor and an effector module. Exemplary natural photoreceptors with modular architecture are the LOV protein YtvA from Bac illus subtilis [61], a rhodops in guanylate cyclase from Blastocladiella emersonii [62], the plant phytochrome Arabidopsis thaliana PhyA [9], the Synechocystis sp. cyanobacteriochrome PixJ [63] and a bacterial phytochrome from Deinococcus ra diodurans [64]
First, the chromophore must be available in sufficient amounts at the target site in situ to be autonomously incorporated into the functional photoreceptor. Plant UV-B receptors [10] employ intrinsic amino acids to absorb light but more commonly, photoreceptors use chromophore cofactors that derive from small metabolites. Specifically, L OV (light-oxygen-voltage), BL UF (blue-light sensors using flavin-adenine dinucleotide), and cryptochrome sensors employ flavin-nucleotide chromophores sensitive to blue light [11–13]; the rhodopsin family use retinal to respond to light from the UV to the red [4]; phytochromes use linear tetrapyrroles (bilins) to respond to red and near-infrared wavelengths [9], further extended to the entire visible spectrum in recently discovered algal phytochromes [14]; and cyanobacteriochromes also use linear tetrapyrroles and exhibit spectral sensiti vity ranging from the UV to the near-infrared [15, 16]. Reduced tetrapyrroles, such as phycocyanobilin , that plant phytochromes and cyanobacteriochromes resort to, are not found in mammalian tissues which are frequent subjects of optogenetics. By contrast, the oxidized tetrapyrrole biliverdin, employed by bacterial phytochromes, retinal and flavin -nucleotide chromophores are apparently present in sufficient quantities in many mammalian tissues investigated to date [17–20].
Second, the wavelength used for photoreceptor activation determines the maximally achievable tissue penetration depth, phototoxicity , and potential combination of several optogenetic actuators and reporters. Limited tissue penetration of light complicates photon delivery to target sites within opaque tissues or deeper tissue layers. In particular, within the spectral region below 700 nm, penetration is substantially impeded by light scattering and absorption by lipids, hemoglobin, and other pigments. Mainly for longer wavelengths above ~700 nm, in a region denoted “near-infrared spectral window,” so far only covered by members of the phytochrome family, high penetration depths are achieved. Especially at lower wavelengths, the absorbed light quanta can elicit inadvertent phototoxic effects, e.g., due to generation of reactive oxygen species. If photoreceptors are to be deployed in parallel and/or in combination with fluorescent reporters, the individual wavelengths used for photoreceptor activation should be spectrally separated such that activation of a selected process does not interfere with other ones; that is, stimulation of a given photoreceptor should be orthogonal without eliciting other responses.
2.2 Photocycle
The term photocycle refers to a series of structural and dynamic changes within the chromophore and the surrounding protein scaffold following light absorption. In addition to the dark-adapted state D and the signaling state S, the photocycle often encompasses short-lived intermediate states. Regardless of the presence of these intermediates, the photochemical reaction towards the signaling state S is generally completed within microseconds at most, which is much faster than many physiological responses; for the purpose of this guideline we hence disregard ph otocycle intermediates.
The absolute light sensitivity of a photoreceptor depends on the absorption coefficient at a given wavelength and on the intrinsic quantum efficiency for formation of the signaling state. Notably, natural photoreceptors are intrinsically optimized for sensitive light reception with suitably high quantum efficiencies, and absolute ligh t sensitivity can usually not be enhanced to significant extent. Instead, to improve photoreceptor activation in optogenetic applications, light power can be ramped up but only to limited extent lest it causes severe biological damage. However, for optogenetic experiments conducted under constant illumination, a second route to optimizing photoreceptor activation is available. At photostationary conditions, an equilibrium is assumed between the dark-adapted and light-adapted states, which is not only determined by the kinetics of the light-driven forward reaction towards S but also by those of the thermally driven reverse reaction towards D (cf. Subheading 2.3). We denote the ratio of forward and reverse kinetics as the effective light sensitivity. For some photosensors , specifically LOV proteins and phytochromes, the dark recovery kinetics can be varied by many orders of magnitude via the introduction of mutations proximal to the chromophore , the reby offering an alternative way of modulating the effective light sensitivity [3].
2.3 Dark-Reversion Kinetics
The reversion from S to D occurs in a thermally driven reaction which can often be greatly accelerated by elevating temperature or changing solvent composition [21]. In addition to this spontaneous reaction, an alternative means of depleting S is offered in photochromic photoreceptors for which the signaling state S can actively be reverted to the dark-adapted state D by a subsequent light stimulus, typically of different color. The group of photochromic photoreceptors comprises phytochromes, cyanobacteriochromes , certain so-called “bistable” rhodopsins, and a re-engineered derivative of the photo-switchable fluorescent protein Dronpa [22]. The light-driven, bidirectional interconversion between D and S allows the regulation of downstream signaling events with superior temporal precision. Likewise, if activating and deactivating wavelengths are interleaved in space rather than time, superior spatial resolution can be obtained [3].
3 The Effector
The selection of a suitable effector module for photoreceptor engineering is largely determined by the desired output that should become subject to light control. The nature of the parental protein, from which the effector derives, governs a number of aspects that we discuss in turn: activity and dynamic range (Subheading 3.1); and availability of efficient activity assays (Subheading 3.2).
3.1 Activity and Dynamic Range
To elicit a suitable response in vivo, effector activity often has to be above certain threshold levels. Accordingly, activity of the photoreceptor in situ may have to be adjusted to match these levels, for example by varying overall expression levels of the photoreceptor and/or the specific activity of the effector. Another key consideration is the factor difference between the activities of an effector module in its low-activity and high-activity states, denoted as the “dynamic range” of the signal receptor. Notably, high dynamic ranges can only be achieved if the basal activity of the low-activity state is sufficiently low; for example, in light-activated receptors the dynamic range is often limited by residual dark activity. For engineered photoreceptors, a low dynamic range of the originally light-inert parental effector often limits the maximally attainable factor of light induction or repression. Vice versa, it is not guaranteed, that photoreceptors engineered on the basis of high-dynamic-range parental proteins will also yield strongly light-regulated derivatives. For example, the overall activity, the substrate affinities, and the maximal two-fold activation by light of E. coli di hydrofolate reductase (DHFR) fused to the Avena sativa phototropin 1 LOV2 (AsLOV2) pale in comparison to the corresponding parameters of wild-type DHFR [23].
In at least certain cases, the dynamic range can be amplified via downstream cellular signaling pathways , e.g., those involving gene expression [24], second messengers [25] or signaling cascades like MAP kinase pathways [26].
3.2 Activity Assay
The engineering of photoreceptors often requires the testing of sizeable numbers of candidate constructs which is greatly aided by the availability of fast and convenient activity assays. In general, high-throughput approaches distribute into two groups: screening systems, often set up inside living cells, which rely on readily detectable reporter readout (e.g., fluorescence); and selection systems in which cell proliferation/survival under set selection settings (e.g., dark vs. light) is conditional on expression and activity of candidate photoreceptors.
An efficient in vivo screening setup can be established provided that the desired effector output is orthogonal to other cellular metabolic pathways; does not harm living cells; and generates a chromogenic, fluorogenic, or other easily detectable readout. High-throughput screening systems are particularly effective if the output of the engineered photoreceptor can be coupled to reporter gene expression [24], thus allowing the screening of large numbers of receptor variants, for example by fluorescence-activated cell sorting. In case of proteins that undergo light-regulated association reactions, several display techniques, i.e. phage, mRNA , or ribosome display, are well suited for screening [27]. Independently of the screening approach, iterative rounds of positive and negative screening under light and dark conditions are often necessary to optimize dynamic range. If high-throughput screening systems cannot be established, photoreceptor engineering can be facilitated by medium-throughput screening systems, e.g., assays that determine the presence of specific metabolites or enzymatic activities in crude or partially purified cell lysates (see Chapter 7 for a recent example).
Selection systems, allowing cell growth under either light or dark conditions, and conversely leading to cell death or growth arrest under the opposite condition, provide an alternative means of accelerating photoreceptor engineering [28]. However, such systems need to be carefully set up and calibrated which is often challenging, in particular when the initial activity difference between dark and light conditions is small [29].
4 Allosteric Mechanisms of Signal Transduction
The transduction of signals in receptors is achieved through allosteric coupling between sensor and effector modules. Regardless of the precise mechanism, in photoreceptors the reception of light generally leads to initial conformational and dynamic transitions within the chromophore -binding pocket and the surrounding photosensor scaffold. Signal transmission to the effector is often achieved through α-helical structures that serve as linkers between photosensor and effector modules. Allosteric coupling mechanisms widely differ across photoreceptors but usually involve conformational and dynamic transitions, such as local unfolding, refolding, domain rearrangement, association, or dissociation [3]. We arrange this section based on whether light absorption causes changes in oligomeric state of the photoreceptor (Associating photoreceptors; Subheading 4.1) or not (Non-associating photoreceptors; Subheadings 4.2.1 and 4.2.2) (Fig. 2).
Allosteric mechanisms of signal transduction in natural photoreceptors and representative strategies derived for photoreceptor engineering. Engineered photoreceptors undergoing light-induced changes in oligomeric state can act as photo-activatable dimerization modules, for example to mediate the assembly of functional oligomeric states or the reconstitution of split proteins. Light-directed local unfolding can be utilized to release steric hindrance with concomitant changes in effector oligomeric state and/or activity. Engineering by domain exchange allows other light-induced tertiary and quaternary transitions to control naturally light-insensitive proteins
4.1 Associating Photoreceptors
For this group of photorecept ors, the reception of light results in association/dissociation reactions, mostly dimerization, mediated by the uncovering or covering of interaction sites. We distinguish between homo- and hetero-oligomerization depending on whether association occurs between alike or different partners. Association processes can be tied to changes in biological activity in different manners, for example by assembly of proteins into their functional oligomeric form; by colocalization of interacting proteins; or by recruitment of proteins to subcellular compartments.
Examples of naturally occurring systems that homo-oligomerize upon light absorption are the blue light-sensing Arabidopsis thaliana cryptochrome 2 (AtCry2) [30] and the L OV photoreceptors Vivid from Neurospora crassa [31], aureochrome from stramenopiles [32], and EL222 from Erythrobacter litoralis [33]. By contrast, the homodimeric photoreceptor AtUVR8 from A. thaliana dissociates into monomers u pon UV-light exposure [34]. In case of hetero-associating systems, the most widely deployed representatives derive from higher plants, exemplified by A. thaliana: AtCry2 not only assembles into homo-oligomers upon light absorption but also forms a heterodimer with its interacting partner AtCIB1 [35]; similarly, upon light-induced dissociation, AtUVR8 forms a heterodimer with AtCOP1 [34]. The LOV protein AtFKF1 interacts with its partner AtGIGANTEA following blu e-light absorption [36]; and the red/far-red sensing phytoc hromes A and B (AtPhyA and AtPhyB) associate in light-dependent manner with their interacting partners, of which AtPIF3 and AtPIF6 are the most popular in photoreceptor engineering (PIF, ph ytochrome interacting partner) [37, 38]. As we discuss in Subheading 5.1, light-regulated association/dissociation reactions have been utilized in numerous photoreceptor engineering studies.
4.2 Non-associating Photoreceptors
This category comprises a diverse group of photoreceptors for which signal transdu ction involves changes o f tertiary and, in case of oligomeric receptors, quaternary structure but no change in oligomeric state . In contrast to the above cases, for non-associating photoreceptors the physical nature of the linker (sequence, length, structure, topology, dynamics) between photosensor and effector modules is of much greater importance, as the linker has to specifically interact with both photosensor and effector sites to enable signal propagation. Put another way, photosensor and effector have to be linked in a manner conducive to efficient thermodynamic coupling between the se modules.
4.2.1 Uncaging of Peptide Epitopes/Active Sites
As a paradigm of this class, AsLOV2 exhibits light-triggered, local unfolding of its C-terminal Jα helix and concomitant dissociation from the LOV prot ein core [39]. In its original biological context within the multidomain receptor phototropin 1, Jα unfolding elicits subdomain rearrangements, but no apparent changes in the oligomeric state of the photoreceptor [40]. By contrast, in certain engineered photoreceptors, AsLOV2 has been converted into an associating photoreceptor (cf. Subheading 5). Notably, light-regulated unfolding mechanisms are not restricted to AsLOV2 but also contribute to signal transduction in other photoreceptors such as certain LOV domains (e.g., LOV2 from A. thalian a phototropin 1 [41], aureochrome 1a from Phaeodactylum tricornutum [42], RsLOV from Rhodobacter sphaeroides [43]) and the photoactive yellow protein (PYP) from purple bacteria [44].
4.2.2 Tertiary and Quaternary Transitions
In this section, we treat a disparate class of photoreceptors in which signal transmission primarily depends neither on changes in oligomeric state nor on local unfolding but rather on other tertiary or quaternary structural transitions that are often transmitted between sensor and effector modules by helical elements. Many of these concepts are exemplified in two recent case studies.
First, the recent crystal structure of the monomeric LOV histidine kinase EL346 from Erythrobacter litoralis HTCC2594 reveals long helices as mediators of photosensor -effector interdomain interactions [45]. These helices form an interface with the photosensory LOV c ore and maintain contact to the CA effector domain on the opposite side. The interdomain interactions stabilize the inhibited kinase form in the dark and weaken step-wise upon light induction, ther eby causing a rearrangement of the CA domain that increases its catalytic activity.
Second, Takala et al. recently presented crystal and solution structures of dark- and light-adapted states of the PSC module of Deinococcus radiodurans bacterial phytochrome (cf. Fig. 1) [46]. The structures of this parallel homodimeric protein support a previously proposed toggle-model for photoconversion in phytochromes [47]. According to this model, signal-induced rotation of the D ring of the tetrapyrrole chromophore causes contact rearrangements of the GAF/PHY interface. These rearrangements are possibly transferred to the C-terminal effector module by causing a tug on the linker helix and a concomitant pivot motion of the effector modules.
5 Design Strategies
Having discussed the properties of natural photoreceptors, we regard how their signaling mechanisms have been co-opted (in some cases, even transcending the natural mechanism) in the engineering of novel photoreceptors [2, 3] (Table 1).
5.1 Association/Dissociation
Our recent survey of photoreceptor engineering highlighted light-regulated association as a particularly versatile and promising design approach [3]. The prevalence of this approach is arguably explained by the frequent o ccurrence of oligomerization reactions in signal transduction and by less exigent demands on the linker connecting sensor and effector modules (cf. Subheading 4). Beyond providing a physical connection, requirements on the linker here are much less demanding, and linkers are often short, flexible, and predominantly hydrophilic. Association-based strategies are particularly well suited for effectors that are regulated by oligomerization reactions in their natural context, e.g., many transcription factors and transmembrane receptors. However, this approach is not restricted to naturally associating proteins but extends to proteins which are not originally regulated by oligomerization processes, in particular to split proteins.
For example, several recent studies described light-regulated variants of receptor tyr osine kinases in which activation is often based on ligand-induced receptor dimerization (RTKs) [26, 48, 49]. In all studies, control by ligand binding has been reprogrammed to control by light via fusion of the RTK to associating photoreceptors. Whereas Grusch et al. fused aureochrome LOV domains to the intracellular part of the membrane-bound RTK, the closely related studies by Chang et al. and Kim et al. accomplished the same via fusion to AtCry2.
Another set of studies employed associating photoreceptors to generate systems for light-induced expression of transgenes in eukaryotes [36, 50, 51]. For example, the Neurospora crassa Vivid LOV domain assembles into homodimers upon blue-light illumination; when linked to a truncated, nonfunctional monomeric form of the GAL4 transcription factor, this LOV d omain mediated light-dependent association and concomitant reconstitution of the functional dimeric form of GAL4. Even earlier, in a pioneering application, Shimizu-Sato et al. exploited the light-regulated association of the A. thalia na phy tochromes A and B with AtPIF3 to furnish a transcription system that can be activated by red and inactivated by far-r ed light [38]. The system is based on the yeast two-hybrid approach, with the GAL4 DNA-binding domain fused to either full-length or the N-terminal PSC of AtPhyA or AtPhyB, and AtPIF3 coupled to the GAL4 trans-activation domain. Conceptually similar, engineered photoreceptors are based on the A. thaliana cryptochrome AtCry2 and AtCIB1 and mediate light-regulated transgene expression and MAP-kinase signaling [35, 52].
5.2 Other Strategies
5.2.1 Local Unfolding
Local unfolding reactions can be harnessed to alter the accessibility of active sites and surface epitopes in a light-dependent manner, and to thereby regulate the activity of effector modules and downstream metabolic pathways. A striking demonstration of this approach is provided by photoactivatable Rac1 , a small GTPase involved in the regulation of cytoskeletal dynamics. Fusion to AsLOV2 led to steric restriction of the active site o f Rac1, which was relieved upon blue-light-induced unfolding of the AsLOV2 Jα helix, cf. Subheading 4.2.1, [53]. Another example for light-dependent control of activity is the inhibition of pot assium channels by peptide toxins, which were colocalized with the channels via fusion to membrane-tethered AsLOV2 [54]. Upon blue-light-induced unfolding of the Jα helix, the increased mobility of the toxin led to a decrease in its local concentration and channel opening. In two conceptually similar studies, systems for light-induced protein degradation were generated on the basis of phototropin LOV2 domains [41, 55]. Degron peptide sequences were interleaved with the LO V Jα helix such that they were little accessible under dark conditions; only upon light-induced Jα unfolding and concomitant exposure of the degron sequences, proteasomal degradation of target constructs was greatly stimulated.
5.2.2 Domain Exchange
In case the above two engineering strategies do not apply, originally light-inert signal receptors can often be subjected under light control if their sensor modules are replaced by suitable (homologous) photosensor modules. For example, GAF (cGMP-specific phosphodiesterases , adenylate cyclases , and FhlA) domains could be exchanged by (bacterial) phytochrome photosensors that comprise structurally homologous GAF and PHY (phytochrome specific) domains; or LOV domains could replace PAS do mains of which they are a subgroup. Often, the availability of three-dimensional structures allows the construction of structure-based alignments that guide the fusion between sensor and effector modules. When no suitable homologous relatives exist, domain exchange can still yield functional proteins but the lack of structure-based alignments complicates the planning of the fusion strategy. For exchange of sensor and effector modules linked by α-helical linkers (e.g., coiled coil linkers), an examination of the linker properties is helpful for the identificatio n of the best fusion site. Linker helices of discrete length widely recur in natural signal receptors [56–59] and crucially determine activity and regulation by light in engineered photoreceptors as the below case studies illustrate.
As an example for a successful domain exchange, the engineered red-light-sensitive photoreceptor Cph8 connects the PCB-binding photosensor of the cyanobacterial phytochrome Cph1 from Synechocystis sp. to the effector module of the histidine kinase EnvZ fro m E. coli [60]. The light-activated cAMP/cGMP-specific phosphodiesterase LAPD represents another example for homologous domain exchange [18], in this case of two GAF domains of the human phosphodiesterase 2A against a biliverdin-binding PAS-GAF-PHY tandem of the D. radiodurans bacterial phytochrome. Notably, in both cases use of the correct linker length was crucial for obtaining light-regulated enzymes. Finally, the light-activated adenylate cyclase IlaC is based on heterologous domain exchange; specifically, the PAS-GAF-PHY PSC module of the R. sphaeroides b acterial phytochrome BphG1 was connected to the adenylate cyclase effector from Nostoc sp. CyaB1 and thereby replaced two regulatory PAS domains [19].
6 Summary
As discussed above, the properties of engineered photoreceptors strongly depend on the intrinsic characteristics of the constituent photosensor and effector modules (Table 1) [3]. Therefore, the attributes of both the photosensor (e.g., genetic encodability; light sensitivity; achievable temporal and spatial resolution) and the effector (e.g., specific activity; possibility of amplification and availability of a screening assay) should be carefully considered at the initial design stages. Additionally, the resultant photoreceptor needs to be optimized regarding expression in situ, cell-type specific or subcellular targeting and dynamic range. Lastly, to engineer highly active and efficiently regulated photoreceptors, the signal transmission mechanisms of sensor and effector must be compatible.To date, mainly three fundamental design strategies have proven successful in the engineering of photoreceptors, and they apply to different scenarios: (a) Association-based approaches, implementable for effectors whose activity is a function of their oligomeric state or subcellular location ; (b) Approaches based on local unfolding that trigger uncaging of effector pepti des or release of steric hindrance; and (c) Exchange of homologous or heterologous sensor and effector modules. We expect optogenetics and photoreceptor engineering to continue their rapid development and to thus grant light control over otherwise light-insensitive processes that were previously inaccessible to optogenetic intervention.
References
Möglich A, Moffat K (2010) Engineered photoreceptors as novel optogenetic tools. Photochem Photobiol Sci 9:1286–1300
Schmidt D, Cho YK (2015) Natural photoreceptors and their application to synthetic biology. Trends Biotechnol 33:80–91
Ziegler T, Möglich A (2015) Photoreceptor engineering. Front Mol Biosci 2:30
Ernst OP, Lodowski DT, Elstner M, Hegemann P, Brown LS, Kandori H (2014) Microbial and animal rhodopsins: structures, functions, and molecular mechanisms. Chem Rev 114:126–163
Fenno L, Yizhar O, Deisseroth K (2011) The development and application of optogenetics. Annu Rev Neurosci 34:389–412
Schneider F, Grimm C, Hegemann P (2015) Biophysics of channelrhodopsin. Annu Rev Biophys 44:167–186
Hegemann P (2008) Algal sensory photoreceptors. Annu Rev Plant Biol 59:167–189
Möglich A, Yang X, Ayers RA, Moffat K (2010) Structure and function of plant photoreceptors. Annu Rev Plant Biol 61:21–47
Rockwell NC, Su Y-S, Lagarias JC (2006) Phytochrome structure and signaling mechanisms. Annu Rev Plant Biol 57:837–858
Brown BA, Cloix C, Jiang GH, Kaiserli E, Herzyk P, Kliebenstein DJ, Jenkins GI (2005) A UV-B-specific signaling component orchestrates plant UV protection. Proc Natl Acad Sci U S A 102:18225–18230
Conrad KS, Manahan CC, Crane BR (2014) Photochemistry of flavoprotein light sensors. Nat Chem Biol 10:801–809
Losi A, Mandalari C, Gärtner W (2015) The evolution and functional role of flavin-based prokaryotic photoreceptors. Photochem Photobiol 91:1021–1031. doi:10.1111/php.12489
Pudasaini A, El-Arab KK, Zoltowski BD (2015) LOV-based optogenetic devices: light-driven modules to impart photoregulated control of cellular signaling. Front Mol Biosci 2:18
Rockwell NC, Duanmu D, Martin SS, Bachy C, Price DC, Bhattacharya D, Worden AZ, Lagarias JC (2014) Eukaryotic algal phytochromes span the visible spectrum. Proc Natl Acad Sci U S A 111:3871–3876
Ikeuchi M, Ishizuka T (2008) Cyanobacteriochromes: a new superfamily of tetrapyrrole-binding photoreceptors in cyanobacteria. Photochem Photobiol Sci 7:1159–1167
Rockwell NC, Lagarias JC (2010) A brief history of phytochromes. Chemphyschem 11:1172–1180
Filonov GS, Piatkevich KD, Ting L-M, Zhang J, Kim K, Verkhusha VV (2011) Bright and stable near-infrared fluorescent protein for in vivo imaging. Nat Biotechnol 29:757–761
Gasser C, Taiber S, Yeh C-M, Wittig CH, Hegemann P, Ryu S, Wunder F, Möglich A (2014) Engineering of a red-light-activated human cAMP/cGMP-specific phosphodiesterase. Proc Natl Acad Sci U S A 111:8803–8808
Ryu M-H, Kang I-H, Nelson MD, Jensen TM, Lyuksyutova AI, Siltberg-Liberles J, Raizen DM, Gomelsky M (2014) Engineering adenylate cyclases regulated by near-infrared window light. Proc Natl Acad Sci U S A 111:10167–10172
Shu X, Royant A, Lin MZ, Aguilera TA, Lev-Ram V, Steinbach PA, Tsien RY (2009) Mammalian expression of infrared fluorescent proteins engineered from a bacterial phytochrome. Science 324:804–807
Alexandre MT, Arents JC, van Grondelle R, Hellingwerf KJ, Kennis JT (2007) A base-catalyzed mechanism for dark state recovery in the Avena sativa phototropin-1 LOV2 domain. Biochemistry 46:3129–3137
Zhou XX, Chung HK, Lam AJ, Lin MZ (2012) Optical control of protein activity by fluorescent protein domains. Science 338:810–814
Lee J, Natarajan M, Nashine VC, Socolich M, Vo T, Russ WP, Benkovic SJ, Ranganathan R (2008) Surface sites for engineering allosteric control in proteins. Science 322:438–442
Ohlendorf R, Vidavski RR, Eldar A, Moffat K, Möglich A (2012) From dusk till dawn: one-plasmid systems for light-regulated gene expression. J Mol Biol 416:534–542
Jansen V, Alvarez L, Balbach M, Strünker T, Hegemann P, Kaupp UB, Wachten D (2015) Controlling fertilization and cAMP signaling in sperm by optogenetics. eLife 4:e05161
Grusch M, Schelch K, Riedler R, Reichhart E, Differ C, Berger W, Inglés-Prieto Á, Janovjak H (2014) Spatio-temporally precise activation of engineered receptor tyrosine kinases by light. EMBO J 33:1713–1726
Guntas G, Hallett RA, Zimmerman SP, Williams T, Yumerefendi H, Bear JE, Kuhlman B (2015) Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc Natl Acad Sci U S A 112:112–117
Cosentino C, Alberio L, Gazzarrini S et al (2015) Optogenetics. Engineering of a light-gated potassium channel. Science 348:707–710
Goldsmith M, Tawfik DS (2012) Directed enzyme evolution: beyond the low-hanging fruit. Curr Opin Struct Biol 22:406–412
Bugaj LJ, Choksi AT, Mesuda CK, Kane RS, Schaffer DV (2013) Optogenetic protein clustering and signaling activation in mammalian cells. Nat Methods 10:249–252
Lamb JS, Zoltowski BD, Pabit SA, Crane BR, Pollack L (2008) Time-resolved dimerization of a PAS-LOV protein measured with photocoupled small angle X-ray scattering. J Am Chem Soc 130:12226–12227
Takahashi F, Yamagata D, Ishikawa M, Fukamatsu Y, Ogura Y, Kasahara M, Kiyosue T, Kikuyama M, Wada M, Kataoka H (2007) AUREOCHROME, a photoreceptor required for photomorphogenesis in stramenopiles. Proc Natl Acad Sci U S A 104:19625–19630
Nash AI, McNulty R, Shillito ME, Swartz TE, Bogomolni RA, Luecke H, Gardner KH (2011) Structural basis of photosensitivity in a bacterial light-oxygen-voltage/helix-turn-helix (LOV-HTH) DNA-binding protein. Proc Natl Acad Sci U S A 108:9449–9454
Christie JM, Arvai AS, Baxter KJ et al (2012) Plant UVR8 photoreceptor senses UV-B by tryptophan-mediated disruption of cross-dimer salt bridges. Science 335:1492–1496
Kennedy MJ, Hughes RM, Peteya LA, Schwartz JW, Ehlers MD, Tucker CL (2010) Rapid blue-light-mediated induction of protein interactions in living cells. Nat Methods 7:973–975
Yazawa M, Sadaghiani AM, Hsueh B, Dolmetsch RE (2009) Induction of protein-protein interactions in live cells using light. Nat Biotechnol 27:941–945
Levskaya A, Weiner OD, Lim WA, Voigt CA (2009) Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461:997–1001
Shimizu-Sato S, Huq E, Tepperman JM, Quail PH (2002) A light-switchable gene promoter system. Nat Biotechnol 20:1041–1044
Harper SM, Neil LC, Gardner KH (2003) Structural basis of a phototropin light switch. Science 301:1541–1544
Christie JM, Blackwood L, Petersen J, Sullivan S (2015) Plant flavoprotein photoreceptors. Plant Cell Physiol 56:401–413
Renicke C, Schuster D, Usherenko S, Essen L-O, Taxis C (2013) A LOV2 domain-based optogenetic tool to control protein degradation and cellular function. Chem Biol 20:619–626
Herman E, Kottke T (2015) Allosterically regulated unfolding of the A′α helix exposes the dimerization site of the blue-light-sensing aureochrome-LOV domain. Biochemistry 54:1484–1492
Conrad KS, Bilwes AM, Crane BR (2013) Light-induced subunit dissociation by a light-oxygen-voltage domain photoreceptor from Rhodobacter sphaeroides. Biochemistry 52:378–391
Rubinstenn G, Vuister GW, Mulder FAA, Düx PE, Boelens R, Hellingwerf KJ, Kaptein R (1998) Structural and dynamic changes of photoactive yellow protein during its photocycle in solution. Nat Struct Mol Biol 5:568–570
Rivera-Cancel G, Ko W, Tomchick DR, Correa F, Gardner KH (2014) Full-length structure of a monomeric histidine kinase reveals basis for sensory regulation. Proc Natl Acad Sci U S A 111:17839–17844
Takala H, Björling A, Berntsson O et al (2014) Signal amplification and transduction in phytochrome photosensors. Nature 509:245–248
Anders K, Gutt A, Gärtner W, Essen L-O (2014) Phototransformation of the red light sensor cyanobacterial phytochrome 2 from Synechocystis species depends on its tongue motifs. J Biol Chem 289:25590–25600
Chang K-Y, Woo D, Jung H et al (2014) Light-inducible receptor tyrosine kinases that regulate neurotrophin signalling. Nat Commun 5:4057
Kim N, Kim JM, Lee M, Kim CY, Chang K-Y, Heo WD (2014) Spatiotemporal control of fibroblast growth factor receptor signals by blue light. Chem Biol 21:903–912
Nihongaki Y, Suzuki H, Kawano F, Sato M (2014) Genetically engineered photoinducible homodimerization system with improved dimer-forming efficiency. ACS Chem Biol 9:617–621
Wang X, Chen X, Yang Y (2012) Spatiotemporal control of gene expression by a light-switchable transgene system. Nat Methods 9:266–269
Aoki K, Kumagai Y, Sakurai A, Komatsu N, Fujita Y, Shionyu C, Matsuda M (2013) Stochastic ERK activation induced by noise and cell-to-cell propagation regulates cell density-dependent proliferation. Mol Cell 52:529–540
Wu YI, Frey D, Lungu OI, Jaehrig A, Schlichting I, Kuhlman B, Hahn KM (2009) A genetically encoded photoactivatable Rac controls the motility of living cells. Nature 461:104–108
Schmidt D, Tillberg PW, Chen F, Boyden ES (2014) A fully genetically encoded protein architecture for optical control of peptide ligand concentration. Nat Commun 5:3019
Bonger KM, Rakhit R, Payumo AY, Chen JK, Wandless TJ (2014) General method for regulating protein stability with light. ACS Chem Biol 9:111–115
Anantharaman V, Balaji S, Aravind L (2006) The signaling helix: a common functional theme in diverse signaling proteins. Biol Direct 1:25
Möglich A, Ayers RA, Moffat K (2009) Design and signaling mechanism of light-regulated histidine kinases. J Mol Biol 385:1433–1444
Möglich A, Ayers RA, Moffat K (2010) Addition at the molecular level: signal integration in designed Per-ARNT-Sim receptor proteins. J Mol Biol 400:477–486
Rockwell NC, Ohlendorf R, Möglich A (2013) Cyanobacteriochromes in full color and three dimensions. Proc Natl Acad Sci U S A 110:806–807
Levskaya A, Chevalier AA, Tabor JJ et al (2005) Synthetic biology: sngineering Escherichia coli to see light. Nature 438:441–442
Losi A, Polverini E, Quest B, Gärtner W (2002) First evidence for phototropin-related blue-light receptors in prokaryotes. Biophys J 82:2627–2634
Avelar GM, Schumacher RI, Zaini PA, Leonard G, Richards TA, Gomes SL (2014) A rhodopsin-guanylyl cyclase gene fusion functions in visual perception in a fungus. Curr Biol 24:1234–1240
Yoshihara S, Suzuki F, Fujita H, Geng XX, Ikeuchi M (2000) Novel putative photoreceptor and regulatory genes required for the positive phototactic movement of the unicellular motile Cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 41:1299–1304
Davis SJ, Vener AV, Vierstra RD (1999) Bacteriophytochromes: phytochrome-like photoreceptors from nonphotosynthetic eubacteria. Science 286:2517–2520
Acknowledgements
Research in our laboratory is generously supported through a Sofja-Kovalevskaya Award by the Alexander-von-Humboldt Foundation (to A.M.) and by the Deutsche Forschungsgemeinschaft within the Cluster of Excellence ‘Unicat—Unifying Concepts in Catalysis’.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Ziegler, T., Schumacher, C.H., Möglich, A. (2016). Guidelines for Photoreceptor Engineering. In: Kianianmomeni, A. (eds) Optogenetics. Methods in Molecular Biology, vol 1408. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3512-3_27
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3512-3_27
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3510-9
Online ISBN: 978-1-4939-3512-3
eBook Packages: Springer Protocols