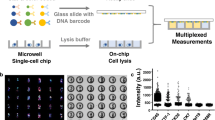
Abstract
The detection of protein translocation (i.e., the movement of intracellular proteins among various subcellular compartments) conventionally relies on imaging and subcellular-fractionation-based techniques that do not generate information on a large cell population with single-cell resolution. Although special flow cytometric tools such as imaging flow cytometry may generate single-cell data on processes such as nucleocytoplasmic transport, such equipment is expensive (thus has limited accessibility) and has low throughput for examining cells due to the reliance on high-speed imaging. Here we describe a protocol for detecting translocation of native proteins using a common flow cytometer which detects fluorescence intensity without imaging. We conduct chemical release of cytosolic proteins and fluorescence immunostaining of a targeted protein. The detected fluorescence intensity is quantitatively correlated to the cytosolic/nuclear localization of the protein at the single cell level. Our technique provides a simple route for studying nucleocytoplasmic transport with single-cell resolution using common flow cytometers.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pawson T (1995) Protein modules and signalling networks. Nature 373:573–580
Lemmon MA, Ferguson KM (2000) Signal-dependent membrane targeting by pleckstrin homology (PH) domains. Biochem J 350(Pt 1):1–18
Nakielny S, Dreyfuss G (1999) Transport of proteins and RNAs in and out of the nucleus. Cell 99:677–690
Nigg EA (1997) Nucleocytoplasmic transport: signals, mechanisms and regulation. Nature 386:779–787
Davis JR, Kakar M, Lim CS (2007) Controlling protein compartmentalization to overcome disease. Pharm Res 24:17–27
Kau TR, Way JC, Silver PA (2004) Nuclear transport and cancer: from mechanism to intervention. Nat Rev Cancer 4:106–117
Bellacosa A, Kumar C, Di Cristofano A, Testa J (2005) Activation of AKT kinases in cancer: implications for therapeutic targeting. Adv Cancer Res 94:29
Baldwin AS (2001) Control of oncogenesis and cancer therapy resistance by the transcription factor NF-kappaB. J Clin Invest 107:241–246
Kim HJ, Hawke N, Baldwin AS (2006) NF-kappa B and IKK as therapeutic targets in cancer. Cell Death Differ 13:738–747
Yamamoto Y, Gaynor RB (2001) Therapeutic potential of inhibition of the NF-kappaB pathway in the treatment of inflammation and cancer. J Clin Invest 107:135–142
Huber LA, Pfaller K, Vietor I (2003) Organelle proteomics – implications for subcellular fractionation in proteomics. Circ Res 92:962–968
Perez O, Nolan G (2006) Phospho-proteomic immune analysis by flow cytometry: from mechanism to translational medicine at the single-cell level. Immunol Rev 210:208–228
Krutzik P, Irish J, Nolan G, Perez O (2004) Analysis of protein phosphorylation and cellular signaling events by flow cytometry: techniques and clinical applications. Clin Immunol 110:206–221
Wang J et al (2008) Detection of kinase translocation using microfluidic electroporative flow cytometry. Anal Chem 80:1087–1093
Deptala A, Bedner E, Gorczyca W, Darzynkiewicz Z (1998) Activation of nuclear factor kappa B (NF-kappa B) assayed by laser scanning cytometry (LSC). Cytometry 33:376–382
Harnett MM (2007) Laser scanning cytometry: understanding the immune system in situ. Nat Rev Immunol 7:897–904
Arechiga AF et al (2005) Cutting edge: FADD is not required for antigen receptor-mediated NF-kappaB activation. J Immunol 175:7800–7804
Fanning SL et al (2006) Receptor cross-linking on human plasmacytoid dendritic cells leads to the regulation of IFN-alpha production. J Immunol 177:5829–5839
Basiji DA, Ortyn WE, Liang L, Venkatachalam V, Morrissey P (2007) Cellular image analysis and imaging by flow cytometry. Clin Lab Med 27:653–670
Vigo J, Salmon J, Lahmy S, Viallet P (1991) Fluorescent image cytometry: from qualitative to quantitative measurements. Anal Cell Pathol 3:145
Falkmer UG (1992) Methodologic sources of errors in image and flow cytometric DNA assessments of the malignancy potential of prostatic carcinoma. Hum Pathol 23:360–367
Wang J et al (2010) Embellishment of microfluidic devices via femtosecond laser micronanofabrication for chip functionalization. Lab Chip 10:1993–1996
Cao ZN, Geng S, Li LW, Lu C (2014) Detecting intracellular translocation of native proteins quantitatively at the single cell level. Chem Sci 5:2530–2535
Hoffmann A, Baltimore D (2006) Circuitry of nuclear factor κB signaling. Immunol Rev 210:171–186
Ghosh S, May MJ, Kopp EB (1998) NF-{kappa} B and Rel proteins: evolutionarily conserved mediators of immune responses. Sci Signal 16:225
Wassler M, Jonasson I, Persson R, Fries E (1987) Differential permeabilization of membranes by saponin treatment of isolated rat hepatocytes. Release of secretory proteins. Biochem J 247:407–415
Jamur MC, Oliver C (2010) Permeablization of cell membrances. Immunocytochemical methods and protocols. pp. 63–66
Goldenthal KL, Hedman K, Chen JW, August JT, Willingham MC (1985) Postfixation detergent treatment for immunofluorescence suppresses localization of some integral membrane proteins. J Histochem Cytochem 33:813–820
Negrutskii BS, Deutscher MP (1992) A sequestered pool of aminoacyl-tRNA in mammalian cells. Proc Natl Acad Sci U S A 89:3601–3604
Acknowledgments
This research was supported in part by NSF Grant 0967069.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Cao, Z., Lu, C. (2015). Quantitative Detection of Nucleocytoplasmic Transport of Native Proteins in Single Cells. In: Singh, A., Chandrasekaran, A. (eds) Single Cell Protein Analysis. Methods in Molecular Biology, vol 1346. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2987-0_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2987-0_16
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2986-3
Online ISBN: 978-1-4939-2987-0
eBook Packages: Springer Protocols