Abstract
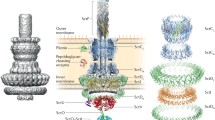
The outer membrane (OM) of gram-negative bacteria is highly packed with OM proteins (OMPs) and the trafficking and assembly of OMPs in gram-negative bacteria is a subject of intense research. Structurally, OMPs vary in the number of β-strands and in the size and complexity of extra-membrane domains, with extreme examples being the members of the type V protein secretion system (T5SS), such as the autotransporter (AT) and intimin/invasin families of secreted proteins, in which a large extracellular “passenger” domain is linked to a β-barrel that inserts in the OM. Despite their structural and functional diversity, OMPs interact in the periplasm with a relatively small set of protein chaperones that facilitate their transport from the inner membrane (IM) to the β-barrel assembly machinery (BAM complex), preventing aggregation and assisting their folding in various aspects including disulfide bond formation. This chapter is focused on the periplasmic folding factors involved in the biogenesis of integral OMPs and members of T5SS in E. coli, which are used as a model system in this field. Background information on these periplasmic folding factors is provided along with genetic methods to generate conditional mutants that deplete these factors from E. coli and biochemical methods to analyze the folding, surface display, disulfide formation and oligomerization state of OMPs/T5SS in these mutants.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Silhavy TJ, Kahne D, Walker S (2010) The bacterial cell envelope. Cold Spring Harb Perspect Biol 2(5):a000414
Hagan CL, Silhavy TJ, Kahne D (2011) beta-Barrel membrane protein assembly by the Bam complex. Annu Rev Biochem 80:189–210
van Ulsen P, Rahman SU, Jong WS et al (2013) Type V secretion: From biogenesis to biotechnology. Biochim et Biophys Acta 1843(8):1592–1611
Rigel NW, Silhavy TJ (2012) Making a beta-barrel: assembly of outer membrane proteins in Gram-negative bacteria. Curr Opin Microbiol 15(2):189–193
Du Plessis DJF, Nouwen N, Driessen AJM (2011) The Sec translocase. Biochimica et Biophysica Acta-Biomembranes 1808(3):851–865
Wu T, Malinverni J, Ruiz N et al (2005) Identification of a multicomponent complex required for outer membrane biogenesis in Escherichia coli. Cell 121(2):235–245
Knowles TJ, Scott-Tucker A, Overduin M et al (2009) Membrane protein architects: the role of the BAM complex in outer membrane protein assembly. Nat Rev Microbiol 7(3):206–214
Bardwell JC, McGovern K, Beckwith J (1991) Identification of a protein required for disulfide bond formation in vivo. Cell 67(3):581–589
Hayano T, Takahashi N, Kato S et al (1991) Two distinct forms of peptidylprolyl-cis-trans-isomerase are expressed separately in periplasmic and cytoplasmic compartments of Escherichia coli cells. Biochemistry 30(12):3041–3048
Missiakas D, Betton JM, Raina S (1996) New components of protein folding in extracytoplasmic compartments of Escherichia coli SurA, FkpA and Skp/OmpH. Mol Microbiol 21(4):871–884
Lazar SW, Kolter R (1996) SurA assists the folding of Escherichia coli outer membrane proteins. J Bacteriol 178(6):1770–1773
Chen R, Henning U (1996) A periplasmic protein (Skp) of Escherichia coli selectively binds a class of outer membrane proteins. Mol Microbiol 19(6):1287–1294
Merdanovic M, Clausen T, Kaiser M et al (2011) Protein quality control in the bacterial periplasm. Annu Rev Microbiol 65(65):149–168
Liechti G, Goldberg JB (2012) Outer membrane biogenesis in Escherichia coli, Neisseria meningitidis, and Helicobacter pylori: paradigm deviations in H. pylori. Front Cell Infect Microbiol 2:29
Geibel S, Procko E, Hultgren SJ et al (2013) Structural and energetic basis of folded-protein transport by the FimD usher. Nature 496(7444):243–246
Noinaj N, Kuszak AJ, Gumbart JC et al (2013) Structural insight into the biogenesis of beta-barrel membrane proteins. Nature 501(7467):385–390
Goemans C, Denoncin K, Collet JF (2013) Folding mechanisms of periplasmic proteins. Biochim Biophys Acta 1843(8):1517–1528
Bodelón G, Palomino C, Fernández LA (2013) Immunoglobulin domains in Escherichia coli and other enterobacteria: from pathogenesis to applications in antibody technologies. FEMS Microbiol Rev 37(2):204–250
Geibel S, Waksman G (2014) The molecular dissection of the chaperone-usher pathway. Biochim Biophys Acta 1843(8):1559–1567
Ruiz N, Silhavy TJ (2005) Sensing external stress: watchdogs of the Escherichia coli cell envelope. Curr Opin Microbiol 8(2):122–126
Bitto E, McKay DB (2002) Crystallographic structure of SurA, a molecular chaperone that facilitates folding of outer membrane porins. Structure 10(11):1489–1498
Behrens S, Maier R, de Cock H et al (2001) The SurA periplasmic PPlase lacking its parvulin domains functions in vivo and has chaperone activity. EMBO J 20(1-2):285–294
Xu XH, Wang SY, Hu YX et al (2007) The periplasmic bacterial molecular chaperone SurA adapts its structure to bind peptides in different conformations to assert a sequence preference for aromatic residues. J Mol Biol 373(2):367–381
Bitto E, McKay DB (2003) The periplasmic molecular chaperone protein SurA binds a peptide motif that is characteristic of integral outer membrane proteins. J Biol Chem 278(49):49316–49322
Ureta AR, Endres RG, Wingreen NS et al (2007) Kinetic analysis of the assembly of the outer membrane protein LamB in Escherichia coli mutants each lacking a secretion or targeting factor in a different cellular compartment. J Bacteriol 189(2):446–454
Ieva R, Bernstein HD (2009) Interaction of an autotransporter passenger domain with BamA during its translocation across the bacterial outer membrane. Proc Natl Acad Sci U S A 106(45):19120–19125
Bodelón G, Marín E, Fernández LA (2009) Role of Periplasmic Chaperones and BamA (YaeT/Omp85) in Folding and Secretion of Intimin from Enteropathogenic Escherichia coli Strains. J Bacteriol 191(16):5169–5179
Sklar JG, Wu T, Kahne D et al (2007) Defining the roles of the periplasmic chaperones SurA, Skp, and DegP in Escherichia coli. Genes Dev 21(19):2473–2484
Bennion D, Charlson ES, Coon E et al (2010) Dissection of beta-barrel outer membrane protein assembly pathways through characterizing BamA POTRA 1 mutants of Escherichia coli. Mol Microbiol 77(5):1153–1171
Heuck A, Schleiffer A, Clausen T (2011) Augmenting beta-augmentation: structural basis of how BamB Binds BamA and may support folding of outer membrane proteins. J Mol Biol 406(5):659–666
Vertommen D, Ruiz N, Leverrier P et al (2009) Characterization of the role of the Escherichia coli periplasmic chaperone SurA using differential proteomics. Proteomics 9(9):2432–2443
Hagan CL, Kim S, Kahne D (2010) Reconstitution of outer membrane protein assembly from purified components. Science 328(5980):890–892
Palomino C, Marín E, Fernández LA (2011) The Fimbrial Usher FimD Follows the SurA-BamB Pathway for Its Assembly in the Outer Membrane of Escherichia coli. J Bacteriol 193(19):5222–5230
Ricci DP, Schwalm J, Gonzales-Cope M et al (2013) The activity and specificity of the outer membrane protein chaperone SurA are modulated by a proline isomerase domain. MBio 4(4)
Walton TA, Sousa MC (2004) Crystal structure of Skp, a prefoldin-like chaperone that protects soluble and membrane proteins from aggregation. Mol Cell 15(3):367–374
Qu J, Mayer C, Behrens S et al (2007) The trimeric periplasmic chaperone skp of Escherichia coli forms 1: 1 complexes with outer membrane proteins via hydrophobic and electrostatic interactions. J Mol Biol 374(1):91–105
Burmann BM, Wang C, Hiller S (2013) Conformation and dynamics of the periplasmic membrane-protein–chaperone complexes OmpX–Skp and tOmpA–Skp. Nat Struct Mol Biol 20(11):1265–1272
Schafer U, Beck K, Muller M (1999) Skp, a molecular chaperone of Gram-negative bacteria, is required for the formation of soluble periplasmic intermediates of outer membrane proteins. J Biol Chem 274(35):24567–24574
Harms N, Koningstein G, Dontje W et al (2001) The early interaction of the outer membrane protein PhoE with the periplasmic chaperone Skp occurs at the cytoplasmic membrane. J Biol Chem 276(22):18804–18811
Ieva R, Tian P, Peterson JH et al (2011) Sequential and spatially restricted interactions of assembly factors with an autotransporter beta domain. Proc Natl Acad Sci U S A 108(31):E383–E391
Jarchow S, Luck C, Gorg A et al (2008) Identification of potential substrate proteins for the periplasmic Escherichia coli chaperone Skp. Proteomics 8(23-24):4987–4994
Ruiz-Perez F, Henderson IR, Leyton DL et al (2009) Roles of periplasmic chaperone proteins in the biogenesis of serine protease autotransporters of Enterobacteriaceae. J Bacteriol 191(21):6571–6583
Denoncin K, Schwalm J, Vertommen D et al (2012) Dissecting the Escherichia coli periplasmic chaperone network using differential proteomics. Proteomics 12(9):1391–1401
Volokhina EB, Grijpstra J, Stork M et al (2011) Role of the periplasmic chaperones Skp, SurA, and DegQ in outer membrane protein biogenesis in Neisseria meningitidis. J Bacteriol 193(7):1612–1621
Wagner JK, Heindl JE, Gray AN et al (2009) Contribution of the periplasmic chaperone Skp to efficient presentation of the autotransporter IcsA on the surface of Shigella flexneri. J Bacteriol 191(3):815–821
Spiess C, Beil A, Ehrmann M (1999) A temperature-dependent switch from chaperone to protease in a widely conserved heat shock protein. Cell 97(3):339–347
Krojer T, Garrido-Franco M, Huber R et al (2002) Crystal structure of DegP (HtrA) reveals a new protease-chaperone machine. Nature 416(6879):455–459
Krojer T, Sawa J, Schafer E et al (2008) Structural basis for the regulated protease and chaperone function of DegP. Nature 453(7197):885–890
Jiang JS, Zhang XF, Chen Y et al (2008) Activation of DegP chaperone-protease via formation of large cage-like oligomers upon binding to substrate proteins. Proc Natl Acad Sci U S A 105(33):11939–11944
Sawa J, Heuck A, Ehrmann M et al (2010) Molecular transformers in the cell: lessons learned from the DegP protease-chaperone. Curr Opin Struct Biol 20(2):253–258
Kim S, Grant RA, Sauer RT (2011) Covalent linkage of distinct substrate degrons controls assembly and disassembly of DegP proteolytic cages. Cell 145(1):67–78
Thompson NJ, Merdanovic M, Ehrmann M et al (2014) Substrate occupancy at the onset of oligomeric transitions of DegP. Structure 22(2):281–290
Kim S, Sauer RT (2012) Cage assembly of DegP protease is not required for substrate-dependent regulation of proteolytic activity or high-temperature cell survival. Proc Natl Acad Sci U S A 109(19):7263–7268
Ge X, Wang R, Ma J et al (2013) DegP primarily functions as a protease for the biogenesis of beta-barrel outer membrane proteins in the Gram-negative bacterium Escherichia coli. FEBS J 281:1226–1240
Rizzitello AE, Harper JR, Silhavy TJ (2001) Genetic evidence for parallel pathways of chaperone activity in the periplasm of Escherichia coli. J Bacteriol 183(23):6794–6800
Bos MP, Robert V, Tommassen J (2007) Biogenesis of the Gram-negative bacterial outer membrane. Annu Rev Microbiol 61:191–214
Schwalm J, Mahoney TF, Soltes GR et al (2013) A role for Skp in LptD assembly in Escherichia coli. J Bacteriol 195(16):3734–3742
Arie JP, Sassoon N, Betton JM (2001) Chaperone function of FkpA, a heat shock prolyl isomerase, in the periplasm of Escherichia coli. Mol Microbiol 39(1):199–210
Dartigalongue C, Missiakas D, Raina S (2001) Characterization of the Escherichia coli sigma(E) regulon. J Biol Chem 276(24):20866–20875
Saul FA, Arie JP, Vulliez-le NB et al (2004) Structural and functional studies of FkpA from Escherichia coli, a cis/trans peptidyl-prolyl isomerase with chaperone activity. J Mol Biol 335(2):595–608
Ruiz-Perez F, Henderson IR, Nataro JP (2010) Interaction of FkpA, a peptidyl-prolyl cis/trans isomerase with EspP autotransporter protein. Gut Microbes 1(5):339–344
Martin JL, Bardwell JCA, Kuriyan J (1993) Crystal structure of the DsbA protein required for disulfide bond formation in vivo. Nature 365(6445):464–468
Hiniker A, Bardwell JCA (2004) In vivo substrate specificity of periplasmic disulfide oxidoreductases. J Biol Chem 279(13):12967–12973
Kadokura H, Tian HP, Zander T et al (2004) Snapshots of DsbA in action: detection of proteins in the process of oxidative folding. Science (New York, NY) 303(5657):534–537
Negoda A, Negoda E, Reusch RN (2010) Resolving the native conformation of Escherichia coli OmpA. FEBS J 277(21):4427–4437
Brandon LD, Goldberg MB (2001) Periplasmic transit and disulfide bond formation of the autotransported Shigella protein IcsA. J Bacteriol 183(3):951–958
Chng SS, Xue M, Garner RA et al (2012) Disulfide rearrangement triggered by translocon assembly controls lipopolysaccharide export. Science (New York, NY) 337(6102):1665–1668
Kadokura H, Beckwith J (2009) Detecting folding intermediates of a protein as it passes through the bacterial translocation channel. Cell 138(6):1164–1173
Rietsch A, Belin D, Martin N et al (1996) An in vivo pathway for disulfide bond isomerization in Escherichia coli. Proc Natl Acad Sci U S A 93(23):13048–13053
McCarthy AA, Haebel PW, Torronen A et al (2000) Crystal structure of the protein disulfide bond isomerase, DsbC, from Escherichia coli. Nat Struct Biol 7(3):196–199
Missiakas D, Schwager F, Raina S (1995) Identification and characterization of a new disulfide isomerase-like protein (DsbD) in Escherichia coli. EMBO J 14(14):3415–3424
Heras B, Shouldice SR, Totsika M et al (2009) DSB proteins and bacterial pathogenicity. Nat Rev Microbiol 7(3):215–225
Totsika M, Heras B, Wurpel DJ et al (2009) Characterization of two homologous disulfide bond systems involved in virulence factor biogenesis in uropathogenic Escherichia coli CFT073. J Bacteriol 191(12):3901–3908
Arts IS, Ball G, Leverrier P et al (2013) Dissecting the machinery that introduces disulfide bonds in Pseudomonas aeruginosa. MBio 4(6):e00912–e00913
Reusch RN (2012) Insights into the structure and assembly of Escherichia coli outer membrane protein A. FEBS J 279(6):894–909
Ge X, Lyu ZX, Liu Y et al (2014) Identification of FkpA as a key quality control factor for the biogenesis of outer membrane proteins under heat shock conditions. J Bacteriol 196(3):672–680
Roux A, Beloin C, Ghigo JM (2005) Combined inactivation and expression strategy to study gene function under physiological conditions: Application to identification of new Escherichia coli adhesins. J Bacteriol 187(3):1001–1013
Veiga E, de Lorenzo V, Fernandez LA (1999) Probing secretion and translocation of a beta-autotransporter using a reporter single-chain Fv as a cognate passenger domain. Mol Microbiol 33(6):1232–1243
Veiga E, de Lorenzo V, Fernandez LA (2004) Structural tolerance of bacterial autotransporters for folded passenger protein domains. Mol Microbiol 52(4):1069–1080
Garmendia J, Phillips AD, Carlier MF et al (2004) TccP is an enterohaemorrhagic Escherichia coli O157: H7 type III effector protein that couples Tir to the actin-cytoskeleton. Cell Microbiol 6(12):1167–1183
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97(12):6640–6645
Cherepanov PP, Wackernagel W (1995) Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158(1):9–14
Donnenberg MS, Kaper JB (1991) Construction of an eae deletion mutant of Enteropathogenic Escherichia coli by using a positive selection suicide vector. Infect Immun 59(12):4310–4317
Frankel G, Phillips AD, Novakova M et al (1996) Intimin from enteropathogenic Escherichia coli restores murine virulence to a Citrobacter rodentium eaeA mutant: Induction of an immunoglobulin a response to intimin and EspB. Infect Immun 64(12):5315–5325
Marín E, Bodelón G, Fernández LA (2010) Comparative analysis of the biochemical and functional properties of C-terminal domains of autotransporters. J Bacteriol 192(21):5588–5602
Murphy KC (1998) Use of bacteriophage lambda recombination functions to promote gene replacement in Escherichia coli. J Bacteriol 180(8):2063–2071
Jurado P, Ritz D, Beckwith J et al (2002) Production of functional single-chain Fv antibodies in the cytoplasm of Escherichia coli. J Mol Biol 320(1):1–10
Nakamura K, Mizushima S (1976) Effects of heating in dodecyl sulfate solution on the conformation and electrophoretic mobility of isolated major outer membrane proteins from Escherichia coli K-12. J Biochem 80(6):1411–1422
Schweizer M, Hindennach I, Garten W et al (1978) Major proteins of the Escherichia coli outer cell envelope membrane. Interaction of protein II with lipopolysaccharide. Eur J Biochem 82(1):211–217
Fairman JW, Dautin N, Wojtowicz D et al (2012) Crystal structures of the outer membrane domain of intimin and invasin from enterohemorrhagic E. coli and enteropathogenic Y. pseudotuberculosis. Structure 20(7):1233–1243
Kenny B, DeVinney R, Stein M et al (1997) Enteropathogenic E. coli (EPEC) transfers its receptor for intimate adherence into mammalian cells. Cell 91(4):511–520
Frankel G, Candy DC, Everest P et al (1994) Characterization of the C-terminal domains of intimin-like proteins of enteropathogenic and enterohemorrhagic Escherichia coli, Citrobacter freundii, and Hafnia alvei. Infect Immun 62(5):1835–1842
Batchelor M, Prasannan S, Daniell S et al (2000) Structural basis for recognition of the translocated intimin receptor (Tir) by intimin from enteropathogenic Escherichia coli. EMBO J 19(11):2452–2464
Yu L, Frey EA, Pfuetzner RA et al (2000) Crystal structure of enteropathogenic Escherichia coli intimin-receptor complex. Nature 405(6790):1073–1077
Lee C. (2007) Western blotting. In: Rosato E (ed) Circadian Rhythms, vol 362. Methods in Molecular Biology, Humana Press, pp 391–399
Schagger H, von Jagow G (1991) Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal Biochem 199(2):223–231
Touze T, Hayward RD, Eswaran J et al (2004) Self-association of EPEC intimin mediated by the β-barrel containing anchor domain: a role in clustering of the Tir receptor. Mol Microbiol 51(1):73–87
Munera D, Hultgren S, Fernandez LA (2007) Recognition of the N-terminal lectin domain of FimH adhesin by the usher FimD is required for type 1 pilus biogenesis. Mol Microbiol 64(2):333–346
Acknowledgements
G.B. and E.M. contribute equally to this work. Work in the laboratory of LAF is supported by research grants from the Spanish “Ministerio de Economía y Competitividad” (MINECO) (BIO2011-26689), “Comunidad Autónoma de Madrid” (S2010-BMD-2312), “La Caixa” Foundation, and the European Research Council (ERC-2012-ADG_20120314).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Bodelón, G., Marín, E., Fernández, L.Á. (2015). Analyzing the Role of Periplasmic Folding Factors in the Biogenesis of OMPs and Members of the Type V Secretion System. In: Buchanan, S., Noinaj, N. (eds) The BAM Complex. Methods in Molecular Biology, vol 1329. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2871-2_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2871-2_7
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2870-5
Online ISBN: 978-1-4939-2871-2
eBook Packages: Springer Protocols