Abstract
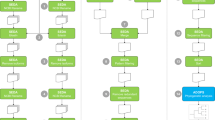
The bioinformatic annotation of Rab GTPases is important, for example, to understand the evolution of the endomembrane system. However, Rabs are particularly challenging for standard annotation pipelines because they are similar to other small GTPases and form a large family with many paralogous subfamilies. Here, we describe a bioinformatic annotation pipeline specifically tailored to Rab GTPases. It proceeds in two steps: first, Rabs are distinguished from other proteins based on GTPase-specific motifs, overall sequence similarity to other Rabs, and the occurrence of Rab-specific motifs. Second, Rabs are classified taking either a more accurate but slower phylogenetic approach or a slightly less accurate but much faster bioinformatic approach. All necessary steps can either be performed locally or using the referenced online tools. An implementation of a slightly more involved version of the pipeline presented here is available at RabDB.org.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
- 2.
- 3.
- 4.
- 5.
- 6.
- 7.
- 8.
- 9.
- 10.
- 11.
- 12.
- 13.
- 14.
- 15.
- 16.
- 17.
- 18.
- 19.
- 20.
- 21.
- 22.
- 23.
- 24.
- 25.
References
Diekmann Y, Seixas E, Gouw M et al (2011) Thousands of Rab GTPases for the cell biologist. PLoS Comput Biol 7:e1002217. doi:10.1371/journal.pcbi.1002217
Kloepper TH, Kienle N, Fasshauer D, Munro S (2012) Untangling the evolution of Rab G proteins: implications of a comprehensive genomic analysis. BMC Biol 10:71. doi:10.1186/1741-7007-10-71
Eliáš M, Brighouse A, Gabernet-Castello C et al (2012) Sculpting the endomembrane system in deep time: high resolution phylogenetics of Rab GTPases. J Cell Sci 125:2500–2508. doi:10.1242/jcs.101378
Pereira-Leal JB (2008) The Ypt/Rab family and the evolution of trafficking in fungi. Traffic 9:27–38. doi:10.1111/j.1600-0854.2007.00667.x
Shintani M, Tada M, Kobayashi T et al (2007) Characterization of Rab45/RASEF containing EF-hand domain and a coiled-coil motif as a self-associating GTPase. Biochem Biophys Res Commun 357:661–667. doi:10.1016/j.bbrc.2007.03.206
Leipe DD, Wolf YI, Koonin EV, Aravind L (2002) Classification and evolution of P-loop GTPases and related ATPases. J Mol Biol 317:41–72. doi:10.1006/jmbi.2001.5378
Pereira-Leal JB, Seabra MC (2000) The mammalian Rab family of small GTPases: definition of family and subfamily sequence motifs suggests a mechanism for functional specificity in the Ras superfamily. J Mol Biol 301:1077–1087. doi:10.1006/jmbi.2000.4010
de Lima Morais DA, Fang H, Rackham OJL et al (2011) SUPERFAMILY 1.75 including a domain-centric gene ontology method. Nucleic Acids Res 39:D427–D434. doi:10.1093/nar/gkq1130
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421. doi:10.1186/1471-2105-10-421
Bailey TL, Gribskov M (1998) Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14:48–54. doi:10.1093/bioinformatics/14.1.48
Bailey TL, Bodén M, Buske FA et al (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. doi:10.1093/nar/gkp335
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. doi:10.1093/molbev/mst010
Löytynoja A, Goldman N (2005) An algorithm for progressive multiple alignment of sequences with insertions. PNAS 102:10557–10562. doi:10.1073/pnas.0409137102
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490. doi:10.1371/journal.pone.0009490
Guindon S, Dufayard J-F, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. doi:10.1093/sysbio/syq010
Finn RD, Bateman A, Clements J et al (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222–D230. doi:10.1093/nar/gkt1223
Andreeva A, Howorth D, Chandonia J-M et al (2008) Data growth and its impact on the SCOP database: new developments. Nucleic Acids Res 36:D419–D425. doi:10.1093/nar/gkm993
Löytynoja A, Goldman N (2010) webprank: a phylogeny-aware multiple sequence aligner with interactive alignment browser. BMC Bioinformatics 11:579. doi:10.1186/1471-2105-11-579
Cock PJA, Antao T, Chang JT et al (2009) Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25:1422–1423. doi:10.1093/bioinformatics/btp163
Stajich JE, Block D, Boulez K et al (2002) The Bioperl toolkit: Perl modules for the life sciences. Genome Res 12:1611–1618. doi:10.1101/gr.361602
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763. doi:10.1093/bioinformatics/14.9.755
Gough J, Chothia C (2002) SUPERFAMILY: HMMs representing all proteins of known structure. SCOP sequence searches, alignments and genome assignments. Nucleic Acids Res 30:268–272. doi:10.1093/nar/30.1.268
Benson DA, Clark K, Karsch-Mizrachi I et al (2014) GenBank. Nucleic Acids Res 42:D32–D37. doi:10.1093/nar/gkt1030
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242. doi:10.1093/nar/28.1.235
UniProt Consortium (2014) Activities at the universal protein resource (UniProt). Nucleic Acids Res 42:D191–D198. doi:10.1093/nar/gkt1140
Wu CH, Yeh L-SL, Huang H et al (2003) The protein information resource. Nucleic Acids Res 31:345–347. doi:10.1093/nar/gkg040
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2:28–36
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152. doi:10.1146/annurev.genet.39.073003.112240
Moore I, Schell J, Palme K (1995) Subclass-specific sequence motifs identified in Rab GTPases. Trends Biochem Sci 20:10–12
Pfeffer SR (2005) Structural clues to Rab GTPase functional diversity. J Biol Chem 280:15485–15488. doi:10.1074/jbc.R500003200
Khan AR, Ménétrey J (2013) Structural biology of Arf and Rab GTPases’ effector recruitment and specificity. Structure 21:1284–1297. doi:10.1016/j.str.2013.06.016
Acknowledgements
We thank Mark Gouw for including the links to the sequence and motif files on the Rabifier website. This work was supported by a grant from Fundação para a Ciência e Tecnologia (PTDC/EBB-BIO/119006/2010)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Diekmann, Y., Pereira-Leal, J.B. (2015). Bioinformatic Approaches to Identifying and Classifying Rab Proteins. In: Li, G. (eds) Rab GTPases. Methods in Molecular Biology, vol 1298. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2569-8_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2569-8_2
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2568-1
Online ISBN: 978-1-4939-2569-8
eBook Packages: Springer Protocols