Abstract
Low temperature (LT) is a major threat that limits growth, development, and distribution leading to plant damage and crop losses. Plants respond to cold stress through a phenomenon known as cold acclimation, which is a complex process involving changes at multiple levels that include physiological and biochemical modifications, alterations in gene expression, and changes in concentrations of proteins and metabolites. Perception of cold stress by the cell membranes results in activation of cold-responsive genes and transcription factors that help in combating cold stress. Transcriptional responses to cold are guided by both ABA-dependent and -independent pathways that induce the expression of cold-regulated (COR) genes, thereby changing protein and metabolite homeostasis. Recent advances in the field of genomics, proteomics, and metabolomics has led to new discoveries, which has augmented our understanding of this intricate phenomenon. Here, we discuss the various aspects of cold stress responses in plants to develop a holistic understanding in the field of stress-mediated signaling.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Abiotic stresses adversely affect growth, productivity and survival of the plants, and hence are considered as key determinants of the crop losses worldwide. The human population is expected to touch 9.6 billion by 2050. To meet the food demand of this rapidly growing population, the agricultural production needs to be double by 2050. However, a continuous increase in the intensity, duration and unpredictability of abiotic stresses impose major limitation to achieve the required food production. Low temperature is one of the most intimidating abiotic stresses that affect the plant growth and development, thereby limiting the distribution of crop species. Based on its intensity, cold stress can be broadly classified into chilling and freezing stresses. Exposure to temperatures below 0 °C results in freezing stress, whereas chilling stress occurs at temperatures ranging from 0 to 20 °C. Plants such as rice, maize, and tomato that grow in tropical and subtropical regions are chilling sensitive whereas the plants from temperate region are chilling tolerant (Chinnusamy et al. 2007; Solanke and Sharma 2008). However, plants have the ability to acquire tolerance to chilling and freezing conditions if they are pre-exposed to non-freezing temperatures, a process known as cold acclimation (Levitt 1980). Cold acclimation helps plants to fine tune their metabolism and improve freezing tolerance by initiating signaling cascades that leads to several biochemical and physiological changes including modification of membrane lipid composition and changes in gene expression (Shinozaki and Yamaguchi-Shinozaki 1996; Thomashow 1998; Gilmour et al. 2000; Chinnusamy et al. 2003). The altered gene expression leads to accumulation of several protective proteins such as antifreeze proteins (AFPs) (Griffith et al. 1997), late embryogenesis abundant (LEA) proteins (Antikainen and Griffith 1997), heat shock proteins (HSP) (Wisniewski et al. 1996), cold-regulated (COR) proteins, and various metabolites such as amino acids, soluble sugars, organic acids, pigments (Krause et al. 1999), polyamines (Bouchereau et al. 1999), and antioxidants (Hausman et al. 2000). These metabolites and proteins help in protecting plant membranes and prevent cell disruption during cold stress by stabilizing the membrane lipids, proteins, maintaining the hydrophobic interactions, ion homeostasis and scavenging the reactive oxygen species (ROS) (Hare et al. 1998; Gusta et al. 2004; Chen and Murata 2008; Janska et al. 2010). In this chapter, we focus on the various aspects of cold acclimation such as physiological and biochemical effects of cold stress, cold sensing and signal transduction, cold-responsive pathway and role of various cold-responsive genes and transcription factors. We have discussed the progress in proteomic, metabolomics, and transcriptomic approaches that have been used to unravel the various biochemical pathways, cellular responses and complex gene interactions during low temperature stress in plants. These system approaches can give a deeper insight into the network of genes, proteins, and metabolites underlying the stress responses (Cramer et al. 2011).
2 Physiological Changes During Cold Stress
Plants respond variably to stress condition depending largely upon their phenological stages and the severity of cold they are subjected to. The various symptoms of chilling injury includes poor germination, stunted seedlings, surface lesions, water soaked appearance, desiccation, discoloration, tissue breakdown, and accelerated senescence among others (Sharma et al. 2005). Cold stress negatively influences reproductive development of plants resulting in flower sterility, as observed in rice at the time of anthesis (flower opening) (Jiang et al. 2002).
Freezing injury in most plants causes membrane damage due to severe cellular dehydration caused by ice formation in inter- and intracellular spaces. Since the chemical potential of ice is less than that of liquid water, extracellular ice formation leads to decrease in water potential outside the cell. Consequently, unfrozen water from inside the cell moves to the intercellular spaces resulting in cellular dehydration. This can potentially result in physical disruption of cells and tissues. Further, the magnitude of injury to the membrane also varies with the freezing temperature and the level of cellular dehydration (Steponkus et al. 1993). At temperatures between −2 to −4 °C, the major injury observed in non-acclimated plants is expansion-induced lysis caused by osmotic contraction and expansion. At lower temperatures of around −4 to −10 °C, the predominant form of injury in non-acclimated plants is freezing-induced lamellar-to-hexagonal-II phase transitions. Another form of injury is the fracture jump lesion that occurs at temperature below −10 °C between the plasma membrane and closely appressed cytoplasmic membrane (Webb and Steponkus 1993). Extreme cold also results in protein denaturation and solutes precipitation due to freezing-induced dehydration in plants (Uemura et al. 1995; Thomashow 1998; Salinas 2002). The lipid composition of plasma membrane is also altered significantly under cold stress (Steponkus et al. 1993; Uemura et al. 1995). Chilling stress also decreases the membrane fluidity by causing fatty acid unsaturation, altering lipid composition and ratio of lipids to proteins in cell membrane (Wang et al. 2006). An increase in the proportion of phospholipids during cold acclimation has been observed in various plant species (Uemura and Yoshida 1984; Uemura and Steponkus 1994; Ishikawa and Yoshida 1985; Uemura et al. 1995). In response to low temperature, the relative proportion of di-unsaturated phosphatidylcholine and phosphatidylethanolamine increases and that of mono-unsaturated phospholipids, cerebrosides and free sterols decreases in Arabidopsis (Uemura et al. 1995). Molecules of 16:0/16:0 phosphatidyl choline, phosphatidyl glycerol or cerebrosides undergo phase transition from a highly fluid liquid crystalline phase to the more rigid gel phase. The gel phase interferes with normal functioning of integral membrane protein in maintaining an efficient permeability barrier. These lead to ion leakage across the membrane and eventually cell dysfunction. Role of lipid unsaturation in cold tolerance was confirmed using transgenic tobacco plants expressing acyl-ACP:glycerol-3-phosphate acyl transferase (GPAT) from chilling-sensitive squash and chilling-tolerant Arabidopsis. Transgenic tobacco carrying GPAT from squash had higher levels of saturated phosphotidyl glycerol, while those with enzyme from Arabidopsis had decreased levels of saturated lipids (Murata et al. 1992). Transgenic tobacco expressing Arabidopsis ω-3 fatty acid desaturase (FAD7) resulted in higher levels of trienoic fatty acids and enhanced freezing tolerance (Kodama et al. 1994). Knockdown mutants of fad2, fad5, and fad6 in Arabidopsis resulted in growth defect and chlorotic plants at low temperature (Hugly and Somerville 1992; Miquel et al. 1993). Mutation in acyllipid desaturase leads to increased sensitivity to chilling and freezing temperatures in Arabidopsis (Chen and Thelen 2013). Cold stress causes inactivation of H+-ATPase in plasma membrane of chilling-sensitive plants (Yoshida 1991; Kasamo 1988). It is evident that an increase in membrane lipid unsaturation and bilayer fluidity are important adaptations for effective cold acclimation (Steponkus et al. 1993). The accumulation of sucrose, simple sugars, osmolytes, and LEA proteins during cold acclimation contributes to stabilization of membranes (Thomashow 1999).
3 Cold Sensing and Signaling
The plasma membrane appears to be the primary site for the perception of temperature change (Sangwan et al. 2002; Uemura et al. 2006; Vaultier et al. 2006; Wang et al. 2006), as low temperature can quickly induce membrane rigidification (Vaultier et al. 2006). Membrane rigidification is caused by the expression of a DMSO (dimethyl sulfoxide)-induced membrane rigidifier protein COR, even at normal growth temperatures, while benzyl alcohol, by negatively regulating the expression of COR gene at low temperatures confers membrane fluidity in alfalfa and Brassica napus (Orvar et al. 2000; Sangwan et al. 2001). Membrane rigidification probably leads to rearrangement of cytoskeletal microtubules and actin filaments, which then may activate mechanosensitive calcium channels to modulate Ca2+ signature (Nick 2000; Orvar et al. 2000; Sangwan et al. 2001).
A transient increase in cytosolic Ca2+ levels is observed as an early response to cold (Knight et al. 1991). Transgenics of Arabidopsis and tobacco expressing the calcium-sensitive luminescent protein aequorin demonstrated a rise in cytosolic Ca2+ concentration in response to low temperature (Knight et al. 1991). Cytosolic Ca2+ is an important second messenger in plant cells involved in cold signal transduction. A positive correlation was observed between cold-induced Ca2+ influx and accumulation of cold-induced transcripts in alfalfa (Monroy and Dhindsa 1995; Reddy and Reddy 2004) and Arabidopsis (Henriksson and Trewavas 2003). Use of various chelators and channel blockers also supported the role of cytosolic Ca2+ influx as second messenger in response to cold in alfalfa (Monroy et al. 1993; Knight et al. 1996) and Arabidopsis (Tahtiharju et al. 1997). The elevation in cytosolic Ca2+ is sensed by Ca2+ sensor proteins, which bind to Ca2+ and undergo conformational changes. The major Ca2+ sensors in plants are calmodulin (CaM), calcium-dependent protein kinases (CDPKs), calcineurin B-like proteins (CBLs), and CBL-interacting protein kinases (CIPKs) (Luan et al. 2002; Pandey 2008; Tuteja and Mahajan 2007). Different Ca2+ binding proteins distinguishes different Ca2+ signatures and protein kinases and decoding of these signals causes changes in gene expression leading to appropriate physiological responses (Yang and Poovaiah 2003; Sathyanarayanan and Poovaiah 2004; Pandey 2008).
CaM is one of the most conserved Ca2+ binding proteins in eukaryotes mediating various signaling responses against both developmental and environmental stimuli (Kim et al. 2009; Das and Pandey 2010). Studies with alfalfa cells (Monroy et al. 1993) and Arabidopsis (Tahtiharju et al. 1997) have shown that CaM antagonist prevents cold acclimation and reduces expression of cold-regulated genes, indicating a positive role of CaM in low temperature stress signaling. However, overexpression of CaM in Arabidopsis resulted in reduced expression of cold-responsive genes, implying it might also function as negative regulator of cold acclimation (Townley and Knight 2002).
Similarly, CDPKs are important sensors in abiotic stress signaling including cold stress (Cheng et al. 2002; Chinnusamy et al. 2004). Studies have shown that, in alfalfa cold induce the expression of CDPKs significantly (Monroy and Dhindsa 1995). Likewise, in rice, the expression of OsCPK4, OsCPK5, and OsCPK13 (OsCDPK7) was found to be upregulated in response to cold (Ray et al. 2007). The overexpression of OsCDPK7 in rice confers enhanced tolerance toward cold as well as salt and drought stresses (Saijo et al. 2001). Overexpression of OsCDPK13 and calreticulin interacting protein (CRTintP1) also conferred cold tolerance in rice (Komatsu et al. 2007).
CBLs were first identified in Arabidopsis. AtCBL1 was identified as a highly induced protein under cold and drought stress conditions (Kudla et al. 1999). Overexpression of AtCBL1 causes freezing sensitivity and alteration of expression of CBF/DREB transcription factor, while mutation in this gene resulted in freezing tolerance and changes in the expression of COR genes (Cheong et al. 2003; Albrecht et al. 2003). CIPKs act downstream of Ca2+ signal but upstream of transcription factors regulating cold stress (Kim et al. 2003; Pandey 2008). CBLs relay the Ca2+ signal by interacting with and regulating the family of CIPKs. The role of CIPK3 in cold signaling was shown via changes in expression pattern of RD29A (Responsive to desiccation 29A), KIN1 (cold-inducible1), and KIN2 (cold-inducible2) genes in Arabidopsis (Kim et al. 2003). In rice, overexpression of OsCIPK03, OsCIPK12, and OsCIPK15 cause increased tolerance toward cold, drought, and salt stress (Xiang et al. 2007). Another study suggested that chilling hypersensitivity of cbl1 mutant (Cheong et al. 2003) indicates its role in the regulation of cold response by interacting with CIPK7 (Huang et al. 2011).
Protein phosphatases also act as Ca2+ sensors. The Arabidopsis protein phosphatase 2C, AtPP2CA, is cold inducible, and attains maximum expression within 12 h and thereafter remain high (Tähtiharju and Palva 2001). Monroy et al. (1998) showed that cold-induced inactivation of protein phosphatase 2A (PP2A) was mediated by Ca2+ influx in alfalfa cells. This suggests a possible role of protein phosphatases in Ca2+-mediated cold stress signaling pathway.
4 Reactive Oxygen Species
Both low and freezing temperatures result in the oxidative stress due to generation of ROS, which includes hydrogen peroxide (H2O2), superoxide anion (O2 −), and hydroxyl radical (HO·) (Halliwell 2007). ROS accumulates in cells exposed to abiotic stresses and mediate signaling and gene expression even at moderate level (Chinnusamy et al. 2007; Renaut et al. 2008). ROS signals can induce the activation of redox-responsive proteins, such as transcription factors and protein kinases (Hung et al. 2005; Chinnusamy et al. 2007). ROS is also known to be involved in the regulation of Ca2+ channels (McAinsh and Pittman 2009), mitogen-activated protein kinase (MAPK) cascade (Knight and Knight 2001; Mittler et al. 2004; Hung et al. 2005; Colcombet and Hirt 2008), and activation of ZAT12 TF (Dat et al. 2000; Mittler et al. 2004; Davletova et al. 2005). The Arabidopsis MPK1, MPK2, MPK3, MPK4, MPK6, and MPK7 are induced by H2O2 (Kovtun et al. 2000; Nakagami et al. 2006; Doczi et al. 2007; Ortiz-Masia et al. 2007). MAP kinase activity has been shown to be enhanced by cold in alfalfa (Jonak et al. 1996; Sangwan et al. 2002) and Arabidopsis (Ichimura et al. 2000). Arabidopsis MPK4 and MPK6 were found to be phosphorylated by MKK2 (MAP kinase kinase2) when exposed to cold stress. Overexpression of MKK2 exhibited upregulation of CBF/DREB1s and cold tolerance (Teige et al. 2004). The cold activation of SAMK, an alfalfa MPK, was inhibited by blocking the influx of extracellular Ca2+ and an antagonist of CDPKs, suggesting that Ca2+ fluxes and CDPKs are essential for the activation of MPK cascades in alfalfa (Sangwan et al. 2002).
Excessive accumulation of ROS leads to cellular injury, which ultimately leads to death of the plant due to damage of photosystem II reaction center and membrane lipids (Prasad et al. 1994; Suzuki and Mittler 2006). The Arabidopsis fro1 (frostbite1) mutant, which constitutively accumulates high levels of ROS, exhibited impaired expression of COR genes and hypersensitivity to chilling and freezing. FRO1 encodes the Fe–S subunit of complex I (NADH dehydrogenase) of the respiratory electron transfer chain in mitochondria, and its disruption leads to high levels of ROS generation (Lee et al. 2002a, b; Chinnusamy et al. 2007). Another Arabidopsis mutant, chy1, which is defective in a peroxisomal β-hydroxyisobutyryl-CoA hydrolase involved in fatty acid β-oxidation and valine catabolism, also accumulates high levels of ROS. The chy1 mutant showed a reduced cold induction of CBF genes, and is defective in chilling and freezing tolerance (Zhu et al. 2007). Arabidopsis AtHAP5A (heme-associated protein HAPs, also known as NUCLEAR FACTOR Y subunit A/B/C) binds to AtXTH21, and these proteins inhibit cold stress-induced ROS accumulation and modulate freezing stress resistance via CBF-independent pathway (Shi et al. 2014).
5 Regulation of Gene Expression During Cold Stress
5.1 ABA-Independent Cold Signaling Pathway
5.1.1 CBF/DREB-Responsive Pathway
Several COR genes such as Arabidopsis COR15A (Baker et al. 1994), COR78/RD29A (Yamaguchi-Shinozaki and Shinozaki 1994), the Brassica napus gene BN115 (Jiang et al. 1996), and the wheat gene WCS120 (Ouellet et al. 1998) are involved in cold acclimation and enhance freezing tolerance of plants in response to low temperature. Yamaguchi-Shinozaki and Shinozaki (1994) identified two 9-bp cis-acting DNA regulatory elements (TACCGACAT), in the promoter region of RD29A gene, which contained the conserved low-temperature DRE core sequence (CCGAC). DRE was named as C-repeat by Thomashow and colleagues (Baker et al. 1994). This cis-element, also referred to as LTRE (Low temperature responsive element), was present in the promoter of other COR genes and was found to be crucial for low temperature response. While many of the changes in gene expression that occur during cold acclimation in response to low temperature are mediated by ABA (Bray 1993; Welin et al. 1994), the C-repeat/DRE appears to be mainly regulated by ABA-independent pathway (Yamaguchi-Shinozaki and Shinozaki 1994). The CBF/DREB1 pathway that regulate ABA-independent expression of COR genes was first identified in Arabidopsis (Stockinger et al. 1997; Liu et al. 1998), and now it is the most characterized pathway involved in cold acclimation and freezing tolerance across evolutionarily diverse plant species.
C-repeat binding factors (CBFs) or dehydration-responsive element binding factor 1 (DREB1) are the transcription factors belonging to ethylene-responsive element binding protein/APETALA2 (EREBP/AP2) family (Stockinger et al. 1997; Liu et al. 1998) and DREB subfamily (Chen et al. 2009). DREB subfamily is further subdivided into six groups A-1 to A-6. DREB1/CBF-like genes belong to the A-1 subgroup and are responsive to low temperature. The CBF/DREB1 were first isolated and characterized in Arabidopsis (Stockinger et al. 1997; Liu et al 1998). In Arabidopsis, three cold inducible CBF/DREB1 genes are known, which are present in tandem on chromosome 4 in the order: DREB1B/CBF1, DREB1A/CBF3, and DREB1C/CBF2 (Medina et al. 1999). The CBFs have two signature sequences, PKK/RPAGRxKFxETRHP and DSAWR, located immediately upstream and downstream of AP2 domain, respectively (Canella et al. 2010). These sequences are highly conserved in the CBF proteins from different plant species. They have recognition sites for protein kinase C and casein kinase II. All the three CBF genes are induced within 15 min of plants exposure to cold, and they induce transcription of CBF/DRE-regulated COR genes accumulated within 2 h of exposure to cold stress (Gilmour et al. 1998). The absence of CRT sequence (CCGAC) in the promoters of CBF genes and negligible effect of overexpression of CBF1 on CBF3 transcript levels suggested that the CBF gene family does not involve auto-regulation (Gilmour et al. 1998), rather it is controlled by a set of interacting transcription factors (Vogel et al. 2005; Chinnusamy et al. 2003, 2010; Agarwal et al. 2006; Doherty et al. 2009). Ectopic expression of CBFs in transgenic Arabidopsis plants activated the expression of COR genes and enhanced freezing tolerance without a prerequisite low temperature acclimation (Jaglo-Ottosen et al. 1998; Liu et al. 1998; Kasuga et al. 1999). Microarray analysis of CBF-overexpressing transgenic plants revealed several CBF target genes involved in various pathways such as signaling, transcription, osmolyte biosynthesis, ROS detoxification, membrane transport, hormone metabolism, and stress response (Fowler and Thomashow 2002; Maruyama et al. 2004). Overexpression of AtCBF1/3 has increased cold, drought, and salt stress tolerance in Brassica species (Jaglo et al. 2001), wheat (Pellegrineschi et al. 2004), tomato (Hsieh et al. 2002), tobacco (Kasuga et al. 2004), and rice (Oh et al. 2005). However, the constitutive overexpression of CBFs under the transcriptional control of the 35S cauliflower mosaic virus promoter in transgenic plants resulted in severe growth retardation under normal growth conditions in diverse plant species such as Arabidopsis, B. napus, tomato, potato, and rice. This negative impact on the plant growth was reduced by expressing the CBF genes under the control of stress inducible RD29A promoter instead of the constitutive CaMV 35S promoter (Kasuga et al. 1999).
Molecular analysis with a cbf2 null mutant of Arabidopsis showed that cbf2 null mutation conferred enhanced expression of DREB1B/CBF1 and DREB1A/CBF3 and leads to freezing tolerance. This suggested that DREB1C/CBF2 is a negative regulator of DREB1B/CBF1 and DREB1A/CBF3 (Novillo et al. 2004). Studies on CBF1-RNAi and CBF3-RNAi lines demonstrated that these are involved in the regulation of other CBF/DREB1s genes and they induce cold acclimation by activating same set of CBF/DREB1 regulon genes. The functions of CBF1 and CBF3 appear to be different from that of CBF2 (Novillo et al. 2007). CBF3 appears to be a negative regulator of CBF2 (Chinnusamy et al. 2003, 2006). Whole transcriptome analysis of transgenic Arabidopsis plant overexpressing CBF have shown that only 12 % of the COR genes are controlled by the CBF regulon, indicating that other regulatory pathways are also activated in response to low temperature. Transcription factors such as AP2/ERF factors RAP2.1 and RAP2.6, and the C2H2-type zinc finger STZ/ZAT10 belong to the CBF regulon (Fowler and Thomashow 2002; Vogel et al. 2005).
The homologs of Arabidopsis CBF have cloned and characterized from several species such as Brassica napus (Jaglo et al. 2001; Gao et al. 2002), barley (Xue 2003), rice (Dubouzet et al. 2003), wheat (Shen et al. 2003), corn (Qin et al. 2004), tomato (Zhang et al. 2004), sweet cherry (Kitashiba et al. 2004), alfalfa (Pennycooke et al. 2008a, b), potato (Pennycooke et al. 2008a, b), etc.. Overexpression of these genes in Arabidopsis and other plants conferred enhanced tolerance to multiple abiotic stresses. Overexpression of OsDREB1A (Dubouzet et al. 2003), maize DREB1 (Qin et al. 2004), a modified AP2/ERF transcription factor from Brassica napus (Xiong et al. 2013), Vaccinium myrtillus CBF/DREB1 (Oakenfull et al. 2013) in transgenic Arabidopsis plants resulted in enhanced freezing tolerance. Transgenic barley lines overexpressing TaCBF14 and TaCBF15 exhibited enhanced freezing tolerance (Soltesz et al. 2013).
5.2 Regulation of CBF/DREB1 Transcription Factors
5.2.1 Inducer of CBF Expression (ICE1)
A systematic genetic analysis of transgenic Arabidopsis plant expressing the firefly luciferase gene under the control of CBF3 promoter led to the identification of a constitutively expressed and nuclear-localized transcription factor, Inducer of CBF expression 1 (ICE1). It encodes an MYC-type basic helix-loop-helix transcription factor that binds to the MYC recognition elements (CANNTG), known as ICE box, present in the promoter of CBF3 gene, which in turn regulates the expression of downstream genes during cold acclimation leading to freezing tolerance (Chinnusamy et al. 2003; Chen et al. 2009). The ice1 mutant is highly sensitive to both chilling and freezing temperature and induction of CBF3 gene was impaired in the mutant. Constitutive overexpression of ICE1 in transgenic Arabidopsis conferred enhanced expression of CBF3, CBF2, and downstream COR genes resulting in freezing tolerance Figure 6.1 (Chinnusamy et al. 2003).
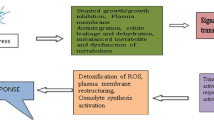
Schematic diagram of regulatory networks involved in low temperature responses. Plants sense low temperature through membrane rigidification and cellular changes that induce calcium signaling and activate protein kinases. Constitutively expressed ICE1 is activated by cold stress via sumoylation and phosphorylation. SIZ1, a SUMO E3 ligase causes sumoylation of ICE1 at K393, which is critical for ICE1 activation of transcription of CBFs and repression of MYB15. HOS1, a RING finger ubiquitin E3 ligase, mediates the ubiquitination and proteosomal degradation of ICE1 and, thus, negatively regulates CBF regulons. CBFs regulate the expression of COR genes that confer freezing tolerance. The expression of CBFs is negatively regulated by MYB15 and ZAT12. CBFs induce the expression of ZAT10, which downregulate the expression of COR genes. Cold-upregulated LOS2 represses the transcription of ZAT10. ZAT10 and ZAT12 are two C2H2 zinc finger transcription factors. Broken arrows indicate negative regulation; purple arrows indicate post-translational regulation; activation solid arrows indicate activation, the (….) indicate unknown cis-elements. Abbreviations: CBF, C-repeat binding factor (an AP2-type transcription factor); CRT, C-repeat elements; DRE, dehydration-responsive elements; HOS1, high expression of osmotically responsive genes1; ICE1, inducer of CBF expression 1 (an MYC-type bHLH transcription factor); LOS2, low expression of osmotically responsive genes 2; MYB, myeloblastosis; MYBR/MYCR, MYB/MYC transcription factor recognition sequence; SIZ1, SAP and MiZ1 (a SUMO E3 ligase); ROS, Reactive oxygen species; Phosphorylation SUMO protein Ubiquitin
Transcriptome analysis of Arabidopsis genome in response to cold revealed that about 40 % of cold-regulated genes, in particular 46 % of cold-regulated transcription factor genes are impaired in dominant ice1 mutant (Lee et al. 2005a, b). Therefore, ICE1 is considered as a master regulator of CBFs and many other cold-responsive regulons. ICE1 is functionally conserved in higher plants. During ambient temperature ICE1 remains in an inactive state at warm temperatures and is activated upon exposure to cold, inducing CBF gene expression. ICE1 mainly affects the expression of CBF3/DREB1A whereas overexpression analysis showed that ICE2 (At1g12860, a homolog of ICE1) induces the expression of CBF1/DREB1B and confers increased freezing tolerance in Arabidopsis after cold acclimation (Fursova et al. 2009).
ICE1 homologs have been identified in barley (Skinner et al. 2006; Tondelli et al. 2006), wheat (Badawi et al. 2008), maize (Hu et al. 2011), rice (Nakamura et al. 2011), tomato (Miura et al. 2012; Feng et al. 2013), tea (Wang et al. 2012), banana (Zhao et al. 2013), wild grapes (Dong et al. 2013; Xu et al. 2014), orchid (Peng et al. 2014), etc. Overexpression of Arabidopsis ICE1 enhanced proline accumulation and improved chilling tolerance in rice (Xiang et al. 2008) and cucumber (Liu et al. 2010). As in case of Arabidopsis, the ICE1 homologs of wheat TaICE141 and TaICE187 are constitutively expressed and induce the expression of the wheat group IV CBFs, which are associated with freezing tolerance. Overexpression of TaICE141 and TaICE187 increased the CBF/DREB1-dependent COR gene expression and freezing tolerance in Arabidopsis (Badawi et al. 2008). Overexpression of SlICE1 enhances chilling tolerance in tomato through increased accumulation of antioxidants, free amino acids, amines and sugars in red tomato fruits of SlICE1-overexpressing plants (Miura et al. 2012). Cold stress upregulates OsICE1 and OsICE2 and OsDREB1B, OsHsfA3 (rice heat shock factor A3), and OsTPP1 (rice trehalose 6-phosphate phosphatase) in rice (Nakamura et al. 2011). Overexpression of ICE1 or CBFs enhanced cold and other abiotic stress tolerance in different species suggesting evolutionarily conserved role of ICE1–CBF pathway in cold tolerance (Table 6.1).
Transcriptome analyses revealed that ICE1 regulates several families of TFs in addition to CBF family (Chinnusamy et al. 2003; Lee et al. 2005a, b). The results from transgenic overexpression in Arabidopsis and other plants support this, as ICE1 overexpression conferred tolerance to cold and other abiotic stresses in these transgenic plants through diverse mechanisms (Table 6.1). In addition to the vegetative tissues, ICE1–CBF pathway also regulates cold tolerance of fruits. Banana ripening-induced MaNAC1, an NAC (NAM, ATAF1/2, and CUC2) transcription factor (TF) gene is involved in cold tolerance of banana fruits. MaICE1 binds to the promoter of MaNAC1 and induces the expression of MaNAC1. Interestingly, MaNAC1 interacted with MaCBF1 in yeast two-hybrid (Y2H) and bimolecular fluorescence complementation (BiFC) analyses. These results suggest that the cold induction of MaNAC1 in banana fruits is mediated by ICE1–CBF cold signaling pathway (Shan et al. 2014).
In addition to chilling temperature (<10 °C)-mediated acclimation, responses to moderate temperature decrease at the non-extreme stress range are also regulated by ICE1 in Arabidopsis. Transgenic Arabidopsis plants expressing CdICE1 when acclimated at 16 °C showed ICE1-mediated induction of miR398–CSD pathway (Chen and Thelen 2013). Shifting of Arabidopsis plants from 28 °C to 22 °C resulted in induction of several genes including the BON1-ASSOCIATED PROTEIN1 (BAP1) gene. BAP1 is under the transcriptional control of ICE1. The ice1 mutant showed low induction of BAP1 and enhanced resistance to Pseudomonas syringae pv tomato (Zhu et al. 2011). Mutational analysis of 125 bp sequence of CBF2 promoter identified two cis-acting elements, ICEr1 and ICEr2 that function together and induce gene expression in response to cold. ICEr1 contains CACATG sequence, which has recognition site for bHLH proteins, CANNTG. So, ICEr1 is the potential binding site for ICE1 (Zarka et al. 2003). In the promoter of maize, ZmDREB1, after cold treatment histones H3 and H4 gets hyperacetylated and DNA demethylation, occur in the ICE1-binding region, also followed by chromatin decondensation, indicating the role of chromatin in the regulation of CBF/DREB1 by ICE1 (Hu et al. 2011).
5.3 Regulation of ICE1
Ubiquitination of ICE1 is mediated by a negative regulator of CBF regulon, HOS1, which is a functional RING-finger protein having E3 ubiquitin ligase activity for proteasomal protein degradation (Dong et al. 2006). The COR genes, such as RD29A, COR15A, KIN1, and ADH, showed enhanced expression in loss-of-function hos1 mutant plants (Ishitani et al. 1998). It encodes a protein of 915 amino acids and contains a short motif near N-terminus, which is similar to RING-finger protein found in the inhibitor of apoptosis (IAP) group of animal protein (Lee et al. 2001). At normal growth temperatures, HOS1 protein resides in the cytoplasm but moves into nucleus upon exposure to cold (Lee et al. 2001). Yeast two-hybrid assays showed that HOS1 physically interacts with ICE1 and mediates its degradation by polyubiquitination both in vitro and in vivo. Once the CBF regulon is activated by ICE1, ICE1 protein is degraded during prolonged duration of cold stress and cold-induced proteolysis of ICE1 is impaired in hos1 mutant. Overexpression of HOS1 in Arabidopsis transgenic plants lowers freezing tolerance and decreases the expression of CBFs and their downstream genes (Dong et al. 2006). The hos1-1 mutant plant showed early flowering under short day conditions, and the expression level of Flowering Locus C (FLC) was downregulated in the mutant plant, indicating its role in vernalization (Ishitani et al. 1998). HOS1 has also been demonstrated to cause the degradation of CO (CONSTANS) under cold stress and regulate Arabidopsis flowering (Jung et al. 2012; Lazaro et al. 2012). The involvement of HOS1 in modulating flowering time in Arabidopsis in response to low ambient temperature was later shown by Lee et al. (2012a, b, c). Under cold stress, HOS1 binds to FLC chromatin, preventing its association with histone deacetylase 6 (HDA6) resulting in FLC induction and delayed flowering (Jung et al. 2013). In wheat, Triticum durum, the RING-finger E3 ligase TdRF1 (RING-finger protein1) interacts with another RING-finger E3 ligase, WVIP2 (wheat viviparus1 interacting protein2), and both of these genes are upregulated by cold treatment (Guerra et al. 2012). HOS1 has been suggested to affect cold signaling partially through altered mRNA export and disruption of circadian clock (MacGregor et al. 2013). A novel HOS1 gene, VvHOS1 from “Muscat Hamburg” grapevine (Vitis vinifera) has been isolated and characterized. Overexpression of VvHOS1 in Arabidopsis reduced the tolerance of plant to cold, drought, and salt stress. It was also shown to negatively regulate the expression level of two stress-responsive genes, AtRD29A and AtCOR47 (Li et al. 2014).
Polyubiquitination of ICE1 is blocked by SUMO (Small Ubiquitin Modifier) E3 ligase SIZ1 (SAP and Miz1)-mediated sumoylation of ICE1 at K393 position, thereby increasing ICE1 activity (Miura et al. 2007). The siz1 null mutant is impaired in the accumulation of SUMO conjugates and result in reduced expression of CBFs and COR genes, thus leads to hypersensitivity to chilling and freezing stresses. However, there is enhanced expression of AtMYB15, a negative regulator of CBF expression, in siz1 mutant (Miura et al. 2007). A K393R substitution in ICE1 blocks SIZ1-mediated sumoylation. It was studied that the serine 403 residue of ICE1 protein regulates the transactivation and stability of ICE1 during cold stress. Substitution of serine 403 by alanine in ICE1 increased ICE1 transactivation activity in the protoplast of Arabidopsis. Transgenic plants overexpressing ICE1 (S403A) showed higher level of freezing tolerance and expression of COR genes as compared to ICE1 (WT) in Arabidopsis. Cold treatment did not alter the protein level of ICE1(S403A) whereas the level of WT ICE1 protein was significantly reduced (Miura et al. 2011).
5.4 CAMTA Transcription Factor
The CaM binding transcriptional activator (CAMTA) family was found to regulate the expression of CBF2 gene during cold stress by binding to the CCGCGT sequence (CM2 motif) present in the promoter of CBF2 gene. Mutation of camta3 resulted in 50 % reduction in cold-induced transcripts of CBF2. However, this mutation has no effect upon cold acclimation of the plant but the double mutant camta1/camta3 altered the freezing tolerance in the plant (Doherty et al. 2009).
5.5 Dead Box RNA Helicase
5.5.1 LOS4 (Low Expression of Osmotically Responsive Gene 4)
LOS4 encodes a DEAD box RNA helicase, AtNUP160, and is localized in nucleus. It is involved in cold-responsive gene expression and freezing tolerance (Gong et al. 2005). It is essential for the mRNA export in the plant cell. Induction of CBFs and the downstream target genes is impaired in los4-1 mutants and plant is hypersensitive to cold. Overexpression of CBF3 gene in los4-1 mutants reverses the sensitivity effect (Gong et al. 2002). Another mutant cryophyte (los4-2) was reported to be more freezing tolerant than the wild type Arabidopsis plant due to enhanced induction of CBF2 in the mutant (Gong et al. 2005).
5.5.2 Regulator of CBF Gene Expression1 (rcf1-1)
A cold inducible DEAD box RNA helicase, RCF1, was identified through a forward genetic approach where CBF2 promoter was fused to a firefly luciferase reporter gene (CBF2:LUC). It functions in maintaining proper splicing of pre-mRNAs of nuclear encoded genes and is important for cold-responsive gene regulation. The rcf1-1 mutant was found to be hypersensitive to chilling and freezing stress, while overexpression of RCF1 in Arabidopsis increased freezing tolerance (Guan et al. 2013).
5.6 Negative Regulators of CBF Regulon
MYB transcription factor: An R2R3 type MYB transcription factor binds to the MYB recognition elements (MYBRS) in the promoter region of CBF/DREB1 and negatively regulates its expression and thereby freezing tolerance. Increased expression of MYB15 in ice1 mutant during cold stress revealed that ICE1 represses the activity of MYB15 either directly by binding to its promoter or indirectly through its downstream genes (Chinnusamy et al. 2003; Badawi et al. 2008). AtMYB15 physically interacts with ICE1 in yeast-two hybrid and in vitro pull down assay (Agarwal et al. 2006). It binds to the promoter region of CBF1, CBF2, and CBF3 genes. Transgenic plants overexpressing MYB15 were sensitive to freezing stress and there was decrease in the transcript levels of CBF genes. Knockout mutants of MYB15 displayed increased freezing tolerance and expression of CBF genes. The downstream cold-responsive genes remain unaffected by overexpression or mutation of MYB15 in plants. MYB15 is localized in nucleus and is present in low levels in all tissues. Upon exposure to cold, its expression is upregulated (Agarwal et al. 2006). In the siz1 mutant, enhanced induction of AtMYB15 was reported (Miura et al. 2007). Another MYB transcription factor, AtMYB14, was found to positively regulate cold tolerance in Arabidopsis. Knockdown mutants of AtMYB14 led to increased cold tolerance and enhanced expression of CBF and downstream genes (Chen and Thelen 2013). Similarly overexpression of a single repeat R3-MYB transcription factor, MYBC1, increased cold tolerance whereas the mybc1 mutants were sensitive to freezing stress (Zhai et al. 2010).
ESK1: The constitutively freezing tolerant eskimo1 (esk1) mutant of Arabidopsis was identified through a freezing tolerant genetic screening. The esk1 plants accumulated 30-fold higher levels of free proline than that in the wild type plants. Additionally, the expression level of RAB18, a cold-responsive LEA II gene was threefold higher. This suggested that ESK1 acts as a negative regulator of cold (Xin et al. 1998). ESK1 encodes a 56.7-kDa protein that contains plant-specific DUF231 (domain of unknown function 231) domain. Transcriptome analysis of the esk1 mutant revealed 312 genes with altered expression, 173 genes were upregulated and 139 were downregulated. The expression of only 12 % of the genes showed overlap with genes regulated by cold acclimation pathway. Other genes were found to be regulated by salt, osmotic and ABA treatment, suggesting that esk1-induced freezing tolerance pathway is distinct from CBF-dependent pathway (Xin et al. 2007).
ZAT12 is a C2H2 zinc finger protein that contains an EAR motif-like sequence, which functions as a repressor domain. It plays a role in cold acclimation and oxidative stress (Vogel et al. 2005). Transcriptome analysis of ZAT12 overexpressing Arabidopsis plant, ZAT12 regulon consists of 24 COS (cold standard set) genes, nine of which are cold-induced and rest are cold-repressed genes (Vogel et al. 2005; Chinnusamy et al. 2007). Constitutive expression of ZAT12 reduced the expression of CBF genes in response to low temperature, indicating that ZAT12 negatively regulates CBF cold-responsive pathway (Vogel et al. 2005; Chinnusamy et al. 2007).
ZAT10/STZ: Molecular analysis of los2 mutant of Arabidopsis led to the identification of another C2H2 zinc finger protein, ZAT10/STZ, as a negative regulator of COR genes. The expression of ZAT10/STZ is induced rapidly and transiently upon cold exposure in the wild type plant and there is increased induction of ZAT10 in los2 mutant. Transient expression studies of ZAT10 in Arabidopsis have shown that ZAT10 represses the expression of RD29A gene (Lee et al. 2002a, b). ZAT10/STZ binds to the STZ recognition site at −554 to −522 in the promoter region of RD29A gene. It binds specifically to A(G/C)T cis-element within the EP2 sequence of RD29A gene (Lee et al. 2002a, b).
LOS2 is a bifunctional enolase, which negatively regulates the expression of ZAT10 by binding to the MYC recognition elements present in the promoter of ZAT10. The expression of COR genes COR/KIN/RD/LTI is impaired in los2 mutant, while there is no difference in the expression levels of CBF genes, indicating that LOS2 has a functional role downstream of the CBF transcription factors and act as a positive regulator of COR genes (Lee et al. 2002a, b). This effect was similar to that of SFR6 (sensitive to freezing) gene. Mutation in SFR6 leads to reduced expression of COR genes having CRT/DRE elements, while the expression of CBF genes remains unaltered, suggesting that SFR6 may be the transcriptional activator of COR genes (Knight et al. 1999).
FIERY2 (FRY2): The FRY2 locus was identified in Arabidopsis by P RD29A ::LUC reporter gene-based genetic screening. The fry2 mutant exhibited an increased expression of COR genes such as RD29A, COR47, COR15A and KIN1 as compared to wild type when treated with cold, salt or ABA. The transcript levels of CBF genes were also higher in fry2 mutant suggesting that FRY2 is a negative regulator of CBF/DREB1 transcription factor (Xiong et al. 2002). Genetic analyses of fry2 mutant in Arabidopsis have shown that FRY2/CPL1 encodes a novel transcriptional repressor having two double-stranded RNA binding domains and a region homologous to the catalytic domain of RNA polymerase II C-terminal domain phosphatases found in yeast and in animals. This region controls transcription and mRNA processing by dephosphorylation of RNA polymerase II (Koiwa et al. 2004; Xiong et al. 2002).
5.7 CBF-Independent Pathways
Cold-responsive pathways are mediated both in ABA-independent and -dependent manner and ABA acts synergistically with cold signaling (Xiong et al. 1999). The ABA-dependent pathway is regulated by transcription factors belonging to bZIP such as ABRE binding factors (AREBs), MYC and MYB families. SCOF1 is a cold-inducible C2H2 zinc finger protein isolated from soybean. Overexpression of SCOF1 in Arabidopsis resulted in increased COR gene expression and enhanced cold tolerance in non-acclimated plants. SCOF1 does not bind directly to the DRE or ABRE sequences, rather facilitates the DNA binding of a cold inducible bZIP transcription factor, soybean G-box binding factor 1 (SGBF1) to ABRE and induces ABA-dependent cold acclimation pathway (Kim et al. 2001).
High expression of osmotically responsive genes, HOS9 was identified by RD29A::LUC genetic screen. HOS9 encodes a homeodomain MYB transcription factor that is localized to the nucleus and is required for basal freezing tolerance (Zhu et al. 2004). Microarray analysis revealed that HOS9 controls expression of about 175 genes that does not belong to the CBF regulon. Also, 41 of these genes targeted by HOS9 were reported to be cold-induced. Knockout mutants of HOS9 gene resulted in reduced basal and acquired cold tolerance. The hos9 mutation also resulted in the alteration of other characteristics including trichome development, growth rate, and flowering time. These results suggested that HOS9 is involved in plant growth and development and affects freezing tolerance in a CBF-independent manner (Zhu et al. 2004).
Overexpression of OsMYB4 in Arabidopsis plants enhanced COR gene expression, increased proline content and exhibited chilling and freezing tolerance (Vannini et al. 2004; Pasquali et al. 2008). OsMYBS3 and OsMYB2 confer increased tolerance in response to cold in rice (Su et al. 2010; Yang et al. 2012).
6 Transcriptomics Approach
Transcriptomics or transcript profiling is a widely used technique to study spatial and temporal gene expression and their regulation by various internal or external stimuli. Several techniques, such as DNA microarrays, Expressed Sequence Tags (ESTs), Serial Analysis of Gene Expression (SAGE), Digital Gene Expression (DGE) profiling, and Next-Generation Sequencing (NGS)-based tools such as RNA sequencing (RNA-seq) are being exploited to capture and identify the differentially regulated genes during stress conditions in plants (Cramer et al. 2011; Lister et al. 2009; Wang et al. 2013) (Table 6.2). It provides an in-depth knowledge of the global gene expression pattern of plant response to stress stimuli (Jung et al. 2003).
Using cDNA microarrays or whole genome arrays, the expression pattern of genes in response to chilling stress has been analyzed in Arabidopsis, rice, sunflower and several other plants (Seki et al. 2002; Rabbani et al. 2003; Fernandez et al. 2008). Seki and coworkers (2001) used a full-length cDNA microarray for 1,300 Arabidopsis genes, and identified 19 COR genes, among which the newly identified genes were ferritin, a nodulin-like protein, LEA protein and glyoxalase. In a different study, Fowler and Thomashow (2002) reported 306 COR genes using microarray for 8,000 genes. Of these 306 genes, 218 were upregulated and 88 were downregulated and 45 of these COR genes were found to be expressing under the control of CBF1. Differentially regulated genes, between two cultivars of wheat, during cold acclimation were identified by microarray and these genes encode protein kinases, transcription factors, calcium binding proteins and proteins involved in photosynthesis (Gulick et al. 2005). A transcriptome profiling of cassava apical shoots subjected to a progressive cold stress was performed using a 60-mer oligonucleotide microarray representing 20,840 cassava genes, which led to the identification of 508 transcripts as early cold-responsive genes including genes for signal transduction components (MAPK4), transcription factors (RAP2.11 and AP2-EREBP), and active oxygen species scavenging enzymes (catalase 2), as well as photosynthesis-related genes (PsaL). An increase in transcripts of genes encoding ROS scavenging enzymes and the genes for accumulation of total soluble sugars (including sucrose and glucose) were also detected (An et al. 2012). Genome-wide transcriptome analysis uncovered various molecular components involved in chilling tolerance and susceptibility in grapefruit. Out of 30,171 transcripts, 1,345 were found to play a role in low temperature response and were termed as chilling response regulons. Transcripts related to cell wall, defense, photosynthesis, respiration and secondary metabolism were observed to be downregulated, while transcripts encoding membrane proteins, lipid and carbohydrate metabolism, hormone biosynthesis and transcription factors were upregulated under cold stress (Maul et al. 2008). Cold-acclimated wheat plants showed the upregulation of CBF genes, WRKY, kinases, phosphatases and genes involved in signal transduction during freezing stress (Skinner 2009).
Based upon ESTs, high expression of dehydrin, COR genes, antifree proteins (AFPs), and pathogenesis-related (PR) proteins was observed in seabuckthorn (Ghangal et al. 2012). Again, the analysis of ESTs in Vitis amurensis showed the upregulation of ERD genes, chitinases, β-glucanase, ubiquitin ligase, and calcium signaling-related genes in response to low temperature (Zhang et al. 2013). Global transcriptome analysis of chickpea during cold acclimation by cDNA-AFLP approach lead to the identification of several transcripts having functions associated with metabolism, transport, signal transduction, and transcription (Dinari et al. 2013).
SAGE was applied to capture the genes involved in freezing tolerance in Arabidopsis, where around 272 differentially expressed genes were identified from cold-treated leaves. The genes induced by cold mostly included COR genes, alcohol dehydrogenase genes involved in glucose metabolism pathways, and lipid transfer protein genes. COR15a was induced over 300-fold, other upregulated genes were COR47, dehydrin Xero 2, and Rab18 (Jung et al. 2003). SAGE analysis revealed that poor accumulation of COR, lipid transfer protein and β-amylase in Arabidopsis pollen are the cause of cold sensitivity (Lee and Lee 2003). Analysis of five different SAGE libraries constructed from Arabidopsis leaf tissues collected at various durations of exposure to cold identified 920 low-temperature responsive genes. Consequently, transcripts of around 63 genes showing COR gene like expression accumulated after 2 days of cold stress treatment and 47, 19, 13 genes exhibiting pattern of CBF1, CBF2, and CBF3, respectively, were identified (Robinson and Parkin 2008). Long SAGE was also performed in Arabidopsis leaf to study the gene expression during early response to cold (Byun et al. 2009).
Global transcriptome analysis and gene expression profile of Jatropha curcas in response to cold were determined by high-throughput sequencing using Illumina Hi-Seq 2000 followed by DGE analysis. Plants were subjected to 12, 24 and 48 h stress at 12 °C. Several genes such as starch catabolism-related genes, phospholipase Dδ (PLDδ), calcium-dependent protein kinase (CDPK), mitogen-activated kinase (MAPK), protein kinase C (PKC), omega-3-fatty acid desaturase, peroxidase, superoxide dismutase, glutathione reductase, osmoprotectants, LEA proteins, CBFs, HOS1, LOS2, LOS4, ZAT10, and ZAT12 were observed to be upregulated. MYB15 was found to be upregulated after 12 and 24 h of stress but after 48 h it gets downregulated (Wang et al. 2013).
7 Proteomics Approach
Although transcriptome studies have enhanced our understanding of stress-responsive gene expression enormously, it is a poor indicator of predicting the expression level of the proteins. Proteins can be identified by differential display pattern following their separation by two-dimensional electrophoresis (2-DE) (Wittmann-Liebold et al. 2006) or liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) (Fournier et al. 2007) and the structures can be predicted by MALDI-TOF/TOF MS. Abundance in protein levels during cold has been studied in several plants such as Arabidopsis thaliana (Bae et al. 2003; Kawamura and Uemura 2003; Amme et al. 2006), rice (Imin et al. 2004; Cui et al. 2005; Yan et al. 2006), peach (Renaut et al. 2008), chicory (Degand et al. 2009), soybean (Cheng et al. 2010), pea (Dumont et al. 2011), and wheat (Rinalducci et al. 2011).
Analysis of Arabidopsis leaves and nuclear proteome during cold stress have revealed 38 plasma membrane proteins and 54 nuclear proteins (Bae et al 2003; Kawamura and Uemura 2003). The nuclear proteome showed abundant change in expression for DEAD box RNA helicase, serine acetyl-transferase, heat-shock proteins, myrosinase-binding proteins, CaMs, and several other families of transcription factors. Cold induced 60S acidic ribosomal protein P2-A, which is involved in elongation step of protein synthesis (Wahl and Moller 2002). A group of putative proteins containing sequences or motifs relevant to RNA-associated functions such as putative U2 snRNP-A, serine-rich protein with a sequence similar to ribosomal protein S1, and putative DEAD box RNA helicase were revealed. Yan et al. (2006) studied total proteins in rice leaves during cold stress using 2D electrophoresis and identified around 85 differentially expressed proteins by mass spectrometry, which included both novel and known cold-responsive proteins. It also showed the involvement of proteins in regulatory and functional network during chilling stress and supported the idea of protein degradation during low temperature. The photosynthetic proteins were found to be highly susceptible to degradation, 19 fragments of Rubisco large subunit were identified during this study. Disassembly of Rubisco under chilling conditions has been suggested to be the cause for reduction in rate of photosynthesis in Arabidopsis (Yi et al. 2011). In addition, glycine dehydrogenase and RcbA, involved in photorespiration, also get degraded during cold stress. It has been shown that ROS may induce degradation of Rubisco by proteases (Desimone et al. 1996). Proteins such as WAK1, armadillo repeat containing protein, HSP, putative acidic ribosomal protein P3a were found to be upregulated in rice during cold. Hashimoto and Komatsu (2007) also reported several differentially expressed proteins in rice seedlings during low temperature. This study also revealed for the first time the downregulation of phosphoglucomutase in response to cold. Proteins involved in cellulose synthesis, such as UDP-glucose pyrophosphorylase were found to be in abundance inside the cells.
Rice roots subjected to 24–72 h of cold stress initiate the activation of metabolic processes as indicated by the presence of several proteins related to metabolism such as phosphogluconate dehydrogenase, NADP-specific isocitrate dehydrogenase, fructokinase, and cytoplasmic malate dehydrogenase. In addition, higher abundance of pyruvate orthophosphate dikinase precursors (PPDK), aconitate hydratase, glycine dehydrogenase, and enolase were also identified in chilling stress (Lee et al. 2009).
Cold stress generates ROS and their precursors that cause oxidative damage to plants. Plants produce antioxidants and detoxifying proteins to combat these damages. Three peroxidases were found in rice proteome, which can scavenge and control ROS levels in plants (Yan et al. 2006). Glyoxalase I and oxalyl-CoA decarboxylase, which are involved in detoxification and prevention of cell damage, were shown to be upregulated in rice roots during cold stress (Lee et al. 2009). Glyoxalase I functions to detoxify methylglyoxal produced during the stress condition whereas oxalyl-CoA decarboxylase causes decarboxylation of activated oxalate molecule that generates ROS. It also causes generation of hydroxyl or carbonate radicals through its interaction with hydrogen peroxide (Espartero et al. 1995; Urzua et al. 1998). The ROS scavengers such as superoxide dismutase, catalase, and ascorbate peroxidase were found in abundance in chicory roots and chickpea during cold stress (Lee et al. 2009; Heidarvand and Maali-Amiri 2013).
Cold stress leads to pollen sterility in rice. Proteome analysis suggest that breakdown of β-expansin and glycogen phosphorylase may possibly lead to incomplete pollen tube growth and starch formation and, thus causing pollen sterility in plants (Imin et al. 2004). Various proteins associated with metabolism, biosynthesis of cell wall components, antioxidative/detoxifying reactions, and energy production were found to be induced in response to cold in rice (Cui et al. 2005). The comparative proteome study of non-transgenic and rd29A:RdreB1B1 transgenics of strawberry under cold stress revealed that the rd29A:RdreB1B1 transgenics showed higher expression level of Cu/Zn superoxide, LEA14-A, eukaryotic translation initiation factor 5A (eIF5A) and photosynthetic proteins which confers higher cold stress tolerance as compared to non-transgenic plants. It was also inferred that the DREB transcription factor regulates the accumulation of eIF5A (Gu et al. 2013). Proteome analysis of spring wheat cultivar revealed increased accumulation of COR proteins, glycine-rich RNA binding protein, proteins involved in amino acid and prosthetic group metabolism, detoxification of ROS and chaperonins (Rinalducci et al. 2011).
Increased level of RNA binding protein, cp29, has been reported during cold (Amme et al. 2006; Gao et al. 2009). It is localized in chloroplast stroma and is involved in photoprotection mechanism that in turn is regulated by phosphorylation. The sensitive genotype of maize lacks phosphorylation of cp29 resulting in photo-inhibitory damage (Mauro et al. 1997). Cold stress leads to degradation of glycine dehydrogenase and RcbA, which are important components in sustaining electron flow (Yan et al. 2006). Goulas et al. (2006) identified 43 (35 in stroma and 8 in lumen of the chloroplast) differentially expressed proteins that participate in photosynthesis, other plastid metabolic functions, hormone biosynthesis and stress sensing and signal transduction during cold acclimation. Proteomic studies during freeze–thaw injury in onion scales revealed the absence of several dehydrins, proteins associated with energy and secondary metabolism, chaperones and antioxidants (Chen and Thelen 2013). Proteins related to recovery after freeze–thaw were also examined. As a result, numerous proteins such as those involved in ion homeostasis, ethylene production, cell wall remodeling, molecular chaperones, detoxifying enzymes, sucrose and fructokinase were identified (Chen and Thelen 2013). Furthermore, reports suggest the activation of defense-related proteins such as protein disulfide isomerase and disease resistance response proteins in pea roots under low temperature stress (Dumont et al. 2011).
The several types of proteins that are expressed under cold stress include AFPs, dehydrins and LEA proteins, HSPs, cold shock domain proteins (CSDPs), chaperones, pathogenesis-related (PR) proteins, and proteins involved in signal transductions (Heidarvand and Maali Amiri 2010).
-
1.
COR/LEA proteins and dehydrins: LEA proteins are low-molecular-weight proteins (10–30 kDa) and provide protection to plants during stresses (Debnath et al. 2011). They are highly hydrophilic and functions as high molecular osmoprotectants (Kosova et al. 2011).
Dehydrins are the subgroup members of LEA proteins. These are heat stable and rich in glycine. During cellular dehydration caused by abiotic stresses, dehydrins play a role in membrane stabilization and protects other proteins from denaturation (Allagulova et al. 2003). DHNs (dehydrins) are characterized by three highly conserved domains known as the K, Y, and S segments. The various DHN subclasses are defined by the number and order of the Y, S, and K: YnSKn, YnKn, SKn, Kn, and KnS. The SKn- and K-type of DHNs are suggested to be involved in CA processes (Rorat 2006). Induction of dehydrins by cold stress has been reported in Arabidopsis (Nylander et al. 2001; Kawamura and Uemura 2003), wheat (Ohno et al. 2003), rice (Lee et al. 2005a, b), and Citrus fruits (Porat et al. 2004). Wisniewski et al. (2006) suggested freeze-responsive role of dehydrins in both herbs and woody plants. They also found intracellular dehydrin in peach had AFPs-like activity. Renaut et al. (2004) reported the expression of single 100 kDa LEA protein in poplar. Similarly, a dehydrin from Solanum sogarandinum increased chilling tolerance in cucumber (Yin et al. 2006). Dehydrins include COR (cold responsive), LTI (Low temperature induced), RAB (responsive to abscisic acid), KIN (cold-induced), or ERD (early responsive to dehydration) (Theocharis et al. 2012). All of the ERD10, ERD14, COR47, and LTI30 accumulate in response to cold stress in plants (Nylander et al. 2001). ERD10 (SK3) and ERD14 (SK2) proteins in Arabidopsis were known to function as chaperones (Kovacs et al. 2008), whereas COR15a prevents protein aggregation (Nakayama et al. 2008) and thus act as a protectant. Thirteen dehydrin genes DHN1 to DHN13 are known in barley. Two of these, DHN5 and DHN8, were reported to express manifold during cold stress (Heidarvand and Maali Amiri 2010) in barley. Three COR genes that are well-identified dehydrins, Wcs120(K6), Wcor410(SK3), and Wcor14 have been shown to accumulate during the initial response to low temperature in wheat (Ganeshan et al. 2008). Wcs120 and Wcor14 have CBF recognition sequences in their promoter region and they are activated by the CBF transcription factors (Heidarvand and Maali Amiri 2010). A COR protein, WCOR18 showed upregulation in wheat leaves subjected to cold stress (Rinalducci et al. 2011). COR WCOR719 is highly induced by low temperature and protects plants from freezing by preserving the structural integrity of plant cells (Danyluk et al. 1996). Cold acclimation-specific protein 15 (CAS15) was identified in chicory roots under chilling stress (Degand et al. 2009). CAS15 protein contains dehydrin K and S segments and contributes to cold tolerance like dehydrins (Pennycooke et al. 2008a, b).
-
2.
Antifreeze proteins: These proteins accumulate in the apoplastic fluid during cold acclimation and inhibit growth of ice crystals (Griffith and Yaish 2004; Griffith et al. 2005; Yaish et al. 2006). These proteins bind to the surface of ice crystals and affect their shape. They also function to depress the freezing point in cold-acclimated leaves and once the leaves are frozen, these prevent ice recrystallization and formation of larger ice crystals. Thus, protecting the plants from mechanical injury. PR proteins like β-1,3-glucanases, chitinases, or thaumatin-like proteins exhibit antifreeze activities (Griffith and Yaish 2004). Wheat TaIRI (ice recrystallization inhibition) proteins have a peculiar bipartite structure. They have a leucine-rich repeat receptor domain at their N terminus and ice recrystallization inhibition domains at their C terminus (Ouellet 2007). It was reported that the hormone ethylene, salicylic acid, ABA, and drought are involved in regulating antifreeze activity in response to cold in plants (Atici and Nalbantoglu 2003).
-
3.
Heat shock proteins (HSPs): Though mainly known to be heat stress inducible, these are also reported to be accumulated in response to other abiotic stresses (Sabehat et al. 1998). HSP90, HSP70, small HSPs, chaperonins 60 and 20 are highly induced by low temperature (Lopez-Matas et al. 2004). Two Hsp70-like proteins were found to be upregulated in response to low temperature in peach (Renaut et al. 2008). Four cytoplasmic and two mitochondrial proteins of Arabidopsis HSP70 family were also revealed to be upregulated in response to low temperature (Heidarvand and Maali Amiri 2010). During analysis of Arabidopsis nuclear proteome, HSP70-1 was identified (Bae et al. 2003) as a dnaK-type molecular chaperone (Sung et al. 2001). These proteins participate in membrane protection, translation, and translocation into organelles, refolding of denatured proteins and prevent aggregation of denatured proteins (Tsvetkova et al. 2002; Renaut et al. 2006; Timperio et al. 2008).
-
4.
Pathogenesis-related (PR) proteins: Apart from being expressed during pathogen attack, these proteins are also produced during various abiotic stresses. These include β-1,3-glucanases, chitinases, thaumatin-like proteins and lipid transfer proteins (Liu et al. 2003). These are involved in signal transduction during cold stress (Hoffmann-Sommergruber 2000). Four PR proteins, two isoforms of β-1,3-glucanases, one chitinase-3-like protein precursor and one glucan endo-1,3-β-glucosidase, were found in onion scales recovered after freeze–thaw injury (Chen and Thelen 2013).
-
5.
Histones: The exploration of the proteomic response to cold stress in rice seedlings for various durations showed the involvement of histone regulation. Histones H4, H2B.9, H3.2, linker histones H1 and H5 were downregulated after cold stress while H2A.3 was found to be upregulated. Additionally, WD-40 repeat protein that is known to modify histones was upregulated after 96 h of stress (Neilson et al. 2011).
-
6.
Vitamin B biosynthetic proteins: Proteins responsible for folate and riboflavin synthesis (vitamin B9 and B2, respectively) were downregulated while thiamine biosynthetic enzyme (vitamin B1) was expressive at all time points of cold stress in rice (Neilson et al. 2011).
7.1 Plasma Membrane Proteome During Cold Stress
Plasma membrane plays a pivotal role in signal perception, cellular homeostasis, and stress tolerance in plants (Takahashi et al. 2013). Plasma membrane proteomics is being exploited to enhance freezing tolerance in plants. An enhanced accumulation of phospholipase Dδ (PLDδ), a plasma membrane-associated protein, which hydrolyses phospholipids and generates phosphotidic acid (PA), was reported to express in Arabidopsis during cold acclimation (Kawamura and Uemura 2003). Li et al. (2004) showed that knockdown of PLDδ gene results in freezing-sensitive plants, whereas overexpression leads to improve freezing tolerance in Arabidopsis. The PLDδ alterations were not associated with change in expression of COR47 or COR78 and also did not affect the soluble sugars or proline. Lipocalin-like protein (temperature-induced lipocalin, TIL) also exhibits enhanced cold tolerance (Kawamura and Uemura 2003; Uemura et al. 2006).
Cold acclimation leads to an increase in P-type ATPase activity, disassembly of microtubules, and accumulation of different types of dehydrin family proteins on the plasma membrane (Ishikawa and Yoshida 1985; Abdrakhamanova et al. 2003; Kosová et al. 2007). Dehydrins, EDR10 and EDR14, protect proteins and membranes against freeze-induced injury and dehydration (Uemura et al. 2006; Kosová et al. 2007). A novel protein expressed during cold acclimation synaptotagmin 1 (SYT1) was identified in plants, an isoform of which is supposed to be involved in membrane resealing in animals. It was later reported that SYT1 is involved in resealing and repairing of membrane and thus conferring freezing tolerance (Yamazaki et al. 2008).
Analysis of plasma membrane proteome in rice roots during low temperature has revealed the role of proteins involved in membrane permeability and signal transduction in cold response (Hashimoto et al. 2009). Members of annexin and hypersensitive-induced response (HIR) protein families fall under this category. Annexins are Ca2+-dependent membrane binding proteins and help in membrane trafficking and organization (Gerke and Moss 2002), while HIR proteins are involved in ion channel activity (Nadimpalli et al. 2000). In Arabidopsis, two cold-responsive plasma membrane proteins have been suggested to have a positive role in freezing tolerance (Tominaga et al. 2006). Outer membrane lipoprotein like protein (lipocalin-like protein, AtLCN) and ERD14 were found to increase during cold acclimation and affect the cryobehavior of plasma membrane (Kawamura and Uemura 2003). Transgenic plants overexpressing AtLCN showed greater freezing tolerance than wild type plants without cold acclimation (Uemura et al. 2006). ERD14 transgenic plants displayed decrease in intracellular freezing in the protoplast (Uemura et al. 2006).
8 Metabolomics Approach
To survive in cold stress environmental conditions, plant needs to adjust its physiological pathways to restore its metabolic homeostasis in order to accomplish its basic requirements of carbon source, energy, membrane stability, osmolarity, and resistance against ROS produced during stress. Various metabolic pathways are altered in response to cold. Some of the important findings of metabolic adaptations revealed from recent metabolomics studies are summarized in Table 6.3.
8.1 Carbohydrates and Soluble Sugar Levels
Starch is the major energy storage molecule found in higher plants. As abiotic stresses inhibit existing photosynthesis while increasing the maintenance respiration for protection and repair of stress-induced damage, stored starch, and other carbohydrates are converted into soluble sugars and additional metabolic pathways are induced to convert the simple soluble sugars such as glucose and fructose into different other sugars and sugar alcohols. Abundance of soluble sugar confers resistance to cold, as it can be transported to the sink tissues for various metabolic and protective requirements. Comparison of metabolome profile of cold acclimating and non-acclimating plants revealed that more than 50 metabolites including trehalose, putrescine, and ascorbate are upregulated in cold-acclimated plants (Cook et al. 2004). Trehalose is a non-reducing disaccharide of glucose. It can reversibly absorb water and thus, protects plant from desiccation-induced damage. In vivo studies also support that trehalose protects membrane and proteins from deleterious alterations that are caused due to oxidative stress (Cook et al. 2004; Janmohammadi 2012).
8.1.1 Raffinose Family Oligosaccharides
It has been widely reported that starch levels fall drastically in LT conditions (Strand et al. 1997, 1999; Goulas et al. 2006) but at the same time levels of soluble sugar and Raffinose Family Oligosaccharides (RFOs) rise (Bachmann et al. 1994). RFOs are alpha-galactosyl derivatives of sucrose. These are soluble sugars such as trisaccharide (Raffinose), tetrasaccharide (Stachyose), and pentasaccharide (Verbascose) (Bachmann et al. 1994). UDP-galactose is the precursor for RFOs. Their synthesis utilizes a series of alpha galactosyl-transferase in a sequential manner (Bachmann et al. 1994). Enzymes involved in RFOs synthesis include Galactinol Synthase (GS), Raffinose Synthase (RS), and Stachyose Synthase (STS). Comparison of plants capable of cold acclimation and cold-sensitive plants revealed that STS in cold-tolerant plants but not in cold-sensitive plants is detectable at temperature below 15 °C. In Ajuga, stachyose was main carbohydrate found in phloem under cold stress. In populus, RFO levels rise in mid-winter and declines post-winter (Cox and Stushnoff 2001). RFOs have been shown to play cryoprotective role in many plants (Stushnoff et al. 1993; dos Santos et al. 2012; Egert et al. 2013). Subsequently, Raffinose was proposed to function in membrane stabilization. It also seems to appear to interact with phospholipid head groups to increase cold tolerance (Morsy et al. 2007). Raffinose was found to be having major role in membrane stabilization than other monosaccharides and disaccharides (Rontein et al. 2002).
8.1.2 Soluble Sugars
In cold hardy plants, synthesis of sucrose and the activity of enzymes involved in sucrose synthesis significantly rise. Mutations in the enzymes of sucrose synthesis make the plant sensitive to cold. On the other hand, mutants overexpressing enzymes of sucrose synthesis were found to be freezing tolerant than wild type plants (Strand et al. 2003).
Transgenic Arabidopsis plants overexpressing Sucrose Phosphate synthase (SPS) exhibited better photosynthesis, mobilization of sucrose and enhanced freezing tolerance as compared with wild type plants when shifted to 5 °C, while antisense plants were impaired in sucrose mobilization and freezing tolerance (Strand et al. 2003). Exogenous application of sucrose conferred cold tolerance to freeze-sensitive plants. Sucrose supplementation to freezing-sensitive plants also exhibited reduced loss of osmotic responsiveness to a level comparable to cold acclimation of wild type (Uemura et al. 2003).
It has been observed that during initial phase of cold acclimation, transcripts of hexose kinase and fructose kinase are abundantly expressed. These enzymes are involved in synthesis of hexose 6-phosphate and fructose 6-phosphate, which are precursors of sucrose biosynthesis. This leads to accumulation of sucrose in plants during cold acclimation (Kaplan et al. 2007). Sucrose and glucose protects a plant by directly acting as a substrate for cellular respiration or as an osmolyte to maintain cell homeostasis (Janmohammadi 2012). Fructan, a polymer of sucrose, synthesized by fructosyl transferase, helps in membrane stabilization by binding to phosphate and choline head groups in lipid membrane (French and Waterhouse 1993). Fructan accumulation in cold-acclimating plants suggests their role in enhancing cold tolerance (Livingston et al. 2009).
Soluble sugars are also speculated to have regulatory effect on genes under cold stress conditions. COR78 gene expression levels increase when plants under LT are supplemented with sucrose. It appears that sucrose imparts control over COR78 gene and upregulates its expression so as to acclimate plant to cold conditions (Rekarte-Cowie et al. 2008). Likewise, β-amylase also regulates some COR genes (Heidarvand and Maali Amiri 2010).
8.2 Photosynthetic Activity
Many studies document the reduction in photosynthetic carbon dioxide uptake in chilling stress conditions as compared to normal conditions. In vitro assay has shown that at LT neither ribulose-1,5-bisphosphate carboxylase nor electron transport from water to Dichlorophenol Indophenol in isolated thylakoid is inhibited. Reduction in photosynthetic activity was not attributed either to low phosphate concentration or to low carbon dioxide supply or to low sink to source ratio under cold stress (Bauer et al. 1996). Study done on conifers revealed that the rate of photosynthesis decreases in winters due to reduction in chlorophyll content (Hansen et al. 1996).
New leaves of plants subjected to 5 °C expressed high levels of Calvin cycle enzymes along with an increase in total protein content. On the other hand, a very small increase of these enzymes is observed in mature leaves of plants that were first grown at normal temperature and later exposed to 5 °C. This suggested that leaves initiated under cold conditions are metabolically more tolerant to cold stress. Moreover, the levels of enzymes involved in sucrose synthesis in cytosol were found to increase more rapidly than enzymes of starch synthesis in chloroplast (Strand et al. 1997). An increase in Calvin cycle’s enzymes was also observed. This increase might be due to overall increase in proteins during cold stress. However, the increase in Calvin cycle enzymes was still less than that of the enzymes of cytosol involved in sucrose synthesis (Goulas et al. 2006).
DIGE analysis indicated that Rubisco small and large subunit proteins increased in cold stress (Goulas et al. 2006). Heidarvand and group (2013) found that Rubisco levels increase when ATP synthesis and Electron Transport activity declines. Warm grown leaves when exposed to cold stress showed high levels of ATP synthase proteins in stromal compartment. Their levels come back to normal when new leaves develop at 5 °C but are high when plant are suddenly exposed to cold stress. The ATP/ADP ratio in plants also increases under cold stress (Hurry et al. 2000).
8.2.1 Xanthophylls
Xanthophylls are yellow pigments found in leaves, synthesized in plastids. These are oxygenated carotenoids and do not require light for synthesis. Their accumulation is known to surge under cold stress. They protect photosystems from oxidative damage by quenching triplet Chl and singlet oxygen during cold stress (Janmohammadi 2012). Xanthophyll acts as natural antioxidants. Free Zeaxanthin and carotienoids also seem to protect thylakoid membrane from any kind of oxidative damage caused due to change in temperature conditions (Theocharis et al. 2012).
8.2.2 Flavonoids
Flavonoids are secondary metabolites, synthesized by polypropanoid pathway from phenylalanine. These molecules have antioxidant activity and their concentrations increases in cold stress. Thus, stabilizing plants by scavenging oxidative species produced in cold stress (Janmohammadi 2012). Transcriptome analysis has revealed that under cold stress transcripts of flavonoid synthesis increase thereby increasing flavonoid concentration and protecting plant from oxidative damage (Theocharis et al. 2012). Flavonoids also appear to be responsible for cell membrane stabilization as they accumulate in lipid phase of cell membrane at sub-zero temperatures (Korn et al. 2008).
8.3 Membrane Lipids and Fluidity
Changes in temperature have a very prominent effect on permeability and composition of membrane lipids in plants. Alteration in membrane lipid composition allows plant to survive not only in cold but also during intense heat conditions (Campos et al. 2003; Sage and Kubien 2007; Heidarvand and Maali-Amiri 2013). Fatty acid composition in membrane is the key factor involved in determining fluidity of the membrane. Saturated fatty acids ensure more rigidness as they favor hydrophobic interactions in membrane whereas unsaturated FAs increase fluidity of membrane (Campos et al. 2003). It has been observed that in cold-tolerant species saturated to unsaturated fatty acid ratio decreases under cold stress. An alteration in unsaturated fatty acid (UFA) levels has also been reported in cold-stressed plants. In Chickpea, fatty acids with longer carbon chain are more abundant than shorter fatty acids. Along with this a high abundance of 18:3 UFA is seen as compared to 18:1 and 18:2. Thus, indicating importance of FAs elongation and desaturation in tolerance against stress (Heidarvand and Maali-Amiri 2013). In C. apoata, C16:1 contributes toward membrane stability (Campos et al. 2003). High C16:1 in thylakoid membrane promotes LHCII oligomerization which enhances photosynthesis in cold stress conditions. However in cereals, C16:1 decreases in cold stress and so does LHCII oligomerization. This was postulated to regulate energy distribution to avoid effects of photoinhibition caused by cold stress in plants. In cold-stressed plants, lipid peroxidation increases as is evident from high MDA levels in plants experiencing cold conditions.
During recovery from cold stress, cold-tolerant plants ensure membrane stability by accumulating more of lipids that are able to withstand lower temperatures. Phosphatidylcholine (PC) and phosphatidyl ethanolamine (PE) are two phospholipids that are highly abundant in plasmalemma and mitochondrial membrane. PC has lower phase transition temperature than PE. So, in recovery phase plants under cold stress have more PC to PE ratio (Campos et al. 2003).
8.3.1 Electrolyte Leakage Index and Double Bond Index
Abundance of UFA in membrane increases permeability of membrane for solutes and ions. Integrity of membrane is evaluated by changes in Electrolyte Leakage Index (ELI) under stress condition. In a study, it was found that initially ELI increased continuously for 2 h and reach to its maximum at the end of this period immediately followed by a decline till 8 h. Initial increase and eventual decrease in ELI suggest induction of cold tolerance and the stability of membranes that plants try to attain after cold shock (Campos et al. 2003; Heidarvand and Maali-Amiri 2013). Degree of unsaturation in FAs also increases in low temperature grown plants. Double Bond index (DBI) is the parameter used to evaluate the level of UFAs. In many species, it was found that monogalactosyl diacylglycerol (MGDG) to digalactosyl diacylglycerol (DGDG) ratio as well as overall phospholipid content decreases under cold stress. In thylakoid membrane DGDG is known to provide high permeability of ions and more stability to lipids and proteins present in membrane (Campos et al. 2003).
Lipoxygenase (LOX) is an enzyme involved in biosynthesis of jasmonic acid and other secondary metabolites involved in antioxidant activity. Increase in LOX activity and DBI is associated with lower levels of membrane damage and ELI. A protein PsbP, oxygen evolving complex (OEC) proteins, stabilizes water-oxidizing complex of PSII and protects Mn cluster from oxidative stress during LT condition. Its level increases till ELI and MDA levels are high. Thus, it plays a cryoprotective role in stabilizing PSII and so electron transfer and energy generation processes for survival of plant. At the same time, an increase in levels of ATP synthase CF1 epsilon subunit was also observed. It is involved in ATP hydrolysis in chloroplast and mitochondria. Increase of epsilon subunit of CF1 indicates high-energy requirement of cells during cold stress. Kinetic study revealed that at an early time point ELI, MDA, and H2O2 levels are high due to sudden change in temperature. To combat temperature fluctuation, the levels of longer UFAs, LOX, and ATP synthesis increase. Cytochrome c oxidase subunit 6b-1 declined initially but regained its levels after 8 h of cold stress (Heidarvand and Maali-Amiri 2013).
8.4 Polyamines
Polyamines are ubiquitous small aliphatic amines, carrying a positive charge at cellular pH. Polyamines viz. putrescine (diamine), spermidine (triamine), and spermine (tetramine) have been reported to play vital role in modulating cellular and physiological functions. Its polycationic nature allows it to interact with other molecules and exists as acid soluble and acid insoluble conjugates (Kasukabe et al. 2004). They can also interact with proteins involved in defense mechanism against various environmental stresses. Polyamines also suppress ROS during cold stress (Cuevas et al. 2008). H2O2 production in cold-stressed plants was reduced when these plants were supplemented with polyamine but its production increased when polyamine synthesis inhibitor was applied. This clearly proves the role of PA as ROS scavenger (Zhang et al. 2009). Exogenous supply of polyamines to chilling-sensitive plants has suppressed electrolyte leakage resulting in higher chilling tolerance. Cold-stressed plants also have reduced soluble protein content, which reaches to normal levels after exogenous supplementation of putrescine and spermidine (Zhang et al. 2009). Spermidine and spermine are biosynthetic precursors of β-alanine. In cold stress, the metabolic pathway of β-alanine biosynthesis also get upregulated. Thus, pointing toward the possible role of polyamines in cold condition (Töpfer et al. 2013).
In cucumber, cold-sensitive and -tolerant cultivars show different levels of H2O2 and polyamines in cold stress. Cold-tolerant cultivar, contrary to cold-sensitive one, has high levels of free polyamine and low level of H2O2. On application of PA synthesis inhibitor, methyglyoxal-bis-(guanylhydrazone) (MGBG), polyamine level decreased and H2O2 levels increased to a level similar to that of cold-sensitive cultivar (Zhang et al. 2009).
A study showed that exogenous application of polyamine especially spermidine improved the functionality of enzymes involved in antioxidant Halliwell–Asada pathway (Kubis 2003). Polyamines have been shown to protect plasma membrane and act as antioxidant in other stress conditions also (Kim et al 2002). In oats and transgenic Arabidopsis putrescine levels increased at early stage of cold stress but there was no prominent increase in spermine or spermidine levels (Alcázar et al. 2010). Putrescine is a precursor for both spermidine and spermine. Thus, it is hypothesized that spermine and spermidine are under strict regulation (Cuevas et al. 2008). Arg decarboxylase (ADC), involved in synthesis of putrescine, is induced under cold stress. Accumulation of ADC isoforms in Arabidopsis under cold stress also indicates increased synthesis of putrescine (Alcazar et al. 2006). Mutations in ADC gene cause cold sensitivity in Arabidopsis (Cuevas et al. 2008).
8.5 Nitric Oxide
Nitric oxide acts as a signaling molecule and triggers downstream pathways to combat various physiological stress and developmental conditions. In Arabidopsis, nitric oxide is known to be involved in cold stress response. Studies have found that NO levels increases in wild type plants in response to cold stress. NO is synthesized by nitric oxide synthase (NOS) or nitrate reductase (NR). NO-synthase-like enzyme was proposed to produce NO by conversion of arginine to citrulline. However, the presence of such a protein is still under question. The other pathway utilizes enzyme nitrate reductase, which assimilate nitrogen by converting nitrate to nitrite (Zhao et al. 2009; Thakur and Nayyar 2013). NR is encoded by gene nitrate reductase NIA that is upregulated under cold conditions. Production of NO in cold stress is entirely dependent only on NR pathway. Arabidopsis NR double mutant nia1nia2 exhibit defective NO production during cold stress and thus results in low freezing tolerance (Zhao et al. 2009). NO activates or inactivates a protein by S-nitrosylation of their thiol group. This leads to regulation of downstream processes (Abat and Deswal 2009). NO is also known to mediate signaling along with ABA, calcium, and hydrogen peroxide. It also plays key role in stomatal opening and closing under stress conditions (Neill et al. 2008).
8.6 Proline
Proline is a unique amino acid derived from “pyrrolidine.” It acts as small organic cation required to maintain cytosolic acidity and membrane integrity (Janmohammadi 2012; Jouve et al. 2004). Accumulation of proline has been observed in plants tolerant to cold stress, indicting its involvement in cold acclimation (Janmohammadi 2012; Kaur et al. 2011). Proline treatment to cold-sensitive plants reduced electrolyte leakage and stabilized membrane during cold stress. Malondialdehyde (MDA) is the end product of lipid peroxidation and is thus used as a marker for osmotic stress. In cell culture exposed to osmotic stress, MDA levels shoot up but significantly decline after proline supplementation.
The water and chlorophyll levels also increase due to proline treatment in cold-stressed plants (Kaur et al. 2011). Decreased levels of ROS in proline-treated plants describe its role in increased activity of antioxidants. The biochemical assays done by Hong et al. (2000) also supports that proline helps in reduction of free radicals that are formed due to osmotic stress. Even in osmotic stress, proline has been documented to play a role as free radical scavenger (Hong et al. 2000; Jouve et al. 2004).
Another in vitro assay, done to examine free radical scavenging activity of proline, proved that proline accumulation during abiotic stress in plant could suppress ROS produced due to stress condition (Kaul et al. 2008). Cold stress induces lipid peroxidation and methylglyoxal (MG) production that decreases upon exposure to proline and betaine. MG is produced in cells due to abiotic stress along with ROS and is highly toxic to plants (Goulas et al. 2006). It reacts with proteins, carbohydrates, and DNA. Enzymes involved in glyoxalase pathway are stabilized by proline supplementation hence again indicating its role in enhancing antioxidant activity of plant. Exposure of proline also enhanced glutathione-S-transferase and glutathione reductase activity in plants. These enzymes are majorly involved in detoxification pathways (Goulas et al. 2006).
Cold has its effect on flowering and pod generation as well. Proline depletion in cold-stressed plants increases their sensitivity to cold and leads to flower and pod abortion (Kaur et al. 2011). Proline levels were found to increase in cold-stressed plants till fourth day and decreased as cold persisted, to a level lower than normal. However, flowering and pod generation retained in proline-treated plants. Lower proline content was earlier linked with male sterility also. Moreover, upon exogenous application of proline enhances pollen germination.
Concentrations of proline in cold-stressed plants also regulate the carbohydrate levels (Kaur et al. 2011). Chilling-stressed plants have a lower level of sucrose and trehalose. These carbohydrates are quite important for acclimatization of plants in cold stress as explained earlier. Upon proline treatment, plants were shown to acquire normal levels of carbohydrates aiding in cold acclimatization. As a result of proline-induced carbohydrate levels, under chilling stress these were made available to other parts of plant especially pods and flower. Availability of sugar to pod and flower prevented their abortion (Kaur et al. 2011). Increase in proline level is directly associated with increasing NO level in cold stress. Mutants with reduced production of NO also exhibit lower proline content. Exogenous application of NO scavenger has shown to decrease the level of proline in cold-stressed plants (Zhao et al. 2009).
9 Conclusions and Perspectives
Stress disrupts metabolic homeostasis and plants require countless adjustments to acclimate these conditions. Here, we have documented various evidences of several genes and gene networks involved in regulating the cold-mediated responses by several “Omic”-based approaches. With the advent of increasing knowledge yielded from transcriptomic and proteomic-based studies now a shift is being taking place mostly toward metabolomic-based studies to understand the adaptive responses during cold stress. Studies in future are expected to enhance our knowledge on the low temperature responsive transcriptome, proteome, and metabolome from different crops plants. These efforts are essential in order to understand the molecular network of interacting proteins that confers freezing tolerance in crop plants. A concerted approach toward deciphering physiological and metabolic aspects of cold stress is needed to comprehend the phenotype of plants under cold stress. Nevertheless, there remains a major task in connecting these “Omic”-based approaches in different crop plants to develop a holistic ideology to tackle the problem of cold- and freezing-associated damages in the plants.
References
Abat JK, Deswal R (2009) Differential modulation of S-nitrosoproteome of Brassica juncea by low temperature: change in S-nitrosylation of Rubisco is responsible for the inactivation of its carboxylase activity. Proteomics 9(18):4368–4380
Abdrakhamanova A, Wang QY, Khokhlova L, Nick P (2003) Is microtubule disassembly a trigger for cold acclimation? Plant Cell Physiol 44:676–686
Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, Zheng X, Zhu JK (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281:37636–37645
Albrecht V, Weinl S, Balzevic D, D’Angelo C, Batistic O, Kolukisaouglu Ü, Bock R, Schulz B, Harter K, Kudla J (2003) The calcium sensor CBL1 integrates plant responses to abiotic stresses. Plant J 36:457–470
Alcazar R, Marco F, Cuevas JC, Patron M, Ferrando A, Carrasco P, Tiburcio AF, Altabella T (2006) Involvement of polyamines in plant response to abiotic stress. Biotechnol Lett 28(23):1867–1876
Alcázar R, Altabella T, Marco F, Bortolotti C, Reymond M, Koncz C, Carrasco P, Tiburcio AF (2010) Polyamines: molecules with regulatory functions in plant abiotic stress tolerance. Planta 231:1237–1249
Allagulova CR, Gimalov FR, Shakirova FM, Vakhitov VA (2003) The plant dehydrins: structure and putative functions. Biochemistry 68:945–951
Amme S, Matros A, Schlesier B, Mock HP (2006) Proteome analysis of cold stress response in Arabidopsis thaliana using DIGE-technology. J Exp Bot 57:1537–1546
An D, Yang J, Zhang P (2012) Transcriptome profiling of low temperature-treated cassava apical shoots showed dynamic responses of tropical plant to cold stress. BMC Genomics 13:64
Antikainen M, Griffith M (1997) Antifreeze protein accumulation in freezing tolerant cereals. Physiol Plant 99:423–432
Atici O, Nalbantoglu B (2003) Antifreeze proteins in higher plants. Phytochemistry 64:1187–1196
Bachmann M, Matile P, Keller F (1994) Metabolism of the raffinose family oligosaccharides in leaves of Ajuga reptans 1. Plant Physiol 105:1335–1345
Badawi M, Reddy YV, Agharbaoui Z, Tominaga Y, Danyluk J, Sarhan F, Houde M (2008) Structure and functional analysis of wheat ICE (inducer of CBF expression) genes. Plant Cell Physiol 49:1237–1249
Bae MS, Cho EJ, Choi EY, Park OK (2003) Analysis of the Arabidopsis nuclear proteome and its response to cold stress. Plant J 36:652–663
Baker SS, Wilhelm KS, Thomashow MF (1994) The 5′-region of Arabidopsis thaliana cor15a has cis-acting elements that confer cold-, drought- and ABA-regulated gene expression. Plant Mol Biol 24:701–713
Bauer H, Pamer R, Perathoner C, Loidolt-Nagele M (1996) Photosynthetic depression in leaves of frost-hardened ivy is not caused by feedback inhibition via assimilate accumulation. J Plant Physiol 149:51–56
Behnam B, Kikuchi A, Celebi-Toprak F, Kasuga M, Yamaguchi-Shinozaki K, Watanabe KN (2007) Arabidopsis rd29A::DREB1A enhances freezing tolerance in transgenic potato. Plant Cell Rep 26:1275–1282
Benedict C, Skinner JS, Meng R, Chang Y, Bhalerao R, Huner NP, Finn CE, Chen TH, Hurry V (2006) The CBF1-dependent low temperature signalling pathway, regulon and increase in freeze tolerance are conserved in Populus spp. Plant Cell Environ 29:1259–1272
Bouchereau A, Aziz A, Larher F, Martin-Tangui J (1999) Polyamines and environmental challenges: recent development. Plant Sci 140:103–125
Bray EA (1993) Molecular responses to water-deficit. Plant Physiol 103:1035–1040
Byun YJ, Kim HJ, Lee DH (2009) Long SAGE analysis of the early response to cold stress in Arabidopsis leaf. Planta 229:1181–1200
Campos PS, Quartin V, Ramalho JC, Nunes MA (2003) Electrolyte leakage and lipid degradation account for cold sensitivity in leaves of Coffea sp. plants. J Plant Physiol 160:283–292
Canella D, Gilmour SJ, Kuhn LA, Thomashow MF (2010) DNA binding by the Arabidopsis CBF1 transcription factor requires the PKKP/RAGRxKFxETRHP signature sequence. Biochim Biophys Acta 1799:454–462
Chen TH, Murata N (2008) Glycinbetaine: an effective protectant against abiotic stress in plants. Trends Plant Sci 13:499–505
Chen M, Thelen JJ (2013) ACYL-LIPID DESATURASE2 is required for chilling and freezing tolerance in Arabidopsis. Plant Cell 25:1430–1444
Chen M, Xu Z, Xia L, Li L, Cheng X, Dong J, Wang Q, Ma Y (2009) Cold-induced modulation and functional analyses of the DRE-binding transcription factor gene, GmDREB3, in soybean (Glycine max L.). J Exp Bot 60:121–135
Chen J, Tian L, Xu H, Tian D, Luo Y, Ren C, Yang L, Shi J (2012a) Cold-induced changes of protein and phosphoprotein expression patterns from rice roots as revealed by multiplex proteomic analysis. Plant Omics J 5:194–199
Chen L, Chen Y, Jiang J, Chen S, Chen F, Guan Z, Fang W (2012b) The constitutive expression of Chrysanthemum dichrum ICE1 in Chrysanthemum grandiflorum improves the level of low temperature, salinity and drought tolerance. Plant Cell Rep 31:1747–1758
Chen Y, Jiang J, Song A, Chen S, Shan H, Luo H, Gu C, Sun J, Zhu L, Fang W, Chen F (2013) Ambient temperature enhanced freezing tolerance of Chrysanthemum dichrum CdICE1 Arabidopsis via miR398. BMC Biol 11:121
Cheng SH, Willmann MR, Chen HC, Sheen J (2002) Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol 129:469–485
Cheng L, Gao X, Li S, Shi M, Javeed H, Jing X, Yang G, He G (2010) Proteomic analysis of soybean [Glycine max (L.) Meer.] seeds during imbibition at chilling temperature. Mol Breeding 26:1–17
Cheong YH, Kim KN, Pandey GK, Gupta R, Grant JJ, Luan S (2003) CBL1, a calcium sensor that differentially regulates salt, drought, and cold responses in Arabidopsis. Plant Cell 15:1833–1845
Chinnusamy V, Ohta M, Kanrar S, Lee BH, Hong X, Agarwal M, Zhu JK (2003) ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev 17:1043–1054
Chinnusamy V, Schumaker K, Zhu JK (2004) Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J Exp Bot 55:225–236
Chinnusamy V, Zhu J, Zhu JK (2006) Gene regulation during cold acclimation in plants. Physiol Plant 126:52–61
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12:444–451
Chinnusamy V, Zhu JK, Sunkar R (2010) Gene regulation during cold stress acclimation in plants. Methods Mol Biol 639:39–55
Colcombet J, Hirt H (2008) Arabidopsis MAPKs: a complex signalling network involved in multiple biological processes. Biochem J 413:217–226
Cook D, Fowler S, Fiehn O, Thomashow MF (2004) A prominent role for the CBF cold response pathway in configuring the low-temperature metabolome of Arabidopsis. Proc Natl Acad Sci U S A 101:15243–15248
Cox SE, Stushnoff C (2001) Temperature-related shifts in soluble carbohydrate content during dormancy and cold acclimation in Populus tremuloides. Can J For Res 31:730–737
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163
Cuevas JC, López-Cobollo R, Alcázar R, Zarza X, Koncz C, Altabella T, Salinas J, Tiburcio AF, Ferrando A (2008) Putrescine is involved in Arabidopsis freezing tolerance and cold acclimation by regulating abscisic acid levels in response to low temperature. Plant Physiol 148:1094–1105
Cui S, Huang F, Wang J, Ma X, Cheng Y, Liu J (2005) A proteomic analysis of cold stress responses in rice seedlings. Proteomics 5:3162–3172
Danyluk J, Carpentier E, Sarhan F (1996) Identification and characterization of a low temperature regulated gene encoding an actin-binding protein from wheat. FEBS Lett 389:324–327
Das R, Pandey GK (2010) Expressional analysis and role of calcium regulated kinases in abiotic stress signaling. Curr Genomics 11(1):2–13
Dat J, Vandenabeele S, Vranova E, Van Montagu M, Inze D, Van Breusegem F (2000) Dual action of the active oxygen species during plant stress responses. Cell Mol Life Sci 57:779–795
Davletova S, Schlauch K, Coutu J, Mittler R (2005) The zinc-finger protein Zat12 plays a central role in reactive oxygen and abiotic stress signaling in Arabidopsis. Plant Physiol 139:847–856
Debnath M, Pandey M, Bisen PS (2011) An omics approach to understand the plant abiotic stress. Omics J 15:739–762
Degand H, Faber AM, Dauchot N, Mingeot D, Watillon B, Cutsem PV, Morsomme P, Boutry M (2009) Proteomic analysis of chicory root identifies proteins typically involved in cold acclimation. Proteomics 9:2903–2907
Desimone M, Henke A, Wagner E (1996) Oxidative stress induces partial degradation of the large subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase in isolated chloroplasts of barley. Plant Physiol 111:789–796
Dinari A, Niazi A, Afsharifar AR, Ramezani A (2013) Identification of upregulated genes under cold stress in cold-tolerant chickpea using the cDNA-AFLP approach. PLoS One 8:e52757
Doczi R, Brader G, Pettko-Szandtner A, Rajh I, Djamei A, Pitzschke A, Teige M, Hirt H (2007) The Arabidopsis mitogen-activated protein kinase kinase MKK3 is upstream of group C mitogen-activated protein kinases and participates in pathogen signaling. Plant Cell 19:3266–3279
Doherty CJ, Van Buskirk HA, Myers SJ, Thomashow MF (2009) Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21:972–984
Dong CH, Hu X, Tang W, Zheng X, Kim YS, Lee BH, Zhu JK (2006) A putative Arabidopsis nucleoporin, AtNUP160, is critical for RNA export and required for plant tolerance to cold stress. Mol Cell Biol 26:9533–9543
Dong C, Zhang Z, Ren J, Qin Y, Huang J, Wang Y, Cai B, Wang B, Tao J (2013) Stress-responsive gene ICE1 from Vitis amurensis increases cold tolerance in tobacco. Plant Physiol Biochem 71:212–217
dos Santos R, Vergauwen R, Pacolet P, Lescrinier E, Van den Ende W (2012) Manninotriose is a major carbohydrate in red deadnettle (Lamium purpureum, Lamiaceae). Ann Bot 111(3):385–393
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Dumont E, Bahrman N, Goulas E, Valot B, Sellier H, Hilbert JL, Vuylsteker C, Lejeune-Henaut I, Delbreil B (2011) A proteomic approach to decipher chilling response from cold acclimation in pea (Pisum sativum L.). Plant Sci 180:86–98
Egert A, Keller F, Peters S (2013) Abiotic stress-induced accumulation of raffinose in Arabidopsis leaves is mediated by a single raffinose synthase (RS5, At5g40390). BMC Plant Biol 13:218. doi:10.1186/1471-2229-13-218
Espartero J, Sanchez-Aguayo I, Pardo JM (1995) Molecular characterization of glyoxalase-I from a higher plant; upregulation by stress. Plant Mol Biol 29:1223–1233
Feng XM, Zhao Q, Zhao LL, Qiao Y, Xie XB, Li HF, Yao YX, You CX, Hao YJ (2012) The cold-induced basic helix-loop-helix transcription factor gene MdCIbHLH1 encodes an ICE-like protein in apple. BMC Plant Biol 12:22
Feng HL, Ma NN, Meng S, Zhang S, Wang JR, Chai S, Meng QW (2013) A novel tomato MYC-type ICE1-like transcription factor, SlICE1a, confers cold, osmotic and salt tolerance in transgenic tobacco. Plant Physiol Biochem 73:309–320
Fernandez P, Di Rienzo J, Fernandez L, Hopp HE, Paniego N, Heinz RA (2008) Transcriptomic identification of candidate genes involved in sunflower responses to chilling and salt stresses based on cDNA microarray analysis. BMC Plant Biol 8:11
Fournier ML, Gilmore JM, Martin-Brown SA, Washburn MP (2007) Multidimensional separations-based shotgun proteomics. Chem Rev 107:3654–3686
Fowler S, Thomashow MF (2002) Arabidopsis transcriptome profiling indicates that multiple regulatory pathways are activated during cold acclimation in addition to the CBF cold response pathway. Plant Cell 14:1675–1690
French AD, Waterhouse AL (1993) Chemical structure and characherisitc. In: Suzuki M, Chatterton NJ (eds) Science and technology of fructans. CRC Press, Florida, pp 41–81
Fursova OV, Pogorelko GV, Tarasov VA (2009) Identification of ICE2, a gene involved in cold acclimation which determines freezing tolerance in Arabidopsis thaliana. Gene 429:98–103
Ganeshan S, Vitamvas P, Fowler DB, Chibbar RN (2008) Quantitative expression analysis of selected COR genes reveals their differential expression in leaf and crown tissues of wheat (Triticum aestivum L.) during an extended low temperature acclimation regimen. J Exp Bot 59:2393–2402
Gao MJ, Allard G, Byass L, Flanagan AM, Singh J (2002) Regulation and characterization of four CBF transcription factors from Brassica napus. Plant Mol Biol 49(5):459–471
Gao F, Zhou Y, Zhu W, Li X, Fan L, Zhang G (2009) Proteomic analysis of cold stress-responsive proteins in Thellungiella rosette leaves. Planta 230:1033–1046
Gerke V, Moss SE (2002) Annexins: from structure to function. Physiol Rev 82:331–371
Ghangal R, Raghuvanshi S, Sharma PC (2012) Expressed sequence tag based identification and expression analysis of some cold inducible elements in seabuckthorn (Hippophae rhamnoides L.). Plant Physiol Biochem 51:123–128
Gilmour SJ, Zarka DG, Stockinger EJ, Salazar MP, Houghton JM, Thomashow MF (1998) Low temperature regulation of the Arabidopsis CBF family of AP2 transcriptional activators as an early step in cold-induced COR gene expression. Plant J 16:433–442
Gilmour SJ, Sebolt AM, Salazar MP, Everard JD, Thomashow MF (2000) Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol 124:1854–1865
Gilmour SJ, Fowler SG, Thomashow MF (2004) Arabidopsis transcriptional activators CBF1, CBF2, and CBF3 have matching functional activities. Plant Mol Biol 54:767–781
Gong Z, Lee H, Xiong L, Jagendorf A, Stevenson B, Zhu JK (2002) RNA helicase-like protein as an early regulator of transcription factors for plant chilling and freezing tolerance. Proc Natl Acad Sci U S A 99:11507–11512
Gong Z, Dong CH, Lee H, Zhu J, Xiong L, Gong D, Stevenson B, Zhu JK (2005) A DEAD box RNA helicase is essential for mRNA export and important for development and stress responses in Arabidopsis. Plant Cell 17:256–267
Goulas E, Schubert M, Kieselbach T, Kleczkowski LA, Gardeström P, Schröder W, Hurry V (2006) The chloroplast lumen and stromal proteomes of Arabidopsis thaliana show differential sensitivity to short- and long-term exposure to low temperature. Plant J 47:720–734
Griffith M, Yaish MW (2004) Antifreeze proteins in overwintering plants: a tale of two activities. Trends Plant Sci 9:399–405
Griffith M, Antikainen M, Hon WC, Pihakaski-Maunsbach K, Yu XM, Chun JU, Yang DSC (1997) Antifreeze proteins in winter rye. Physiol Plant 100:327–332
Griffith M, Lumb C, Wiseman SB, Wisniewski M, Johnson RW, Marangoni AG (2005) Antifreeze proteins modify the freezing process in planta. Plant Physiol 138:330–340
Gu X, Gao Z, Zhuang W, Qiao Y, Wang X, Mi L, Zhang Z, Lin Z (2013) Comparative proteomic analysis of rd29A:RdreB1BI transgenic and non-transgenic strawberries exposed to low temperature. J Plant Physiol 170:696–706
Guan Q, Wu J, Zhang Y, Jiang C, Liu R, Chai C, Zhu J (2013) A DEAD box RNA helicase is critical for pre-mRNA splicing, cold-responsive gene regulation, and cold tolerance in Arabidopsis. Plant Cell 25(1):342–356
Guerra D, Mastrangelo AM, Lopez-Torrejon G, Marzin S, Schweizer P, Stanca AM, del Pozo JC, Cattivelli L, Mazzucotelli E (2012) Identification of a protein network interacting with TdRF1, a wheat RING ubiquitin ligase with a protective role against cellular dehydration. Plant Physiol 158:777–789
Gulick PJ, Drouin S, Yu Z, Danyluk J, Poisson G, Monroy AF, Sarhan F (2005) Transcriptome comparison of winter and spring wheat responding to low temperature. Genome 48:913–923
Gusta LV, Wisniewski M, Nesbitt NT, Gusta ML (2004) The effect of water, sugars, and proteins on the pattern of ice nucleation and propagation in acclimated and nonacclimated canola leaves. Plant Physiol 135:1642–1653
Gutha LR, Reddy AR (2008) Rice DREB1B promoter shows distinct stress-specific responses, and the overexpression of cDNA in tobacco confers improved abiotic and biotic stress tolerance. Plant Mol Biol 68:533–555
Halliwell B (2007) Biochemistry of oxidative stress. Biochem Soc Trans 35:1147–1150
Hansen J, Vogg G, Beck E (1996) Assimilation, allocation and utilization of carbon by 3-year-old Scots pine (Pinus sylvestris L.) tree during winter and early spring. Trees 11:83–90
Hare PD, Cress WA, Van Staden J (1998) Dissecting the roles of osmolyte accumulation during stress. Plant Cell Environ 21:535–553
Hashimoto M, Komatsu S (2007) Proteomic analysis of rice seedlings during cold stress. Proteomics 7:1293–1302
Hashimoto M, Toorchi M, Matsushita K, Iwasaki Y, Komatsu S (2009) Proteome analysis of rice root plasma membrane and detection of cold stress responsive proteins. Protein Peptide Lett 16:685–697
Hausman JF, Evers D, Thiellement H, Jouve L (2000) Compared responses of poplar cuttings and in vitro raised shoots to short-term chilling treatments. Plant Cell Rep 19:954–960
Heidarvand L, Maali Amiri R (2010) What happens in plant molecular responses to cold stress? Acta Physiol Plant 32:419–431
Heidarvand L, Maali-Amiri R (2013) Physio-biochemical and proteome analysis of chickpea in early phases of cold stress. J Plant Physiol 170:459–469
Henriksson NK, Trewavas AJ (2003) The effect of short-term low-temperature treatments on gene expression in Arabidopsis correlates with changes in intracellular Ca2+ levels. Plant Cell Environ 26:485–496
Hoffmann-Sommergruber K (2000) Plant allergens and pathogenesis-related proteins. What do they have in common? Int Arch Allergy Immunol 122:155–166
Hong Z, Lakkineni K, Zhang Z, Verma DP (2000) Removal of feedback inhibition of delta(1)-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol 122:1129–1136
Houde M, Dallaire S, N’Dong D, Sarhan F (2004) Overexpression of the acidic dehydrin WCOR410 improves freezing tolerance in transgenic strawberry leaves. Plant Biotechnol J 2:381–387
Hsieh TH, Lee JT, Yang PT, Chiu LH, Charng YY, Wang YC, Chan MT (2002) Heterology expression of the Arabidopsis C-repeat/dehydration response element binding factor 1 gene confers elevated tolerance to chilling and oxidative stresses in transgenic tomato. Plant Physiol 129:1086–1094
Hu Y, Zhang L, Zhao L, Li J, He S (2011) Trichostatin a selectively suppresses the cold-induced transcription of the ZmDREB1 gene in maize. PLoS One 6(7):e22132
Huang C, Ding S, Zhang H, Du H, An L (2011) CIPK7 is involved in cold response by interacting with CBL1 in Arabidopsis thaliana. Plant Sci 181:57–64
Huang XS, Wang W, Zhang Q, Liu JH (2013) A basic helix-loop-helix transcription factor, PtrbHLH, of Poncirus trifoliata confers cold tolerance and modulates peroxidase-mediated scavenging of hydrogen peroxide. Plant Physiol 162:1178–1194
Hugly S, Somerville C (1992) A role for membrane lipid polyunsaturation in chloroplast biogenesis at low temperature. Plant Physiol 99:197–202
Hung SH, Yu CW, Lin CH (2005) Hydrogen peroxide functions as a stress signal in plants. Bot Bull Acad Sin 46:1–10
Hurry V, Strand A, Furbank R, Stitt M (2000) The role of inorganic phosphate in the development of freezing tolerance and the acclimatization of photosynthesis to low temperature is revealed by the pho mutants of Arabidopsis thaliana. Plant J 24:383–396
Ichimura K, Mizoguchi T, Yoshida R, Yuasa T, Shinozaki K (2000) Various abiotic stresses rapidly activate Arabidopsis MAP kinases ATMPK4 and ATMPK6. Plant J 24:655–665
Imin N, Kerim T, Rolfe BG, Weinman JJ (2004) Effect of early cold stress on the maturation of rice anthers. Proteomics 4:1873–1882
Ishikawa M, Yoshida S (1985) Seasonal changes in plasma membranes and mitochondria isolated from jerusalem artichoke tubers. Possible relationship to cold hardiness. Plant Cell Physiol 26:1331–1344
Ishitani M, Xiong L, Lee H, Stevenson B, Zhu JK (1998) HOS1, a genetic locus involved in cold-responsive gene expression in arabidopsis. Plant Cell 10:1151–1161
Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol 47:141–153
Jaglo KR, Kleff S, Amundsen KL, Zhang X, Haake V, Zhang JZ, Deits T, Thomashow MF (2001) Components of the Arabidopsis C-repeat/dehydration-responsive element binding factor cold-response pathway are conserved in Brassica napus and other plant species. Plant Physiol 127:910–917
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
Janmohammadi M (2012) Metabolomic analysis of low temperature responses in plants. Curr Opin Agric 1:1–6
Janska A, Marsik P, Zelenkova S, Ovesna J (2010) Cold stress and acclimation—what is important for metabolic adjustment? Plant Biol 12:395–405
Jiang C, Iu B, Singh J (1996) Requirement of a CCGAC cis-acting element for cold induction of the BN115 gene from winter Brassica napus. Plant Mol Biol 30:679–684
Jiang QW, Kiyoharu O, Ryozo I (2002) Two novel mitogen-activated protein signaling components, OsMEK1 and OsMAP1, are involved in a moderate low-temperature signaling pathway in rice. Plant Physiol 129:1880–1891
Jonak C, Kiegerl S, Ligterink W, Barker PJ, Huskisson NS, Hirt H (1996) Stress signaling in plants: a mitogen-activated protein kinase pathway is activated by cold and drought. Proc Natl Acad Sci U S A 93:11274–11279
Jouve L, Hoffmann L, Hausman J-F (2004) Polyamine, carbohydrate, and proline content changes during salt stress exposure of aspen (Populus tremula L.): involvement of oxidation and osmoregulation metabolism. Plant Biol (Stuttg) 6:74–80
Jung SH, Lee JY, Lee DH (2003) Use of SAGE technology to reveal changes in gene expression in Arabidopsis leaves undergoing cold stress. Plant Mol Biol 52:553–567
Jung JH, Seo PJ, Park CM (2012) The E3 ubiquitin ligase HOS1 regulates Arabidopsis flowering by mediating CONSTANS degradation under cold stress. J Biol Chem 287:43277–43287
Jung JH, Park JH, Lee S, To TK, Kim JM, Seki M, Park CM (2013) The cold signaling attenuator HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENE1 activates FLOWERING LOCUS C transcription via chromatin remodeling under short-term cold stress in Arabidopsis. Plant Cell 25:4378–4390
Kaplan F, Kopka J, Haskell DW, Zhao W, Schiller KC, Gatzke N, Sung DY, Guy CL (2004) Exploring the temperature-stress metabolome. Plant Physiol 136:4159–4168
Kaplan F, Kopka J, Sung DY, Zhao W, Popp M, Porat R, Guy CL (2007) Transcript and metabolite profiling during cold acclimation of Arabidopsis reveals an intricate relationship of cold-regulated gene expression with modifications in metabolite content. Plant J 50:967–981
Kasamo K (1988) Inhibition of tonoplast and plasma membrane H+-ATPase activity in rice (Oryza sativa L.) culture cells by local anesthetics. Plant Cell Physiol 29:215–221
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17(3):287–291
Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2004) A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low-temperature stress tolerance in tobacco by gene transfer. Plant Cell Physiol 45:346–350
Kasukabe Y, He L, Nada K, Misawa S, Ihara I, Tachibana S (2004) Overexpression of spermidine synthase enhances tolerance to multiple environmental stresses and up-regulates the expression of various stress-regulated genes in transgenic Arabidopsis thaliana. Plant Cell Physiol 45:712–722
Kaul S, Sharma SS, Mehta IK (2008) Free radical scavenging potential of L-proline: evidence from in vitro assays. Amino Acids 34:315–320
Kaur G, Kumar S, Thakur P, Malik JA, Bhandhari K, Sharma KD, Nayyar H (2011) Involvement of proline in response of chickpea (Cicer arietinum L.) to chilling stress at reproductive stage. Sci Hortic (Amsterdam) 128:174–181
Kawamura Y, Uemura M (2003) Mass spectrometric approach for identifying putative plasma membrane proteins of Arabidopsis leaves associated with cold acclimation. Plant J 36:141–154
Kim JC, Lee SH, Cheong YH, Yoo CM, Lee SI, Chun HJ, Yun DJ, Hong JC, Lee SY, Lim CO, Cho MJ (2001) A novel cold-inducible zinc finger protein from soybean, SCOF-1, enhances cold tolerance in transgenic plants. Plant J 25(3):247–259
Kim TE, Kim S-K, Han TJ, Lee JS, Chang SC (2002) ABA and polyamines act independently in primary leaves of cold-stressed tomato (Lycopersicon esculentum). Physiol Plant 115:370–376
Kim KN, Cheong YH, Grant JJ, Pandey GK, Luan S (2003) CIPK3, a calcium sensor-associated protein kinase that regulates abscisic acid and cold signal transduction in Arabidopsis. Plant Cell 15:411–423
Kim MC, Chung WS, Yun DJ, Cho MJ (2009) Calcium and calmodulin-mediated regulation of gene expression in plants. Mol Plant 2:13–21
Kitashiba H, Ishizaka T, Isuzugawa K, Nishimura K, Suzuki T (2004) Expression of a sweet cherry DREB1/CBF ortholog in Arabidopsis confers salt and freezing tolerance. J Plant Physiol 161:1171–1176
Knight H, Knight MR (2001) Abiotic stress signalling pathways: specificity and cross-talk. Trends Plant Sci 6:262–267
Knight MR, Campbell AK, Smith SM, Trewavas AJ (1991) Transgenic plant aequorin reports the effects of touch and cold-shock and elicitors on cytoplasmic calcium. Nature 352:524–526
Knight H, Trewavas AJ, Knight MR (1996) Cold calcium signaling in Arabidopsis involves two cellular pools and a change in calcium signature after acclimation. Plant Cell 8:489–503
Knight H, Veale EL, Warren GJ, Knight MR (1999) The sfr6 mutation in Arabidopsis suppresses low-temperature induction of genes dependent on the CRT/DRE sequence motif. Plant Cell 11:875–886
Kodama H, Hamada T, Horiguchi G, Nishimura M, Iba K (1994) Genetic enhancement of cold tolerance by expression of a gene for chloroplast v-3 fatty acid desaturase in transgenic tobacco. Plant Physiol 105:601–605
Koiwa H, Hausmann S, Bang WY, Ueda A, Kondo N, Hiraguri A, Fukuhara T, Bahk JD, Yun DJ, Bressan RA, Hasegawa PM, Shuman S (2004) Arabidopsis C-terminal domain phosphatase-like 1 and 2 are essential Ser-5-specific C-terminal domain phosphatases. Proc Natl Acad Sci U S A 101:14539–14544
Kolbe A, Tiessen A, Schluepmann H, Paul M, Ulrich S, Gelgenberger P (2005) Trehalose 6-phosphate regulates starch synthesis via posttranslational redox activation of ADP-glucose pyrophosphorylase. Proc Natl Acad Sci U S A 102:11118–11123
Komatsu S, Yang G, Khan M, Onodera H, Toki S, Yamaguchi M (2007) Over-expression of calcium dependent protein kinase 13 and calreticulin interacting protein 1 confers cold tolerance on rice plants. Mol Genet Genomics 277:713–723
Korn M, Peterek S, Mock HP, Heyer AG, Hincha DK (2008) Heterosis in the freezing tolerance, and sugar and flavonoid contents of crosses between Arabidopsis thaliana accessions of widely varying freezing tolerance. Plant Cell Environ 31(6):813–827
Korn M, Gartner T, Erban A, Kopka J, Selbig J, Hincha DK (2010) Predicting Arabidopsis freezing tolerance and heterosis in freezing tolerance from metabolite composition. Mol Plant 3(1):224–235
Kosová K, Vítámvás P, Prášil IT (2007) The role of dehydrins in plant response to cold. Biol Plant 51:601–617
Kosova K, Vitamvas P, Prasil IT, Renaut J (2011) Plant proteome changes under abiotic stress–contribution of proteomics studies to understanding plant stress response. J Proteomics 74:1301–1322
Kovacs D, Kalmar E, Torok Z, Tompa P (2008) Chaperone activity of ERD10 and ERD14, two disordered stress-related plant proteins. Plant Physiol 147:381–390
Kovtun Y, Chiu WL, Tena G, Sheen J (2000) Functional analysis of oxidative stress-activated mitogen-activated protein kinase cascade in plants. Proc Natl Acad Sci U S A 97:2940–2945
Krause E, Dathe M, Wieprecht T, Bienert M (1999) Noncovalent immobilized artificial membrane chromatography, an improved method for describing peptide-lipid bilayer interactions. J Chromatogr A 849:125–133
Kubis J (2003) Polyamines and “scavenging system”: influence of exogenous spermidine on catalase and guaiacol peroxidase activities, and free polyamine level in barley leaves under water deficit. Acta Physiol Plant 25:337–343
Kudla J, Xu Q, Harter K, Gruissem W, Luan S (1999) Genes for calcineurin B-like proteins in Arabidopsis are differentially regulated by stress signals. Proc Natl Acad Sci U S A 96:4718–4723
Laudencia-Chingcuanco D, Ganeshan S, You F, Fowler B, Chibbar R, Anderson O (2011) Genome-wide gene expression analysis supports a developmental model of low temperature tolerance gene regulation in wheat (Triticum aestivum L.). BMC Genomics 12:299
Lazaro A, Valverde F, Pineiro M, Jarillo JA (2012) The Arabidopsis E3 ubiquitin ligase HOS1 negatively regulates CONSTANS abundance in the photoperiodic control of flowering. Plant Cell 24:982–999
Lee JY, Lee DH (2003) Use of serial analysis of gene expression technology to reveal changes in gene expression in Arabidopsis pollen undergoing cold stress. Plant Physiol 132:517–529
Lee H, Xiong L, Gong Z, Ishitani M, Stevenson B, Zhu JK (2001) The Arabidopsis HOS1 gene negatively regulates cold signal transduction and encodes a RING finger protein that displays cold-regulated nucleo–cytoplasmic partitioning. Genes Dev 15:912–924
Lee H, Guo Y, Ohta M, Xiong L, Stevenson B, Zhu JK (2002a) LOS2, a genetic locus required for cold-responsive gene transcription encodes a bi-functional enolase. EMBO J 21:2692–2702
Lee BH, Lee H, Xiong L, Zhu JK (2002b) A mitochondrial complex I defect impairs cold-regulated nuclear gene expression. Plant Cell 14:1235–1251
Lee SC, Lee MY, Kim SJ, Jun SH, An G, Kim SR (2005a) Characterization of an abiotic stress-inducible dehydrin gene, OsDhn1, in rice (Oryza sativa L.). Mol Cells 19:212–218
Lee BH, Henderson DA, Zhu JK (2005b) The Arabidopsis cold-responsive transcriptome and its regulation by ICE1. Plant Cell 17:3155–3175
Lee D-G, Ahsan N, Lee S-H, Lee J, Bahk JD, Kang KY, Lee BH (2009) Chilling stress-induced proteomic changes in rice roots. J Plant Physiol 166:1–11
Lee YP, Babakov A, de Boer B, Zuther E, Hincha DK (2012a) Comparison of freezing tolerance, compatible solutes and polyamines in geographically diverse collections of Thellungiella sp. and Arabidopsis thaliana accessions. BMC Plant Biol 12:131
Lee JH, Kim JJ, Kim SH, Cho HJ, Kim J, Ahn JH (2012b) The E3 ubiquitin ligase HOS1 regulates low ambient temperature-responsive flowering in Arabidopsis thaliana. Plant Cell Physiol 53:1802–1814
Lee JH, Kim SH, Kim JJ, Ahn JH (2012c) Alternative splicing and expression analysis of High expression of osmotically responsive genes1 (HOS1) in Arabidopsis. BMB Rep 45:515–520
Levitt J (1980) Responses of plants to environmental stress, vol 1, 2nd edn. Academic, New York, pp 166–222
Li W, Li M, Zhang W, Welti R, Wang X (2004) The plasma membrane-bound phospholipase Ddelta enhances freezing tolerance in Arabidopsis thaliana. Nat Biotechnol 22:427–433
Li J, Wang L, Zhu W, Wang N, Xin H, Li S (2014) Characterization of two VvICE1 genes isolated from ‘Muscat Hamburg’ grapevine and their effect on the tolerance to abiotic stresses. Sci Hortic 165:266–273
Lin Y, Zheng H, Zhang Q, Liu C, Zhang Z (2014) Functional profiling of EcaICE1 transcription factor gene from Eucalyptus camaldulensis involved in cold response in tobacco plants. J Plant Biochem Biotechnol 23:141–150
Lister R, Gregory BD, Ecker JR (2009) Next is now: new technologies for sequencing of genomes, transcriptomes, and beyond. Curr Opin Plant Biol 12:107–118
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA-binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression in Arabidopsis. Plant Cell 10:1391–1406
Liu JJ, Ekramoddoullah AKM, Yu X (2003) Differential expression of multiple PR10 proteins in western white pine following wounding, fungal infection and cold-hardening. Physiol Plant 119:544–553
Liu L, Duan L, Zhang J, Zhang Z, Mi G, Ren H (2010) Cucumber (Cucumis sativus L.) over-expressing cold-induced transcriptome regulator ICE1 exhibits changed morphological characters and enhances chilling tolerance. Sci Hortic 124:29–33
Liu MQ, Shi J, Lu CF (2013) Identification of stress-responsive genes in Ammopiptanthus mongolicus using ESTs generated from cold- and drought-stressed seedlings. BMC Plant Biol 13
Livingston DP, Hincha DK, Heyer AG (2009) Fructan and its relationship to abiotic stress tolerance in plants. Cell Mol Life Sci 66:2007–2023
Lopez-Matas MA, Nunez P, Soto A, Allona I, Casado R, Collada C, Guevara MA, Aragoncillo C, Gomez L (2004) Protein cryoprotective activity of a cytosolic small heat shock protein that accumulates constitutively in chestnut stems and is up-regulated by low and high temperatures. Plant Physiol 134:1708–1717
Luan S, Kudla J, Rodriguez-Concepcion M, Yalovsky S, Gruissem W (2002) Calmodulins and calcineurin B-like proteins: calcium sensors for specific signal response coupling in plants. Plant Cell 14(Suppl):S389–S400
MacGregor DR, Gould P, Foreman J, Griffiths J, Bird S, Page R, Stewart K, Steel G, Young J, Paszkiewicz K, Millar AJ, Halliday KJ, Hall AJ, Penfield S (2013) High expression of osmotically responsive genes1 is required for circadian periodicity through the promotion of nucleo-cytoplasmic mRNA export in Arabidopsis. Plant Cell 25:4391–4404
Maruyama K, Sakuma Y, Kasuga M, Ito Y, Seki M, Goda H, Shimada Y, Yoshida S, Shinozaki K, Yamaguchi-Shinozaki K (2004) Identification of cold-inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J 38:982–993
Maruyama K, Takeda M, Kidokoro S, Yamada K, Sakuma Y, Urano K, Fujita M, Yoshiwara K, Matsukura S, Morishita Y et al (2009) Metabolic pathways involved in cold acclimation identified by integrated analysis of metabolites and transcripts regulated by DREB1A and DREB2A. Plant Physiol 150(4):1972–1980
Maul P, McCollum GT, Popp M, Guy CL, Porat R (2008) Transcriptome profiling of grapefruit flavedo following exposure to low temperature and conditioning treatments uncovers principal molecular components involved in chilling tolerance and susceptibility. Plant Cell Environ 31:752–768
Mauro S, Dainese P, Lannoye R, Bassi R (1997) Cold-resistant and cold-sensitive maize lines differ in the phosphorylation of the photosystem II subunit, CP29. Plant Physiol 115:171–180
McAinsh MR, Pittman JK (2009) Shaping the calcium signature. New Phytol 181:275–294
Medina J, Bargues M, Terol J, Perez-Alonso M, Salinas J (1999) The Arabidopsis CBF gene family is composed of three genes encoding AP2 domain-containing proteins whose expression is regulated by low temperature but not by abscisic acid or dehydration. Plant Physiol 119:463–470
Miquel M, James D, Dooner H, Browse J (1993) Arabidopsis requires polyunsaturated lipids for low-temperature survival. Proc Natl Acad Sci U S A 90:6208–6212
Mittal D, Madhyastha DA, Grover A (2012) Genome-wide transcriptional profiles during temperature and oxidative stress reveals coordinated expression patterns and overlapping regulons in rice. PLoS One 7:e40899
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Miura K, Jin JB, Hasegawa PM (2007) Sumoylation, a post-translational regulatory process in plants. Curr Opin Plant Biol 10:495–502
Miura K, Ohta M, Nakazawa M, Ono M, Hasegawa PM (2011) ICE1 Ser403 is necessary for protein stabilization and regulation of cold signaling and tolerance. Plant J 67:269–279
Miura K, Okamoto H, Okuma E, Shiba H, Kamada H, Hasegawa PM, Murata Y (2012) SIZ1 deficiency causes reduced stomatal aperture and enhanced drought tolerance via controlling salicylic acid-induced accumulation of reactive oxygen species in Arabidopsis. Plant J doi: 10.1111/tpj.12014
Monroy AF, Dhindsa RS (1995) Low temperature signal transduction: induction of cold acclimation-specific genes of alfalfa by calcium at 25 °C. Plant Cell 7:321–331
Monroy AF, Sarhan F, Dhindsa RS (1993) Cold-induced changes in freezing tolerance, protein phosphorylation, and gene expression (evidence for a role of calcium). Plant Physiol 102:1227–1235
Monroy AF, Sangwan V, Dhindsa RS (1998) Low temperature signal transduction during cold acclimation: protein phosphatase 2A as an early target for cold-inactivation. Plant J 13:653–660
Morran S, Eini O, Pyvovarenko T, Parent B, Singh R, Ismagul A, Eliby S, Shirley N, Langridge P, Lopato S (2011) Improvement of stress tolerance of wheat and barley by modulation of expression of DREB/CBF factors. Plant Biotechnol J 9(2):230–249
Morsy MR, Jouve L, Hausman J-F, Hoffmann L, Stewart JM (2007) Alteration of oxidative and carbohydrate metabolism under abiotic stress in two rice (Oryza sativa L.) genotypes contrasting in chilling tolerance. J Plant Physiol 164:157–167
Murata N, Ishizaki-Nishizawa O, Higashi S, Hayashi H, Tasaka Y, Nishida I (1992) Genetically engineered alteration in the chilling sensitivity of plants. Nature 356:710–713
Murchie EH, Sarrobert C, Contard P, Betsche T, Foyer CH, Galtier N (1999) Overexpresion of sucrose-phosphate synthase in tomato plants grown with Co2 enrichment leads to decreased foliar carbohydrate accumulation relative to transfromed controls. Plant Physiol Biochem 37(4):251–260
Nadimpalli R, Yalpani N, Johal GS, Simmons CR (2000) Prohibitins, stomatins, and plant disease response genes compose a protein superfamily that controls cell proliferation, ion channel regulation, and death. J Biol Chem 275:29579–29586
Nakagami H, Soukupova H, Schikora A, Zarsky V, Hirt H (2006) A mitogen-activated protein kinase kinase kinase mediates reactive oxygen species homeostasis in Arabidopsis. J Biol Chem 281:38697–38704
Nakamura J, Yuasa T, Thi HT, Harano K, Tanaka S, Iwata T, Phan T, Mari II (2011) Rice homologs of inducer of CBF expression (OsICE) are involved in cold acclimation. Plant Biotechnol 28:303–309
Nakayama K, Okawa K, Kakizaki T, Inaba T (2008) Evaluation of the protective activities of a late embryogenesis abundant (LEA) related protein, Cor15am, during various stresses in vitro. Biosci Biotechnol Biochem 72:1642–1645
Neill SJ, Desikan R, Clarke A, Hurst RD, Hancock JT (2002) Hydrogen peroxide and nitric oxide as signalling molecules in plants. J Exp Bot 53(372):1237–1247
Neill S, Barros R, Bright J, Desikan R, Hancock J, Harrison J, Morris P, Riberiro D, Wilson I (2008) Nitric oxide, stomatal closure, and abiotic stress. J Exp Bot 59:165–176
Neilson KA, Mariani M, Haynes PA (2011) Quantitative proteomic analysis of cold-responsive proteins in rice. Proteomics 11:1696–1706
Nick P (2000) Control of the response to low temperatures. In: Nick P (ed) Plant microtubules: potential for biotechnology. Springer, Berlin, pp 121–135
Novillo F, Alonso JM, Ecker JR, Salinas J (2004) CBF2/DREB1C is a negative regulator of CBF1/DREB1B and CBF3/DREB1A expression and plays a central role in stress tolerance in Arabidopsis. Proc Natl Acad Sci U S A 101:3985–3990
Novillo F, Medina J, Salinas J (2007) Arabidopsis CBF1 and CBF3 have a different function than CBF2 in cold acclimation and define different gene classes in the CBF regulon. Proc Natl Acad Sci U S A 104:21002–21007
Nylander M, Svensson J, Palva ET, Welin BV (2001) Stress-induced accumulation and tissue-specific localization of dehydrins in Arabidopsis thaliana. Plant Mol Biol 45:263–279
O’Kane D, Gill V, Boyd P, Burdon R (1996) Chilling, oxidative stress and antioxidant responses in Arabidopsis thaliana callus. Planta 198:371–377
Oakenfull RJ, Baxter R, Knight MR (2013) A C-repeat binding factor transcriptional activator (CBF/DREB1) from European bilberry (Vaccinium myrtillus) induces freezing tolerance when expressed in Arabidopsis thaliana. PLoS One 8:e54119
Oh SJ, Song SI, Kim YS, Jang HJ, Kim SY, Kim M, Kim YK, Nahm BH, Kim JK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138:341–351
Oh SJ, Kwon CW, Choi DW, Song SI, Kim JK (2007) Expression of barley HvCBF4 enhances tolerance to abiotic stress in transgenic rice. Plant Biotechnol J 5:646–656
Ohno R, Takumi S, Nakamura C (2003) Kinetics of transcript and protein accumulation of a low-molecular-weight wheat LEA D-11 dehydrin in response to low temperature. J Plant Physiol 160:193–200
Oliveira G, Peñuelas J (2004) Effects of winter cold stress on photosynthesis and photochemical efficiency of PSII of the Mediterranean Cistus albidus L. and Quercus ilex L. Plant Eco 175:179–191
Ortiz-Masia D, Perez-Amador MA, Carbonell J, Marcote MJ (2007) Diverse stress signals activate the C1 subgroup MAP kinases of Arabidopsis. FEBS Lett 581:1834–1840
Orvar BL, Sangwan V, Omann F, Dhindsa RS (2000) Early steps in cold sensing by plant cells: the role of actin cytoskeleton and membrane fluidity. Plant J 23:785–794
Ouellet F (2007) Cold acclimation and freezing tolerance in plants. Encyclopaedia of Life Sciences. 1–6
Ouellet F, Vazquez-Tello A, Sarhan F (1998) The wheat wcs120 promoter is cold-inducible in both monocotyledonous and dicotyledonous species. FEBS Lett 423:324–328
Owens CL, Thomashow MF, Hancock JF, Iezzoni AF (2002) CBF1 orthologs in sour cherry and strawberry and the heterologous expression of CBF1 in strawberry. J Am Soc Hortic Sci 127:489–494
Pandey GK (2008) Emergence of a novel calcium signaling pathway in plants: CBL-CIPK signaling network. Physiol Mol Biol Plants 14:51–68
Pang T, Ye CY, Xia XL, Yin WL (2013) De novo sequencing and transcriptome analysis of the desert shrub, Ammopiptanthus mongolicus, during cold acclimation using Illumina/Solexa. BMC Genomics 14:488
Pasquali G, Biricolti S, Locatelli F, Baldoni E, Mattana M (2008) Osmyb4 expression improves adaptive responses to drought and cold stress in transgenic apples. Plant Cell Rep 27:1677–1686
Paul MJ (2008) Trehalose 6-phosphate: a signal of sucrose status. Biochem J 412:e1–e2
Pellegrineschi A, Reynolds M, Pacheco M, Brito RM, Almeraya R, Yamaguchi-Shinozaki K, Hoisington D (2004) Stress-induced expression in wheat of the Arabidopsis thaliana DREB1A gene delays water stress symptoms under greenhouse conditions. Genome 47:493–500
Peng T, Zhu XF, Fan QJ, Sun PP, Liu JH (2012) Identification and characterization of low temperature stress responsive genes in Poncirus trifoliata by suppression subtractive hybridization. Gene 492:220–228
Peng PH, Lin CH, Tsai HW, Lin TY (2014) Cold response in phalaenopsis aphrodite and characterization of PaCBF1 and PaICE1. Plant Cell Physiol pii: pcu093. [Epub ahead of print]
Pennycooke JC, Cheng H, Roberts SM, Yang Q, Rhee SY, Stockinger EJ (2008a) The low temperature-responsive, Solanum CBF1 genes maintain high identity in their upstream regions in a genomic environment undergoing gene duplications, deletions, and rearrangements. Plant Mol Biol 67:483–497
Pennycooke JC, Cheng H, Stockinger EJ (2008b) Comparative genomic sequence and expression analyses of Medicago truncatula and alfalfa subspecies falcata COLD-ACCLIMATION-SPECIFIC genes. Plant Physiol 146:1242–1254
Porat R, Pasentsis K, Rozentzvieg D, Gerasopoulos D, Falara V, Samach A, Lurie S, Kanellis AK (2004) Isolation of a dehydrin cDNA from orange and grapefruit citrus fruit that is specifically induced by the combination of heat followed by chilling temperatures. Physiol Plant 120:256–264
Prasad TK, Anderson MD, Martin BA, Stewart CR (1994) Evidence for chilling-induced oxidative stress in Maize seedlings and a regulatory role for hydrogen peroxide. Plant Cell 6:65–74
Qin F, Sakuma Y, Li J, Liu Q, Li YQ, Shinozaki K, Yamaguchi-Shinozaki K (2004) Cloning and functional analysis of a novel DREB1/CBF transcription factor involved in cold-responsive gene expression in Zea mays L. Plant Cell Physiol 45:1042–1052
Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, Ito Y, Yoshiwara K, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133:1755–1767
Ray S, Agarwal P, Arora R, Kapoor S, Tyagi AK (2007) Expression analysis of calcium-dependent protein kinase gene family during reproductive development and abiotic stress conditions in rice (Oryza sativa L. ssp. indica). Mol Genet Genomics 278(5):493–505
Reddy VS, Reddy AS (2004) Proteomics of calcium signaling components in plants. Phytochemistry 65:1745–1776
Rekarte-Cowie I, Ebshish OS, Mohamed KS, Pearce RS (2008) Sucrose helps regulate cold acclimation of Arabidopsis thaliana. J Exp Bot 59:4205–4217
Renaut J, Lutts S, Hoffmann L, Hausman JF (2004) Responses of poplar to chilling temperatures: proteomic and physiological aspects. Plant Biol 6:81–90
Renaut J, Hausman J-F, Wisniewski ME (2006) Proteomics and low-temperature studies: bridging the gap between gene expression and metabolism. Physiol Plant 126:97–109
Renaut J, Hausman JF, Bassett C, Artlip T, Cauchie HM, Witters E, Wisniewski M (2008) Quantitative proteomic analysis of short photoperiod and low-temperature responses in bark tissues of peach (Prunus persica L. Batsch). Tree Genet Genomes 4:589–600
Rinalducci S, Egidi MG, Karimzadeh G, Jazii FR, Zolla L (2011) Proteomic analysis of a spring wheat cultivar in response to prolonged cold stress. Electrophoresis 32:1807–1818
Robinson SJ, Parkin IA (2008) Differential SAGE analysis in Arabidopsis uncovers increased transcriptome complexity in response to low temperature. BMC Genomics 9:434
Rontein D, Basset G, Hanson AD (2002) Metabolic engineering of osmoprotectant accumulation in plants. Metab Eng 4:49–56
Rorat T (2006) Plant dehydrins–tissue location, structure and function. Cell Mol Biol Lett 11:536–556
Ruiz JM, Sánchez E, García PC, Lefebre LRL, Rivero RM, Romero L (2002) Proline metabolism and NAD kinase activity in greenbean plants subjected to cold-shock. Phytochemistry 59:473–478
Sabehat A, Lurie S, Weiss D (1998) Expression of small heat-shock proteins at low temperatures. A possible role in protecting against chilling injuries. Plant Physiol 117:651–658
Sage RF, Kubien DS (2007) The temperature response of C(3) and C(4) photosynthesis. Plant Cell Environ 30:1086–1106
Saijo Y, Kinoshita N, Ishiyama K, Hata S, Kyozuka J, Hayakawa T, Nakamura T, Shimamoto K, Yamaya T, Izui K (2001) A Ca2+ dependent protein kinase that endows rice plants with cold- and salt-stress tolerance functions in vascular bundles. Plant Cell Physiol 42:1228–1233
Salinas J (2002) Molecular mechanisms of signal transduction in cold acclimation. In: Scheel D, Wasternack C (eds) Plant signal transduction. Oxford University Press, Oxford, pp 116–139
Sangwan V, Foulds I, Singh J, Dhindsa RS (2001) Cold-activation of Brassica napus BN115 promoter is mediated by structural changes in membranes and cytoskeleton, and requires Ca2+ influx. Plant J 27:1–12
Sangwan V, Orvar BL, Beyerly J, Hirt H, Dhindsa RS (2002) Opposite changes in membrane fluidity mimic cold and heat stress activation of distinct plant MAP kinase pathways. Plant J 31:629–638
Sathyanarayanan PV, Poovaiah BW (2004) Decoding Ca2+ signals in plants. Crit Rev Plant Sci 23:1–11
Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K, Carninci P, Hayashizaki Y, Shinozaki K (2001) Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13:61–72
Seki M, Ishida J, Narusaka M, Fujita M, Nanjo T, Umezawa T, Kamiya A, Nakajima M, Enju A, Sakurai T et al (2002) Monitoring the expression pattern of around 7,000 Arabidopsis genes under ABA treatments using a full-length cDNA microarray. Funct Integr Genomics 2(6):282–291
Shan W, Kuang JF, Lu WJ, Chen JY (2014) Banana fruit NAC transcription factor MaNAC1 is a direct target of MaICE1 and involved in cold stress through interacting with MaCBF1. Plant Cell Environ 37(9):2116–2127
Sharma P, Sharma N, Deswal R (2005) The molecular biology of the low-temperature response in plants. Bioessays 27:1048–1059
Shen YG, Zhang WK, He SJ, Zhang JS, Liu Q, Chen SY (2003) An EREBP/AP2-type protein in Triticum aestivum was a DRE-binding transcription factor induced by cold, dehydration and ABA stress. Theor Appl Genet 106:923–930
Shi H, Ye T, Zhong B, Liu X, Jin R, Chan Z (2014) AtHAP5A modulates freezing stress resistance in Arabidopsis through binding to CCAAT motif of AtXTH21. New Phytol 203:554–567
Shimamura C, Ohno R, Nakamura C, Takumi S (2006) Improvement of freezing tolerance in tobacco plants expressing a cold-responsive and chloroplast-targeting protein WCOR15 of wheat. J Plant Physiol 163:213–219
Shinozaki K, Yamaguchi-Shinozaki K (1996) Molecular responses to drought and cold stress. Curr Opin Biotechnol 7:161–167
Si J, Wang J-H, Zhang L-J, Zhang H, Liu Y-J, An L-Z (2009) CbCOR15, A cold-regulated gene from alpine chorispora bungeana, confers cold tolerance in transgenic tobacco. J Plant Biol 52:593–601
Siddiqua M, Nassuth A (2011) Vitis CBF1 and Vitis CBF4 differ in their effect on Arabidopsis abiotic stress tolerance, development and gene expression. Plant Cell Environ 34:1345–1359
Signora L, Galtier N, Skøt L, Lucas H, Foyer CH (1998) Over-expression of sucrose phosphate synthase in Arabidopsis thaliana results in increased foliar sucrose/starch ratios and favours decreased foliar carbohydrate accumulation in plants after prolonged growth with CO enrichment. J Exp Bot 49:669–680
Skinner DZ (2009) Post-acclimation transcriptome adjustment is a major factor in freezing tolerance of winter wheat. Funct Integr Genomics 9:513–523
Skinner JS, von Zitzewitz J, Szucs P, Marquez-Cedillo L, Filichkin T, Amundsen K, Stockinger EJ, Thomashow MF, Chen TH, Hayes PM (2005) Structural, functional, and phylogenetic characterization of a large CBF gene family in barley. Plant Mol Biol 59:533–551
Skinner JS, Szucs P, von Zitzewitz J, Marquez-Cedillo L, Filichkin T, Stockinger EJ, Thomashow MF, Chen TH, Hayes PM (2006) Mapping of barley homologs to genes that regulate low temperature tolerance in Arabidopsis. Theor Appl Genet 112:832–842
Solanke AU, Sharma AK (2008) Signal transduction during cold stress in plants. Physiol Mol Biol Plants 14:69–79
Soltesz A, Smedley M, Vashegyi I, Galiba G, Harwood W, Vagujfalvi A (2013) Transgenic barley lines prove the involvement of TaCBF14 and TaCBF15 in the cold acclimation process and in frost tolerance. J Exp Bot 64:1849–1862
Song Y, Chen Q, Ci D, Zhang D (2013) Transcriptome profiling reveals differential transcript abundance in response to chilling stress in Populus simonii. Plant Cell Rep 32:1407–1425
Steponkus PL, Uemura M, Webb MS (1993) A contrast of the cryostability of the plasma membrane of winter rye and spring oat-two species that widely differ in their freezing tolerance and plasma membrane lipid composition. In: Steponkus PL (ed) Advances in low-temperature biology, vol 2. JAI Press, London, pp 211–312
Stockinger EJ, Gilmour SJ, Thomashow MF (1997) Arabidopsis thaliana CBF1 encodes an AP2 domaincontaining transcription activator that binds to the C repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl Acad Sci U S A 94:1035–1040
Strand A, Hurry V, Gustafsson P, Gardeström P (1997) Development of Arabidopsis thaliana leaves at low temperatures releases the suppression of photosynthesis and photosynthetic gene expression despite the accumulation of soluble carbohydrates. Plant J 12:605–614
Strand A, Hurry V, Henkes S, Huner N, Gustafsson P, Gardeström P, Stitt M (1999) Acclimation of Arabidopsis leaves developing at low temperatures. Increasing cytoplasmic volume accompanies increased activities of enzymes in the Calvin cycle and in the sucrose-biosynthesis pathway. Plant Physiol 119:1387–1398
Strand A, Zrenner R, Trevanion S, Stitt M, Gustafsson P, Gardeström P (2000) Decreased expression of two key enzymes in the sucrose biosynthesis pathway, cytosolic fructose-1,6-bisphosphatase and sucrose phosphate synthase, has remarkably different consequences for photosynthetic carbon metabolism in transgenic Arabidopsis thaliana. Plant J 23:759–770
Strand Å, Foyer CH, Gustafsson P, Gardeström P, Hurry V (2003) Altering flux through the sucrose biosynthesis pathway in transgenic Arabidopsis thaliana modifies photosynthetic acclimation at low temperatures and the development of freezing tolerance. Plant Cell Envi 26:523–535
Stushnoff C, Remmele RL Jr, Essensee V, Mcneil M (1993) Low temperature induced biochemical mechanisms: implications for cold acclimation and de-acclimation. Springer NATO ASI series 16:647–657
Su CF, Wang YC, Hsieh TH, Lu CA, Tseng TH, Yu SM (2010) A novel MYBS3-dependent pathway confers cold tolerance in rice. Plant Physiol 153:145–158
Sung DY, Vierling E, Guy CL (2001) Comprehensive expression profile analysis of the Arabidopsis Hsp70 gene family. Plant Physiol 126:789–800
Suzuki N, Mittler R (2006) Reactive oxygen species and temperature stresses: a delicate balance between signaling and destruction. Physiol Plant 126:45–51
Tähtiharju S, Palva T (2001) Antisense inhibition of protein phosphatase 2C accelerates cold acclimation in Arabidopsis thaliana. Plant J 26:461–470
Tahtiharju S, Sangwan V, Monroy AF, Dhindsa RS, Borg M (1997) The induction of kin genes in cold-acclimating Arabidopsis thaliana. Evidence of a role for calcium. Planta 203:442–447
Takahashi D, Li B, Nakayama T, Kawamura Y, Uemura M (2013) Plant plasma membrane proteomics for improving cold tolerance. Front Plant Sci 4:90
Takumi S, Shimamura C, Kobayashi F (2008) Increased freezing tolerance through up-regulation of downstream genes via the wheat CBF gene in transgenic tobacco. Plant Physiol Biochem 46:205–211
Teige M, Scheikl E, Eulgem T, Doczi R, Ichimura K, Shinozaki K, Dangl JL, Hirt H (2004) The MKK2 pathway mediates cold and salt stress signaling in Arabidopsis. Mol Cell 15:141–152
Thakur P, Nayyar H (2013) Facing the cold stress by plants in the changing environment: sensing, signalling, and defending mechanisms. In: Tuteja N, Gill SS (eds) Plant acclimation to environmental stress. Springer, New York, pp 29–69
Theocharis A, Clement C, Barka EA (2012) Physiological and molecular changes in plants grown at low temperatures. Planta 235:1091–1105
Thomashow MF (1998) Role of cold-responsive genes in plant freezing tolerance. Plant Physiol 118:1–8
Thomashow MF (1999) Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50:571–599
Timperio AM, Egidi MG, Zolla L (2008) Proteomics applied on plant abiotic stresses: role of heat shock proteins (HSP). J Proteomics 71:391–411
Tominaga Y, Nakagawara C, Kawamura Y, Uemura M (2006) Effect of plasma membrane-associated proteins on acquisition of freezing tolerance in Arabidopsis thaliana. In: Chen THH, Uemura M, Fujikawa S (eds) Cold hardiness in plants: molecular genetics, cell biology and physiology. CABI Publishing, Oxfordshire, pp 235–249
Tondelli A, Francia E, Barabaschi D, Aprile A, Skinner JS, Stockinger EJ, Stanca AM, Pecchioni N (2006) Mapping regulatory genes as candidates for cold and drought stress tolerance in barley. Theor Appl Genet 112:445–454
Töpfer N, Caldana C, Grimbs S, Willmitzer L, Fernie AR, Nikoloski Z (2013) Integration of genome-scale modeling and transcript profiling reveals metabolic pathways underlying light and temperature acclimation in Arabidopsis. Plant Cell 25:1197–1211
Townley HE, Knight MR (2002) Calmodulin as a potential negative regulator of Arabidopsis COR gene expression. Plant Physiol 128:1169–1172
Tsvetkova NM, Horvath I, Torok Z, Wolkers WF, Balogi Z, Shigapova N, Crowe LM, Tablin F, Vierling E, Crowe JH et al (2002) Small heat-shock proteins regulate membrane lipid polymorphism. Proc Natl Acad Sci U S A 99(21):13504–13509
Tuteja N, Mahajan S (2007) Further characterization of calcineurin B-like protein and its interacting partner CBL-interacting protein kinase from Pisum sativum. Plant Signal Behav 2:358–361
Uemura M, Steponkus PL (1994) A contract of the plasma membrane lipid composition of oat and rye leaves inrelation to freezing tolerance. Plant Physiol 104:479–496
Uemura M, Yoshida S (1984) Involvement of plasma membrane alterations in cold acclimation of winter rye seedlings (Secale cereale L. cv Puma). Plant Physiol 75:818–826
Uemura M, Joseph RA, Steponkus PL (1995) Cold acclimation of Arabidopsis thaliana (effect on plasma membrane lipid composition and freeze-lnduced lesions). Plant Physiol 109:15–30
Uemura M, Warren G, Steponkus PL (2003) Freezing sensitivity in the sfr4 mutant of Arabidopsis is due to low sugar content and is manifested by loss of osmotic responsiveness. Plant Physiol 131:1800–1807
Uemura M, Tominaga Y, Nakagawara C, Shigematsu S, Minami A, Kawamura Y (2006) Responses of the plasma membrane to low temperatures. Physiol Plant 126:81–89
Urzua U, Kersten PJ, Vicuna R (1998) Manganese peroxidase-dependent oxidation of glyoxylic and oxalic acids synthesized by ceriporiopsis subvermispora produces extracellular hydrogen peroxide. Appl Environ Microbiol 64:68–73
Vannini C, Locatelli F, Bracale M, Magnani E, Marsoni M, Osnato M, Mattana M, Baldoni E, Coraggio I (2004) Overexpression of the rice Osmyb4 gene increases chilling and freezing tolerance of Arabidopsis thaliana plants. Plant J 37:115–127
Vaultier MN, Cantrel C, Vergnolle C, Justin AM, Demandre C, Benhassaine-Kesri G, Cicek D, Zachowski A, Ruelland E (2006) Desaturase mutants reveal that membrane rigidification acts as a cold perception mechanism upstream of the diacylglycerol kinase pathway in Arabidopsis cells. FEBS Lett 580:4218–4223
Venu RC, Sreerekha MV, Sheshu Madhav M, Nobuta K, Mohan KM, Chen S, Jia Y, Meyers BC, Wang G-L (2013) Deep transcriptome sequencing reveals the expression of key functional and regulatory genes involved in the abiotic stress signaling pathways in rice. J Plant Biol 56:216–231
Vogel JT, Zarka DG, Van Buskirk HA, Fowler SG, Thomashow MF (2005) Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis. Plant J 41:195–211
Wahl MC, Moller W (2002) Structure and function of the acidic ribosomal stalk proteins. Curr Protein Pept Sci 3:93–106
Wang X, Li W, Li M, Welti R (2006) Profiling lipid changes in plant response to low temperatures. Physiol Plant 126:90–96
Wang Y, Jiang CJ, Li YY, Wei CL, Deng WW (2012) CsICE1 and CsCBF1: two transcription factors involved in cold responses in Camellia sinensis. Plant Cell Rep 31:27–34
Wang XC, Zhao QY, Ma CL, Zhang ZH, Cao HL, Kong YM, Yue C, Hao XY, Chen L, Ma JQ, Jin JQ, Li X, Yang YJ (2013) Global transcriptome profiles of Camellia sinensis during cold acclimation. BMC Genomics 14:415
Webb MS, Steponkus PL (1993) Freeze-induced membrane ultrastructural alterations in rye (Secale cereale) leaves. Plant Physiol 101:955–963
Welin BV, Olson A, Nylander M, Palva ET (1994) Characterization and differential expression of dhn/lea/rab-like genes during cold acclimation and drought stress in Arabidopsis thaliana. Plant Mol Biol 26:131–144
Welling A, Palva ET (2008) Involvement of CBF transcription factors in winter hardiness in birch. Plant Physiol 147:1199–1211
Wisniewski J, Orosz A, Allada R, Wu C (1996) The C-terminal region of Drosophila heat shock factor (HSF) contains a constitutively functional transactivation domain. Nucleic Acids Res 24:367–374
Wisniewski ME, Bassett CL, Renaut J, Farrell R, Tworkoski T, Artlip TS (2006) Differential regulation of two dehydrin genes from peach (Prunus persica) by photoperiod, low temperature and water deficit. Tree Physiol 26:575–584
Wittmann-Liebold B, Graack HR, Pohl T (2006) Two-dimensional gel electrophoresis as tool for proteomics studies in combination with protein identification by mass spectrometry. Proteomics 6:4688–4703
Xiang Y, Huang Y, Xiong L (2007) Characterization of stress-responsive CIPK genes in rice for stress tolerance improvement. Plant Physiol 144:1416–1428
Xiang DJ, Hu XY, Zhang Y et al (2008) Overexpression of ICE1 gene in transgenic rice improves cold tolerance. Rice Sci 15(3):173–178
Xiang D, Man L, Yin K, Song Q, Wang L, Zhao M, Xu Z (2013) Overexpression of a ItICE1 gene from Isatis tinctoria enhances cold tolerance in rice. Mol Breeding 32:617–628
Xin Z, Browse J (1998) Eskimo1 mutants of Arabidopsis are constitutively freezing-tolerant. Proc Natl Acad Sci U S A 95(13):7799–7804
Xin Z, Mandaokar A, Chen J, Last RL, Browse J (2007) Arabidopsis ESK1 encodes a novel regulator of freezing tolerance. Plant J 49(5):786–799
Xin H, Zhu W, Wang L, Xiang Y, Fang L, Li J, Sun X, Wang N, Londo JP, Li S (2013) Genome wide transcriptional profile analysis of Vitis amurensis and Vitis vinifera in response to cold stress. PLoS One 8:e58740
Xiong Y, Fei SZ (2006) Functional and phylogenetic analysis of a DREB/CBF-like gene in perennial ryegrass (Lolium perenne L.). Planta 224:878–888
Xiong L, Ishitani M, Zhu JK (1999) Interaction of osmotic stress, temperature, and abscisic acid in the regulation of gene expression in Arabidopsis. Plant Physiol 119:205–212
Xiong L, Lee H, Ishitani M, Tanaka Y, Stevenson B, Koiwa H, Bressan RA, Hasegawa PM, Zhu JK (2002) Repression of stress-responsive genes by FIERY2, a novel transcriptional regulator in Arabidopsis. Proc Natl Acad Sci U S A 99:10899–10904
Xiong AS, Jiang HH, Zhuang J, Peng RH, Jin XF, Zhu B, Wang F, Zhang J, Yao QH (2013) Expression and function of a modified AP2/ERF transcription factor from Brassica napus enhances cold tolerance in transgenic Arabidopsis. Mol Biotechnol 53:198–206
Xu W, Jiao Y, Li R, Zhang N, Xiao D, Ding X, Wang Z (2014) Chinese wild-growing vitis amurensis ICE1 and ICE2 encode MYC-type bHLH transcription activators that regulate cold tolerance in Arabidopsis. PLoS One 9:e102303
Xue GP (2003) The DNA-binding activity of an AP2 transcriptional activator HvCBF2 involved in regulation of low-temperature responsive genes in barley is modulated by temperature. Plant J 33:373–383
Yaish MW, Doxey AC, McConkey BJ, Moffatt BA, Griffith M (2006) Cold-active winter rye glucanases with ice-binding capacity. Plant Physiol 141:1459–1472
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264
Yamazaki T, Kawamura Y, Minami A, Uemura M (2008) Calcium-dependent freezing tolerance in Arabidopsis involves membrane resealing via synaptotagmin SYT1. Plant Cell 20:3389–3404
Yan SP, Zhang QY, Tang ZC, Su WA, Sun WN (2006) Comparative proteomic analysis provides new insights into chilling stress responses in rice. Mol Cell Proteomics 5:484–496
Yang T, Poovaiah BW (2003) Calcium/calmodulin mediated signal network in plants. Trends Plant Sci 8:505–512
Yang W, Liu XD, Chi XJ, Wu CA, Li YZ, Song LL, Liu XM, Wang YF, Wang FW, Zhang C, Liu Y, Zong JM, Li HY (2011) Dwarf apple MbDREB1 enhances plant tolerance to low temperature, drought, and salt stress via both ABA-dependent and ABA-independent pathways. Planta 233:219–229
Yang A, Dai X, Zhang WH (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Yi AB, Yu LX, Hua T, Hong LW, Wen SL (2011) Comparative profile of Rubisco-interacting proteins from Arabidopsis : photosynthesis under cold conditions. Progr Biochem Biophys 38(5):455–463
Yin Z, Rorat T, Szabala BM, Ziółkowska A, Malepszy S (2006) Expression of a Solanum sogarandinum SK3-type dehydrin enhances cold tolerance in transgenic cucumber seedlings. Plant Sci 170:1164–1172
Yoshida S (1991) Chilling induced inactivation and its recovery of tonoplast H + -ATPase in mung bean cell suspension cultures. Plant Physiol 95:456–460
Zarka DG, Vogel JT, Cook D, Thomashow MF (2003) Cold induction of Arabidopsis CBF genes involves multiple ICE (inducer of CBF expression) promoter elements and a cold-regulatory circuit that is desensitized by low temperature. Plant Physiol 133:910–918
Zhai H, Bai X, Zhu Y, Li Y, Cai H, Ji W, Ji Z, Liu X, Liu X, Li J (2010) A single-repeat R3-MYB transcription factor MYBC1 negatively regulates freezing tolerance in Arabidopsis. Biochem Biophys Res Commun 394(4):1018–1023
Zhang JZ, Creelman RA, Zhu JK (2004) From laboratory to field. Using information from Arabidopsis to engineer salt, cold, and drought tolerance in crops. Plant Physiol 135(2):615–621
Zhang W, Jiang B, Li W, Song H, Yu Y, Chen J (2009) Polyamines enhance chilling tolerance of cucumber (Cucumis sativus L.) through modulating antioxidative system. Sci Hortic (Amsterdam) 122:200–208
Zhang J, Mao Z, Chong K (2013) A global profiling of uncapped mRNAs under cold stress reveals specific decay patterns and endonucleolytic cleavages in Brachypodium distachyon. Genome Biol 14:R92
Zhao H, Bughrara SS (2008) Isolation and characterization of cold-regulated transcriptional activator LpCBF3 gene from perennial ryegrass (Lolium perenne L.). Mol Genet Genomics 279:585–594
Zhao M-G, Chen L, Zhang L-L, Zhang W-H (2009) Nitric reductase-dependent nitric oxide production is involved in cold acclimation and freezing tolerance in Arabidopsis. Plant Physiol 151:755–767
Zhao Z, Tan L, Dang C, Zhang H, Wu Q, An L (2012) Deep-sequencing transcriptome analysis of chilling tolerance mechanisms of a subnival alpine plant, Chorispora bungeana. BMC Plant Biol 12:222
Zhao ML, Wang JN, Shan W, Fan JG, Kuang JF, Wu KQ, Li XP, Chen WX, He FY, Chen JY, Lu WJ (2013) Induction of jasmonate signalling regulators MaMYC2s and their physical interactions with MaICE1 in methyl jasmonate-induced chilling tolerance in banana fruit. Plant Cell Environ 36:30–51
Zhu J, Shi H, Lee BH, Damsz B, Cheng S, Stirm V, Zhu JK, Hasegawa PM, Bressan RA (2004) An Arabidopsis homeodomain transcription factor gene, HOS9, mediates cold tolerance through a CBF-independent pathway. Proc Natl Acad Sci U S A 101(26):9873–9878
Zhu J, Dong CH, Zhu JK (2007) Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr Opin Plant Biol 10:290–295
Zhu Y, Yang H, Mang HG, Hua J (2011) Induction of BAP1 by a moderate decrease in temperature is mediated by ICE1 in Arabidopsis. Plant Physiol 155(1):580–588
Acknowledgements
Research in MA lab is funded by the Department of Biotechnology and Delhi University.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this chapter
Cite this chapter
Sinha, S. et al. (2015). The Omics of Cold Stress Responses in Plants. In: Pandey, G. (eds) Elucidation of Abiotic Stress Signaling in Plants. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-2540-7_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2540-7_6
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-2539-1
Online ISBN: 978-1-4939-2540-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)