Abstract
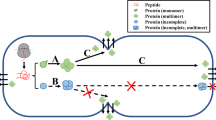
Recent development of large-scale analyses such as the secretome analysis has enabled the discovery of a vast number of intracellular proteins that are secreted outside the cell. Often, these proteins do not contain any known signal sequence required for conventional protein secretion. In order to avoid misidentification of such “leaked” proteins as “secreted” proteins, reconstructing the process of protein secretion is essential. Here, we describe methods for the detection of reconstructed unconventional protein secretion and determination of regulatory proteins of secretion in Saccharomyces cerevisiae. We show that conjugating target proteins with a tag-sequence and utilizing various reagents and tools can facilitate quantitative detection of the secretion of target proteins. We expect that these methods will reveal novel unconventional secretion pathways of proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
van Deventer HJ, Goessens WH, van Vliet AJ et al (1996) Anti-enolase antibodies partially protective against systemic candidiasis in mice. Clin Microbiol Infect 2:36–43
Torimura T, Ueno T, Kin M et al (2001) Autocrine motility factor enhances hepatoma cell invasion across the basement membrane through activation of beta1 integrins. Hepatology 34:62–71
Dobashi Y, Watanabe H, Matsubara M et al (2006) Autocrine motility factor/glucose-6-phosphate isomerase is a possible predictor of metastasis in bone and soft tissue tumours. J Pathol 208:44–53
Capello M, Ferri-Borgogno S, Cappello P et al (2011) α-Enolase: a promising therapeutic and diagnostic tumor target. FEBS J 278:1064–1074
Villarreal L, Méndez O, Salvans C et al (2013) Unconventional secretion is a major contributor of cancer cell line secretomes. Mol Cell Proteomics 12:1046–1060
Jeffery CJ (2013) New ideas on protein moonlighting. In: Henderson B (ed) Moonlighting cell stress proteins in microbial infections. Springer, New York, pp 51–66
Nickel W, Rabouille C (2009) Mechanisms of regulated unconventional protein secretion. Nat Rev Mol Cell Biol 10:148–155
Duran JM, Anjard C, Stefan C et al (2010) Unconventional secretion of Acb1 is mediated by autophagosomes. J Cell Biol 188:527–536
Manjithaya R, Anjard C, Loomis WF et al (2010) Unconventional secretion of Pichiapastoris Acb1 is dependent on GRASP protein, peroxisomal functions, and autophagosome formation. J Cell Biol 188:537–546
Zhang M, Schekman R (2013) Unconventional secretion, unconventional solutions. Science 340:559–561
Miura N, Kirino A, Endo S et al (2012) Tracing putative trafficking of the glycolytic enzyme enolase via SNARE-driven unconventional secretion. Eukaryot Cell 11:1075–1082
Schekman R (2010) Charting the secretory pathway in a simple eukaryote. Mol Biol Cell 21:3781–3784
Oliveira DL, Nakayasu ES, Joffe LS et al (2010) Characterization of yeast extracellular vesicles: evidence for the participation of different pathways of cellular traffic in vesicle biogenesis. PLoS ONE 5:e11113
Giardina BJ, Stanley BA, Chiang HL (2014) Glucose induces rapid changes in the secretome of Saccharomyces cerevisiae. Proteome Sci 12:9
Shinya R, Morisaka H, Kikuchi T et al (2013) Secretome analysis of the pine wood nematode Bursaphelenchus xylophilus reveals the tangled roots of parasitism and its potential for molecular mimicry. PLoS ONE 21:e67377
de Wit M, Kant H, Piersma SR et al (2014) Colorectal cancer candidate biomarkers identified by tissue secretome proteome profiling. J Proteomics 99C:26–39
Wasinger VC, Zeng M, Yau Y (2013) Current status and advances in quantitative proteomic mass spectrometry. Int J Proteomics 2013:180605
Miura N, Shinohara M, Tatsukami Y et al (2013) Spatial reorganization of Saccharomyces cerevisiae enolase to alter carbon metabolism under hypoxia. Eukaryot Cell 12:1106–1119
Muthusamy BP, Natarajan P, Zhou X, Graham TR (2009) Linking phospholipid flippases to vesicle-mediated protein transport. Biochim Biophys Acta 1791:612–619
Bröcker C, Engelbrecht-Vandré S, Ungermann C (2010) Multisubunit tethering complexes and their role in membrane fusion. Curr Biol 20:R943–R952
Novick P, Schekman R (1979) Secretion and cell-surface growth are blocked in a temperature-sensitive mutant of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 76:1858–1862
Novick P, Field C, Schekman R (1980) Identification of 23 complementation groups required for post-translational events in the yeast secretory pathway. Cell 21:205–215
Bryant NJ, Stevens TH (1998) Vacuole biogenesis in Saccharomyces cerevisiae: protein transport pathways to the yeast vacuole. Microbiol Mol Biol Rev 62:230–247
Hua Z, Fatheddin P, Graham TR (2002) An essential subfamily of Drs2p-related P-type ATPases is required for protein trafficking between Golgi complex and endosomal/vacuolar system. Mol Biol Cell 13:3162–3177
Gall WE, Geething NC, Hua Z, Ingram MF, Liu K, Chen SI, Graham TR (2002) Drs2p-dependent formation of exocytic clathrin-coated vesicles in vivo. Curr Biol 12:1623–1627
Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y (2009) Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol 10:458–467
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132
Entian KD, Schuster T, Hegemann JH et al (1999) Functional analysis of 150 deletion mutants in Saccharomyces cerevisiae by a systematic approach. Mol Gen Genet 262:683–702
Matsui K, Kuroda K, Ueda M (2009) Creation of a novel peptide endowing yeasts with acid tolerance using yeast cell-surface engineering. Appl Microbiol Biotechnol 82:105–113
Ito H, Fukuda Y, Murata K et al (1983) Transformation of intact yeast cells treated with alkali cations. J Bacteriol 153:163–168
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Miura, N., Ueda, M. (2015). Evaluation of Unconventional Protein Secretion in Saccharomyces cerevisiae . In: Tang, B. (eds) Membrane Trafficking. Methods in Molecular Biology, vol 1270. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2309-0_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2309-0_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2308-3
Online ISBN: 978-1-4939-2309-0
eBook Packages: Springer Protocols