Abstract
In most archaeons and many bacteria, the cell surface is covered by a monomolecular and paracrystalline protein layer, called S-layer. S-layers serve as an outer protective shell and also act as molecular sieves and/or as cell-adhesion sites for exoenzymes. In bacteria, S-layers are associated either with peptidoglycan or with secondary cell wall polymers of the outer membrane. In archaea, the S-layer proteins (SLPs) are anchored to the cytoplasmic membrane and form the major cell wall component. A prominent feature of SLPs is their self-assembly into mono- or bilayers in solution or at interfaces. The unique self-assembling feature of S-layers provides the basis for a wide range of applications in biotechnology and nanobiology. This chapter covers the structural aspects of SLPs and S-layer assembly.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Surface Layer Protein
- Methanosarcina Acetivorans
- Methanosarcina Mazei
- Secondary Cell Wall Polymer
- Secondary Cell Wall Polysaccharide
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
2.1 Introduction
The majority of bacteria and archaea develop multicomponent cell envelopes with layered supramolecular architectures surrounding the cytoplasmic membrane. Of the finest examples of such supramolecular cell wall components are S-layers or surface layers, regular paracrystalline pericellular structures, first observed in the bacterium Spirillum serpens and in the archaeum Halobacterium salinarum (Houwink 1956, 1953) . Following their first observation and the realization that S-layers were of a proteinaceous nature, the first S-layer protein (SLP) gene sequence to be published was that of Brevibacillus brevis 47 in 1986 (Tsuboi et al. 1986) . It is now well established that S-layers are composed of monolayers of glycoproteins/proteins with an Mr range of 40–200 kDa, many of which have been thoroughly characterized by genetic and morphogenetic studies as well as by structural, biophysical and biochemical means (Altman et al. 1990; Fagan and Fairweather 2014; Kuen et al. 1994; Messner et al. 1986a, b; Sleytr et al. 1986, 1993) . Generally, different SLPs give rise to defined monomolecular lattices, although bimolecular S-layers are known for Clostridium difficile (Takeoka et al. 1991) and Bacillus anthracis (Etienne-Toumelin et al. 1995) . The S-layer two-dimensional (2D) arrays are formed through non-covalent self-assembly events of the SLP subunits, though exceptionally covalent cross links between the protein subunits are found, as in Methanosprillum hungatei (Beveridge et al. 1985; Messner et al. 2008; Sleytr and Beveridge 1999; Stewart et al. 1985) . Bacterial S-layer lattices have p1, p2 (oblique), p4 (square) or p3 or p6 (hexagonal) symmetries associated with them, whilst archaeal S-layers are predominantly hexagonal in symmetry (Albers and Meyer 2011; Sleytr et al. 1993) . Topologically, S-layers completely encapsulate their producing cell (Figure 2.1). Their mode of anchoring to the cell envelope can be broadly classified in accordance with the archaeal, Gram-positive or Gram-negative cell architecture (Pum et al. 2013) . In archaea, the S-layer is directly associated with the cytoplasmic membrane through a single-span transmembrane segment and forms the dominant component of the cell wall. The S-layer functions primarily as a protective barrier, though in Methanosarcina mazei the SLPs were also shown to be involved in cell–cell association (Mayerhofer et al. 1992) . In Gram-positive bacteria, the S-layers lie atop the peptidoglycan layer and are anchored to the peptidoglycan or the associated nonclassical secondary cell wall polymers (SCWPs; Mesnage et al. 2000; Schaffer and Messner 2005; Zhao et al. 2006) . In Gram-negatives, the S-layers are associated with the lipopolysaccharides (LPS) in the outer membrane (OM; Ford et al. 2007) . SLPs reach the cell surface using the general secretory pathway (SEC translocon), except in Gram-negative bacteria, where the diderm cell walls require more elaborate translocative routes such as type I and II secretion pathways (Noonan and Trust 1995; Thompson et al. 1998) .
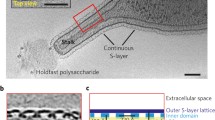
a Cryo electron tomogram of the intact cell-bound S-layer (wt-SbpA) of Lysinibacillus sphaericus (formerly, Bacillus sphaericus), a Gram-positive soil bacterium that forms a 2D crystalline envelope with p4 symmetry (cell constant: ~ 13 nm). Inset shows the close-up view of the cell wall and the enveloping SbpA S-layer. Scale bar: 50 nm (image courtesy of Luis R. Comolli). b Electron tomogram of Caulobacter crescentus cell, overlaid with the 3D reconstruction of the RsaA S-layer, which can be seen to form a hexagonal repeating units (six RsaA monomers) interconnected at their junctions by p3 symmetry. The cutaway region at the cell pole shows the S-layer as the outermost layer of the cell envelope. Scale bar: 250 nm (image courtesy of Luis R. Comolli). c Negative stain EM micrograph of an in vitro self-assembly product of the Geobacillus stearothermophilus PV72/p2 SLP SbsB (mutant S347-HA). (Reproduced from Kinns et al. 2010). Scale bar: 100 nm
2.2 SLP Composition and Modification
SLPs are distributed over a wide range of phylogenetic branches (Messner and Sleytr 1992; Sleytr et al. 1999) and generally show little (< 20 % sequence identity) to no apparent sequence conservation, even within a single species or closely related taxons . The recent advances in SLP structural biology (Baranova et al. 2012; Ethordic et al. 2012; Fagan et al. 2009; Jing et al. 2002; Kern et al. 2011; Pavkov et al. 2003, 2008; Stetefeld et al. 2000; see below) reveal a diverse domain composition, indicating different SLPs do not generally have a common origin. Relatively, higher similarity is observed in S-layers of Gram-positive bacteria as compared to different S-layers from distant bacteria and archaea (Engelhardt and Peters 1998) . Nevertheless, SLPs show similar overall amino acid compositions that are low in methionine and negligible cysteine. For the majority of SLPs, an enrichment in aspartic and glutamic acid results in isoelectric points (pIs) that lie in the weakly acidic range (Messner et al. 1997; Sara and Sleytr 2000) , with exceptions for Methanothermus fervidus (pI: 8.4; Brockl et al. 1991) and lactobacilli (pI: 9–11; Boot et al. 1995) . An exception to the low overall sequence similarity is the ~ 55-amino-acid S-layer homology (SLH) motif , frequently found in tandem repeats near the N-terminus of many bacterial SLPs and surface-anchored enzymes (Mesnage et al. 2000; Schaffer and Messner 2005; Zhao et al. 2006) . The X-ray structure of the three tandem SLH motifs in the B. anthracis surface array protein (Sap) shows that these form helix-loop-helix structures that together organize into a single three-prong spindle domain (Kern et al. 2011) . The SLH domains are cell-wall-anchoring domains that bind the peptidoglycan, either directly as in Paenibacillus alvei CCM 2051 (Janesch et al. 2013a, b) or through the associated pyruvylated carbohydrate moieties in the nonclassical SCWPs (Mesnage et al. 2000; Schaffer and Messner 2005; Zhao et al. 2006) . SLH domains are not universal S-layer-anchoring modules, however. In Geobacillus stearothermophilus SbsC, an extended three-helix bundle is found as N-terminal glycan-binding domain (Pavkov et al. 2008), and in Corynebacterium glutamicum this role is taken by the C-terminal hydrophobic domain (Bahl et al. 1997; Chami et al. 1997) . In Gram-negatives, the S-layers attach to the OM LPS via direct or divalent cation-mediated charge interactions as in Campylobacter fetus or Aeromonas salmonicida (Garduno et al. 1992) , or by covalently bound fatty acids as in Deinococcus radiodurans Hpi (Peters et al. 1987) .
Both bacterial and archaeal SLPs are frequently glycosylated and S-layer glycoproteins (SLGPs) have provided a major impetus to the prokaryotic glycoprotein research. SLGPs have been described for halophilic archaeons such as H. salinarum (glycoprotein with a M r of 200,000 and carbohydrate content of 10–12 %; Mescher and Strominger 1976) , Haloferax volcanii (presence of O-glycosidic linkages like “Thr/Ser-Gal” and 1, 2-linked glucosyl-galactose as a structural element in S-layer; Eichler and Adams 2005; Sumper et al. 1990) , hyperthermophilic methanogens such as M. fervidus (Brockl et al. 1991) , Methanothermus sociabilis and Methanococcus voltae (Konisky et al., 1994) and the thermoacidophilic crenarchaeote Sulfolobus acidocaldarius (oligosaccharide is a complex, branched, six-membered glycan tree; Peyfoon et al. 2010) . In Gram-positive bacteria, SLP glycosylation is found in clostridia and bacilli such as Thermoanaerobacter, Geobacillus, Aneurinibacillus, Paenibacillus and Lactobacillus species, whereas in Gram-negatives SLP glycosylation has only been described for Tannerella forsythia TfsA and TfsB (Posch et al. 2011; Schaffer and Messner 2004; Sleytr and Thorne 1976) . The glycan chains can be branched, though in bacteria typically contain an extended linear stretch of repeating units (10–50 units) that is frequently capped by O-methylation in the 2′ or 3′ position of the nonreducing glycan and is anchored to the protein via a core of two to four sugar residues and an O-glycosidic linkage (galactose–tyrosine, glucose–tyrosine, N-acetyl galactosamine–serine, N-acetyl galactosamine–threonine) . Archaeal glycans are shorter and generally attached by an N-glycosidic bond to asparagine, though O-linked chains have also been observed (Altman et al. 1996; Messner 1997; Wugeditsch et al. 1999) . The number of glycosylation sites can vary from two to four in Gram-positive SLPs, to up to 25 in archaeal SLPs (Lechner et al. 1986) . Additional post-translational modifications of SLPs include sulphated glycan chains of Halobacter halobium to provide stability to the S-layer or phosphorylation of tyrosine residues in Aeromonas hydrophila to decrease the pI (6.7–4.6) .
2.3 Functional Diversity of S-Layers
The primary function of S-layers is taken to be a role in cell stabilization, which can be mechanical, thermal and osmotic in nature (Engelhardt 2007) . From the accumulated data, it is evident that S-layers exhibit high levels of physical and chemical stability, indicating they can act as protective and/or shape-maintaining exoskeleton (Engelhardt and Peters 1998) . Experiments in Halobacteria indeed found that the rod-like shape was lost upon S-layer lattice degradation, supporting the notion that S-layers can impart shape maintenance (Engelhardt 2007; Mescher and Strominger 1976) . Even though S-layers can maintain the shape of the cell and to a certain extent be shape-modifying, the primary shape determination is endoskeletal in origin (Baumeister and Lembcke 1992; Engelhardt 2007; Messner et al. 1986; Peters et al. 1995; Pum et al. 1991) . Another protective role of S-layers can come from a shielding function against environmental factors. S-layers in Gram-negative bacteria such as A. salmonicida, C. fetus and Caulobacter crescentus have been observed to form a protective covering for bacterial parasites like Bdellovibrio bacteriovorous (Koval 1988; Koval and Bayer 1997; Koval and Hynes 1991) , though the S-layers did not provide protection from protozoans. S-layers have also been observed to attenuate host immune responses in the periodontal pathogen T. forsythia (Sekot et al. 2011) . Also, in the opportunistic pathogen C. fetus, the surface expression of SLPs prevented binding of complement component C3b and in A. salmonicida the S-layer (A-layer) imparts high to moderate levels of anti-bactericidal activity against complement systems (Dworkin and Blaser 1997) . The A-layer also holds an adhesive property, able to bind laminins, fibronectins (Doig et al. 1992) and basement membrane protein collagen type IV (Trust et al. 1993) . Also in other pathogens like T. forsythia, C. difficile and B. anthracis, the S-layers have been found to contribute to virulence by mediating host cell adhesion (Calabi et al. 2002; Kern and Schneewind 2010; Sakakibara et al. 2007) .
Besides a supporting role in maintaining cell integrity or in bacterial virulence, several S-layers have also been attributed a scaffolding function. Archaeal S-layers are composed of long hydrophobic protrusions entering the plasma membrane, which have been proposed to provide a contact zone for macromolecules that function in folding and export of proteins and/or nutrient degradation and transport mechanisms (Lechner and Sumper 1987) . Also, in many Bacillaceae, S-layers provide adhesion sites for cell-associated exoenzymes (Sleytr et al. 1999) . Well-documented examples include the high-molecular-weight amylases in G. stearothermophilus strains DSM 2358 and ATCC 12980 (Egelseer et al. 1995, 1996) , endo-xylanase in Thermoanaerobacterium thermohydrosulfurigen strain JW/SL-YS 485 (Liu et al. 1996) and endo-glucanase in Clostridium thermocellum (Leibovitz et al. 1997) . Finally, an interesting feature observed in the unicellular cyanobacterium Synechococcus strain GL24 (strain isolated from a meromictic Fayetteville Green Lake, New York, USA, with high levels of calcium and sulphate ions) is that the hexagonal S-layer acts as a template for sulphate and carbonate mineral formation over its surface (Schultzelam and Beveridge 1994a, b) .
Thus, although a general function of an S-layer appears to be the provision of a 2D monomolecular array outside the cell, various genetic and functional studies found that they are phylogenetically and structurally dissimilar and attain varied roles in cellular activities, leading to a high degree of functional heterogeneity .
2.4 Structural Biology of S-Layers
Over the past two decades, there has been a steady increase in the level of molecular and structural understanding of the surface layer proteins (Table 2.1) . In the initial phase, S-layer structural biology was mainly based on freeze-etching, freeze-drying, negative staining followed by 2D and 3D transmission electron microscopy (TEM) and on scanning probe microscopy (Beveridge 1993; Beveridge et al. 1993; Firtel et al. 1994) . This was followed by efforts towards the 3D prediction of the building components using the mean force approach, and by simulations of self-assembly process by Monte-Carlo simulations (Horejs et al. 2008, 2011) . Cryo-electron microscopy and tomography (cryo-EM, cryo-ET) in turn provided depth to the topographical information derived from the SLPs (Baumeister and Lembcke 1992; Kinns et al. 2010; Norville et al. 2007; Rachel et al. 1986; Shin et al. 2013; Smit et al. 1992; Figure 2.1). 3D crystallization of SLPs has long been plagued by the inherent self-aggregation of SLPs. Nevertheless, in recent years, X-ray crystallography has started to reveal the molecular architecture of the SLPs and/or non-assembling SLP fragments (Baranova et al. 2012; Ethordic et al. 2012; Fagan et al. 2009; Jing et al. 2002; Kern et al. 2011; Pavkov et al. 2003, 2008; Stetefeld et al. 2000; Figures 2.2, 2.3, 2.4, 2.5, 2.6, 2.7, 2.8) and has started to provide more molecular insights into the build-up of the supramolecular assemblies . In the following paragraphs, these recent advances in SLP structural biology are systematically reviewed. As common structural architecture, bacterial SLPs appear to have the self-assembling or “crystallization domain” located in the C-terminal part of the protein, preceded by the cell-wall-anchoring domain(s), mostly SLH domains, at the N-terminus. The SLH domains are present in the majority of bacterial SLPs, though there are exceptions like SbsC in G. stearothermophilus (Pavkov et al. 2008) .
a Schematic diagram of the S. marinus SLP, which forms a filiform tetrameric unit called tetrabrachion (70 nm in length). The tetrabrachion is composed of an extended parallel-coiled coil stalk region with C-terminal membrane anchor region and four extending arms (24 nm each in length) at its N-terminal end (Peters et al. 1995). The tetrameric stalk contains a protease (represented as globular structures)-binding domain, the atomic structure of which (grey boxed area) is shown in panel b. b Crystal structure of the right-handed parallel coiled coil region (RHCC) (PDB code: 1FE6) corresponding to the protease-binding domain (grey boxed area), shown in lateral and axial view. (Stetefeld et al. 2000)
a Schematic representation of the domain composition of the M. mazei SLP MA1904 (Jing et al. 2002). At its N-terminus, the SLP contains two “YVTN” β-propeller domains (each with seven blades composed of four-stranded β-sheets labelled W), followed by twelve (1+11) consecutive PKD domains. b Crystal structure of a 42 kDa N-terminal SLP fragment (boxed area in panel a) reveals the molecular structure of a “YVTN” β-propeller domain (blades labelled W1 to W7) and a single-polycystic-kidney disease (PKD1) domain (PDB code: 1L06)
a Domain architecture of the SlpA preprotein from C. difficile CD 630. The preprotein is proteolytically processed into a high- and low-molecular weight SLP (HMW- and LMW-SLP, resp.). Cleavage sites are represented by small triangles. b Crystal structure of LMW-SLP1-262 (PDB code: 3CVZ) encompassing domains I and II, spanning ~ 90 Å in length
a A schematic representation of the M. acetivorans MA0829 S-layer protein with N-terminal signal sequence, followed by a tandem repeat of DUF1608 domains (NTR and CTR), a tether sequence and finally, the C-terminal membrane-anchoring region. b The crystal structure of DUF1608 (CTR)/MA0829 CTR (294–570 residues) (PDB code: 3U2G) reveals two subdomains, domain I and II. c Putative model of the MA0829 S-layer lattice generated based on crystallographic dimers of the CTR. The model shows a semiporous lattice with three distinct pores, labelled P (primary), A (asymmetric) and T (trimer). (Image courtesy of Mark A. Arbing)
a A schematic representation of domain composition of SbsC from G. stearothermophilus, with experimentally observed domains labelled I–VI, followed by a C-terminal fragment of unknown domain structure, coloured grey. Domains I–III correspond to rSbsC(31-443) and domains I–VI correspond to rSbsC(31-844). b Crystal structure of rSbsC(31-443) (PDB code: 2RA1) revealing the N-terminal cell wall attachment domain (domain I, α1–α7) and partial C-terminal crystallization domains (II and III). c Crystal structure rSbsC(31-844) (adapted from Pavkov et al. 2008), with the modelled structure comprising the domains I, II and III shown in ribbon representation and the unmodelled density for domains IV, V and VI in surface representation. Domains II–VI form a planar, ring-like structure similar to that seen in SbsB (see Figure 2.8). Adjacent view is by a lateral rotation of 90°
a A schematic representation of domain composition of mature SbsB from G. stearothermophilus show the N-terminal cell wall attachment domain, domain I (composed of three SLH domains), and the C-terminal crystallization domain composed of domains II–VII that can self-assembly to form the S-layer lattice. b Crystal structure of SbsB32-920 (PDB code: 4AQ1) in complex with the single domain antibody NbKB6 (not shown in the figure), with domains II–VII in a φ-shaped, disk-like quaternary structure. c Ribbon and space-filling representation of the calculated S-layer model in extracellular view (top panel) and side view, with the cell-wall facing surface pointing down (lower panel). For clarity, one protomer is coloured in rainbow from N- to C-terminus
2.4.1 Structure of SLP Fragment from Staphylothermus marinus
Staphylothermus marinus is a hyperthermophilic archaeobacterium. EM studies provided initial insight into the S-layer of S. marinus (Peters et al. 1995) . The SLP subunits organize into ~ 70-nm-long filiform “tetrabrachions” (M r 92,000) by tetramerization via a parallel, four-stranded α-helical structure (Figure 2.2). The presence of leucine and isoleucine throughout the hydrophobic core in the long α-helical stalk provides an extreme level of thermal stability to the structure, resistance to heat denaturation at 130 °C in the presence of 1 % (w/v) dodecyl sulphate or 6M guanidine (Peters et al. 1996) . The 1.8 Å X-ray structure of a 52-amino-acid recombinant fragment encompassing a protease-binding region of the tetrabrachion (PDB code: 1FE6) shows a right-handed coiled coil structure (RHCC; with dimensions of ~ 72 Å length and ~ 25 Å in diameter, respectively; Stetefeld et al. 2000; Figure 2.2b). The RHCC structure exhibits an 11-residue repeat (Ile 19 to Leu 29), which is similar in properties to the 11-residue repeat (Leu 3 to Ile 13) of the de novo designed RH4 peptide tetramer. Three intra-helical (Glu 24—Lys27; Asp 10—Arg14 and Asp 21—Arg 22 pairs) and four inter-helical (Arg 33—Asp 35; Arg 36—Glu 38; Asp 20—Arg 22 and Asp 9—Arg 14 pairs) salt bridges provide electrostatic interactions in the coiled coil structure. The RHCC is further stabilized by a series of complex hydrophobic interactions between the four helices, which play a major role in thermal stability of the tetrabrachion stalk domain (Peters et al. 1995, 1996) . This RHCC crystal structure provides the template for proper understanding of the stability shown by hyperthermophilic proteins and has helped inspire de novo designing strategies for thermostable proteins scaffolds .
2.4.2 Structure of SLP from M. mazei
The 2.4-Å resolution structure of an N-terminal fragment from the MA1904 SLP from the methanogenic archaeobacterium M. mazei provides structural information on an architecturally different class of archaeal SLPs (PDB code: 1L0Q; Jing et al. 2002) . MA1904 is a 1564-residue mature protein that reveals repetitive domain build-up composed of two YVTN heptarepeat domains and a series of polycystic-kidney disease (PKD)-like domains (Figure 2.3a). Jing et al. (2002) crystallized a 42-kDa MA1904 fragment encompassing the N-terminal YVTN heptarepeat domain (residues 1–302) and the subsequent PKD domain (residues 303–391). The YVTN repeats give rise to a seven-bladed β-propeller, with each propeller blade composed of a four-stranded antiparallel β-sheet, whilst the PKD domain is formed of an eight-stranded β-sandwich (Figure 2.3b). Based on these domain structures, the full-length M. mazei SLP can be discerned to comprise 14 YVTN repeats grouped in two β-propeller domains and 12 PKD domains. Apart from MA1904, genome analysis of metazoans reveals multiple proteins that possess similar six-bladed YWTN β-propeller domains in cell surface and extracellular matrix-associated proteins. These are sequentially similar to the YVTN β-propeller domain seen in the MA1904 structure and are expected to differ only at the position of a β-bulge in β-strand 4 (Jing et al. 2002) and the number of propeller blades. Along with these two domains, many SLPs in archaea also exhibit a third kind of repeat, which is predicted to form a right-handed parallel β-helix domain (Galagan et al. 2002) .
2.4.3 Structure of SlpA from C. difficile
SlpA is a surface layer protein from the Gram-positive, spore-forming, anaerobic bacterium C. difficile, which causes opportunistic gastrointestinal tract infections called C. difficile–associated disease (CDAD; Fagan et al. 2009; Poxton et al. 2001) . The C. difficile S-layer contains a 374-residue “high molecular weight SLP” (HMW-SLP) and a 321-residue “low molecular weight SLP” (LMW-SLP), which associate to form a tight non-covalent complex (H/L) that gives rise to an elongated particle, as revealed by small-angle X-ray scattering (SAXS) analysis (Fagan et al. 2009) . A 3D crystal structure is available of a LMW-SLP (321 amino acid residues in length, 34.2 kDA without signal peptide) from C. difficile CD630 determined at 2.4 Å resolution (PDB code: 3CVZ). The crystal structure reveals that LMW-SLP is composed of two non-contiguous domains (Figure 2.4b; Fagan et al. 2009) . Domain 1 encompasses residues 1–88 and 239–249 that form a two-layered sandwich structure composed of five-stranded mixed β-sheets packed against a pair of antiparallel α-helices. Domain 2 encompasses residues 89–238 and exhibits a novel fold with two β-hairpins (residues 150–170 and 211–227) and an elongated α-helix (residues 97–111), interspersed by extended loop regions. Mutagenesis studies show the C-terminal end of LMW-SLP to be involved in the formation of the LMW–HMW complex. Further, the HMW-SLP contains the “cell-wall-binding” motifs (PF04122; Calabi et al. 2001) that establish binding with the cell wall .
2.4.4 Structure of the SLH Domain of Sap from B. anthracis
Sap is an SLP in the Gram-positive, rod-shaped, spore-forming mammalian pathogen B. anthracis . As many other bacterial SLPs, Sap utilizes three consecutive surface-layer homology (SLH) repeats to be tethered to the secondary cell wall polysaccharide (SCWP) in the bacterial cell envelope (Mesnage et al. 2000) . Kern et al. provided the first crystal structure of an SLH domain (SapSLH, residues 31–210, N-terminal His-tag), solved at a resolution of 1.8 Å (Kern et al. 2011; PDB code: 3PYW; Figure 2.5a, b). The 3D structure is defined to be a three-pronged spindle with each prong arising from a single SLH repeat. The spindle’s base is formed by a three-helical bundle that is contributed by the three SLH domains, respectively (each with one helix) . Solvent-accessible surface area analysis revealed each SLH repeat contains a small tunnel. The SLH domains (SLH1, residues 31–90; SLH2, residues 91–151; SLH3, residues 152–209) exhibit a partially conserved ITRAE motif (not shown in figure). Both the tunnels and ITRAE motifs contribute positively charged residues to the surface structure of inter-prong grooves formed by all the three prongs of SLP. Kern et al. (2011) propose that these positively charged residues play a major role in binding of SLP to the pyruvyl-ketal of SCWP .
2.4.5 Structure of the DUF1608 Domain of Methanosarcina acetivorans
M. acetivorans is a methanogenic archaeon with the cytoplasmic membrane surrounded by an S-layer composed of a single protein, MA0829. Depending on the environmental conditions, the S-layer may additionally be covered with a layer of heteropolysaccharide (methanochondroitin; Francoleon et al. 2009; Sowers et al. 1993a, b) . Two crystal structures of the C-terminal DUF1608 domain of M. acetivorans MA0829 were reported at 2.3 and 2.36 Å resolutions, respectively (Arbing et al. 2012; PDB codes: 3U2H and 3U2G; Figure 2.6). The mature MA0829 (671 residues) consists of a signal sequence followed by an N-terminal DUF1608 domain (DUF1608 (NTR)/MA0829 NTR) and a C-terminal DUF1608 domain (DUF1608 (CTR)/MA0829 CTR; Arbing et al. 2012; Bateman et al. 2010) . The crystal structure of MA0829 CTR exhibits two structurally related domains (rmsd: 3.1 Å, sequence identity: 3 %) comprising two antiparallel β-sheets opposed in a β-sandwich fold. Both the N- and C-terminal ends are located nearer to the domain I. The polypeptide chain crosses domain I to domain II and folds back to form the bipartite CTR structure and, during the process, creates a connector domain (~ 40-amino-acid, three-stranded β-sheet). Domain II has an additional three-stranded β-sheet observed to be attached to one of the β-sheets.
The MA0829 CTR possesses a hexagonal lattice and is used to model the basic repeating unit of the S-layer (Figure 2.6b). Around a threefold crystallographic axis, three CTR dimers (trimeric unit) are arranged to form the repeating unit. Using this trimeric unit, a translation in two dimensions creates a sheet with sixfold symmetry that is similar to the architecture of hexagonal S-layers visualized by EM for archaea (Figure 2.6c). The 2D S-layer lattice is stabilized by a series of extensive intermolecular interactions including hydrogen bonds, salt bridges and van der Waals interactions. Thus, modelled S-layer reveals pores are classified as P or primary pores (pores on sixfold axis; ~ 13 Å in diameter), T or trimer pores (pores on the three-fold axis at the center of trimer; ~ 8 Å in diameter) and A or asymmetric pores (pores at the interface of two trimers; 5 × 14 Å in dimensions). All these pores would assist in the passage of small molecules like nutrients and due to large pore size; P pores may assist in passage of siderophores, oligos and lipids.
2.4.6 Structures of SbsB and SbsC from G. stearothermophilus
G. stearothermophilus is a Gram-positive, rod-shaped thermophilic bacterium encased in a rigid cell wall composed of peptidoglycan (A1-γ chemotype) and an SCWP composed of 2,3-diacetamido mannosamine uronic acid, N-acetyl glucosamine and glucose in wild-type strains; or N-acetyl glucosamine, N-acetyl mannosamine and pyruvic acid in variant strains (Egelseer et al. 1998; Mader et al. 2004; Schaffer et al. 1999) . Five different SLPs have been identified in various strains of G. stearothermophilus, namely SbsA (wild-type strain PV72/p6, hexagonal lattice type; Kuen et al. 1994) , SbsB (oxygen-induced variant strain PV72/p2, oblique lattice type; Scholz et al. 2001) , SbsC (strain ATCC 12980, oblique lattice type; Jarosch et al. 2000, 2001) , SbsD (strain ATCC 12980 under high temperatures of 67 °C, oblique lattice type; Egelseer et al. 2001) and SgsE (strain NRS 2004/3a, oblique lattice type; Schaffer et al. 2002; Pavkov et al. 2011) . The N-terminal ends of SbsA, SbsC, SbsD and SgsE share higher sequence similarity and are capable of binding to the same SCWP (Egelseer et al. 2001; Pavkov et al. 2011; Schaffer et al. 2002) . The C-terminal crystallization domains of these SLPs show little sequence conservation, except for SbsD and SgsE which share 94 % sequence identity. SbsB shows an overall low sequence similarity with SbsA, SbsC, SbsD and SgsE.
A partial crystal structure of SbsC was reported at 2.4 Å resolution, corresponding to a C-terminal truncation mutant spanning the first 412 residues of the mature protein (rSbsC(31–443); PDB code: 2RA1; Pavkov et al. 2008; Figure 2.7). The structure reveals an overall architecture that is composed of an α-helical N-terminal domain corresponding to the cell-wall-anchoring domain, followed by a string of consecutive β-sandwich domains that are part of the C-terminal crystallization domain. The N-terminal domain of rSbsC(31–443) (domain I, residues 32–260) comprises seven α-helices organized into three left-handed antiparallel three-helix bundles that give rise to a unique “banana-shaped” conformation (Figure 2.7a). The interface of the first and second triple-helical bundle is interspersed with aromatic residues, and the presence of kinks in α4 and α5 in the second triple-helical bundle due to larger residues (Tyr 130, Arg 167, Arg 184 and Lys188) results in an increased exposure of the hydrophobic core region between the α-helices. The third triple-helical bundle exhibits a canonical coiled-coil structure (Lupas and Gruber 2005; Pavkov et al. 2008) . Surface plasmon resonance (SPR) studies revealed that specific binding of SCWP to SbsC is facilitated by the N-terminal end of the SbsC (rSbsC(31–270); Ferner-Ortner-Bleckmann et al. 2009) . Binding experiments with SCWP and further biochemical analyses reveal that the N-terminal end stabilizes upon binding to the SCWP and this is ascribed to the presence of regularly spaced, positively charged residues on the putative ligand-binding surface matching the negatively charged residues on elongated SCWP. In the crystallization domain, domains II (residues 261–331; two anti-parallel β-sheets) and III (residues 332–443; three anti-parallel β-sheets) adopt immunoglobulin-like (Ig-like) folds (Figure 2.4b) and are believed to take part in intermolecular domain–domain interactions in the S-layer (Luo et al. 2000; Pavkov et al. 2008) . The crystal structure of a larger SbsC fragment (rSbsC(31–844)) reveals an elongated molecule with an additional three compact β-domains compared to rSbsC(31–443). Although poor electron density prevented model building for domains IV–VI, these can be seen to form a ring-like structure together with domains II and III (Figure 2.7c).
Subsequently, the crystal structure of full-length SbsB (residues 32–920 in mature SbsB) was reported at 2.4 Å (Figure 2.8) and provided the first structural insight into a full-length SLP (Baranova et al. 2012) . To avoid the formation of S-layer self-assembly products, the protein was crystallized in complex with a single-domain antibody, or nanobody (SbsB32–920:NbKB6). The mature SbsB comprises a functional N-terminal cell-wall-anchoring region with three SLH motifs and a C-terminal crystallization domain that can assemble in an oblique (p1 symmetry) 2D lattice with unit cell vectors a = 104 Å, b = 79 Å and base angle γ = 81° (Moll et al. 2002) . The crystal structure reveals seven domains of which the N-terminal domain (domain I, 32–201 residues) is not resolved due to poor electron density. The domains II–VII are arranged in a φ-shaped, disk-like quaternary structure that corresponds to the self-assembling crystallization domain of the protein (dimensions of 70 Å × 110 Å × 35 Å; Figure 2.8b). This arrangement creates a single plane (domains IV to VII) with an annular structure that encloses a cavity of ~ 24 Å in diameter. A nine-residue linker between domains IV and III (present at the rim of the ring) passes across the cavity and the domain II protruding away from the ring structure gives the final φ-shaped structure. The domains II and III correspond to C1-type Ig folds, domains IV, V and VII correspond to I-type Ig fold, whereas the domain VII is a mixed fold with an Ig-like β-sandwich core along with a random-coil subdomain. The modular build-up of the crystallization domain by consecutive Ig folds and their supramolecular organization is highly reminiscent of the ring-like architecture formed by domains II–VI in SbsC. Domains II and III in SbsB are structurally equivalent to those in SbsC, with rmsd values of 1.35 and 2.57 Å, respectively, suggesting that despite the low sequence identity SbsB and SbsC form a structurally similar SLP.
Chemically denatured SbsB refolds rapidly in the absence of a chaotropic agent and in the presence of Ca2+ ions (Runzler et al. 2004) . The SbsB crystals structure reveals four bound Ca2+ ions that mediate inter-domain and intra-domain contacts (CA1 to CA3 ions; through a pentagonal bipyramidal coordination and CA4 ion through an octahedral coordination). CA1 is shared by domains IV and VII and is coordinated with a water molecule (W2063) along with the residues Gln 406, Thr 440 from domain IV and Asp 779, Asp 781 in domain VII. CA2 ion is at the interface of domains V and VI (located closer to domain V) and exhibits coordination with a water molecule (W2125) and residues from domain V (Asp517, Gln 518, Asp 592, Asn 624 and Val 625). CA3 ion shares coordination with two water molecules (W2153, W2154) and residues from domain VI (Ala 646, Thr 649, Ser 651 and Arg 654). Finally, CA4 ion is coordinated by the residues from domain VII (Glu 784, Gly 822, Asn 824, Asp 835, Glu 836 and Glu 837; Baranova et al. 2012) . Circular dichroism and SAXS analysis showed the importance of Ca2+ ions for the adoption of the SbsB quaternary structure. Ethylenediaminetetraacetic acid (EDTA)-treated SbsB samples retained the secondary structure in the individual β-sandwich domains, but lost the φ-shaped quaternary structure and instead resulted in a dynamic beads-on-a-string architecture. Cryo-EM and chemical cross-linking data from SbsB S-layers showed that the φ-shaped conformers form the self-assembling species, which are juxtaposed in the plane of the disk-like crystallization domain (Baranova et al. 2012) . The Ca2+-triggered conformational reorganization of domains II–VII primes the SbsB protomers for self-assembly by prepositioning the interactive surfaces in the β-sandwich domains. In this way, a semi-porous monolayer is formed (Figure 2.8c). Intermolecular contacts are formed by domains IV and VII, and domains IV and II. The latter ducks underneath an adjacent monomer and connects the crystallization domain to the N-terminal cell wall attachment domain. Interestingly, the domain II position was not rigid throughout the recombinant SbsB S-layer, possibly providing a conformational hinge between the SLP lattice and the nonuniform topology of the supporting cell wall.
2.5 Applications
The intrinsic capability of SLPs to self-assemble into semi-porous monolayers with defined structural properties has raised interest from the fields of material sciences and biotechnology (Ilk et al. 2011; Pavkov et al. 2011) . Ordered S-layer fragments can be extracted from their bacterial hosts, or can be produced by (re)crystallizing the isolated proteins directly in solution or more commonly at liquid–air, liquid–solid and liquid–lipid interfaces (reviewed in Pum et al. 2013) . Due to the presence of pores with identical shape and morphology, S-layers have been utilized as efficient isoporous ultrafiltration membranes (Sara and Sleytr 1996; Sleytr et al. 1997, 1999) . Such S-layer ultrafiltration membranes (SUMs) have been created by the deposition of S-layer assembly products over microfiltration membranes (Weigert and Sara 1995, 1996) . Integration of functional groups in S-layers has also enabled a broad spectrum of applications owing to the topological alignment of the introduced functionalities. In affinity matrices, this property helps in binding molecules with higher specificity and affinity (Sleytr et al. 1999) . S-layer microparticles or SMPs are cell wall fragments with S-layers having both the outer and inner S-layer cross-linked by glutaraldehyde (Breitwieser et al. 1996; Kupcu et al. 1995, 1996) . These SMPs are used as affinity particles for covalent attachment of biologically active macromolecules (Kupcu et al. 1995) . Monoclonal antibodies from serum have been isolated and purified using SMPs which are covalently linked to protein A that allows identification of the Fc portion of the majority of mammalian antibodies (Weiner et al. 1994) . Affinity microparticles of 1–2-µm size have been prepared from hexagonal S-layer lattice of Clostridium thermohydrosulfuricum L111–69, and used to isolate and purify IgG from artificial IgG–human serum albumin mixtures by affinity cross-flow filtration (Weiner et al. 1994a, b). Furthermore, S-layers have been considered for S-layer vaccine technology, allowing a high display density of recombinantly or chemically introduced epitopes. Smith et al. (1993) observed that when tumour-associated glycans (T- or Lewis Y antigen) are coupled with glutaraldehyde-cross-linked SLPs of Gram-positive bacteria, they gave rise to a stronger hapten-specific delayed-type hypersensitivity (DTH) response (Smith et al. (1993). The availability of a number of immunologically non-cross-reactive S-layers that can be utilized as combined carrier/adjuvants system has been suggested to provide molecular tools for anti-allergic immunotherapy (Jahn-Schmid et al. 1996, 1997) and for therapeutic cancer vaccines (Smith et al. 1993). Recrystallizing S-layer subunits on liposomes and cross-linking with glutaraldehyde produced closed biomimetic structures that resemble archaeal cells (Kupcu et al. 1995, 1998) . In particular, the S-layer subunits of B. stearothermophilus PV72/p2 has been recrystallized on liposomes and used as a matrix for binding and entrapping functional molecules (Mader et al. 1999, 2000) .
In material sciences, S-layers have been used as patterning resists for the deposition of inorganic materials into regular 2D arrays (Allred et al. 2008; Shenton et al. 1997) The SLPs of G stearothermophilus have also been recrystallized using gold colloids, cross-linked to each other and applied as electrochemical sensing tools by wrapping them around single-walled carbon nanotubes (CNT) allowing novel approaches in nanoelectronic biosensor applications (Park et al. 2011) . By recrystallizing the SbpA (B. sphaericus) on amorphous and semicrystalline polyactide derivatives, new bio-supramolecular structures could be fabricated (Lejardi et al. 2013) . In the creation of biomimetic sensors that involve sensitive microelectronic devices attached to biological systems, an attached lipid membrane intact with integral proteins (natural/designed) offers a good platform to perform scientific studies. These lipid membranes can be supported by S-layer assembled on metal or semiconductor surfaces that can be used as a separating layer in order to retain the fluidity and stability of the lipid membrane, thereby providing an ion reservoir and required space for the proteins within the membrane (Schuster et al. 2004) .
Available examples demonstrate that the scaffolding and organizing properties of S-layers can be employed to devise distinct biomimetic structures in a wide range of applications. It is expected that the increasing understanding of the SLP structure and dynamics will further enhance their application.
References
Al-Karadaghi S, Wang DN, Hovmoller S (1988) 3-Dimensional structure of the crystalline surface-layer from Aeromonas hydrophila. J Ultra Mol Struct R 101:92–97
Albers SV, Meyer BH (2011) The archaeal cell envelope. Nat Rev Microbiol 9:414–426
Allred DB, Cheng A, Sarikaya M et al (2008) Three-dimensional architecture of inorganic nanoarrays electrodeposited through a surface-layer protein mask. Nano Lett 8:1434–1438
Altman E, Brisson JR, Messner P et al (1990) Chemical characterization of the regularly arranged surface layer glycoprotein of Clostridium thermosaccharolyticum D120–70. Eur J Biochem 188:73–82
Altman E, Schaffer C, Brisson JR et al (1996) Isolation and characterization of an amino sugar-rich glycopeptide from the surface layer glycoprotein of Thermoanaerobacterium thermosaccharolyticum E207–71. Carbohydr Res 295:245–253
Amat F, Comolli LR, Nomellini JF et al (2010) Analysis of the intact surface layer of caulobacter crescentus by cryo-electron tomography. J Bacteriol 192:5855–5865
Anzengruber J, Pabst M, Neumann L et al (2013) Protein O-glucosylation in Lactobacillus buchneri. Glycoconj J 32:117–131
Arbing MA, Chan S, Shin A et al (2012) Structure of the surface layer of the methanogenic archaean Methanosarcina acetivorans. Proc Natl Acad Sci U S A 109:11812–11817
Bahl H, Scholz H, Bayan N et al (1997) Molecular biology of S-layers. FEMS Microbiol Rev 20:47–98
Baranova E, Fronzes R, Garcia-Pino A et al (2012) SbsB structure and lattice reconstruction unveil Ca2+ triggered S-layer assembly. Nature 487:119–122
Bateman A, Coggill P, Finn RD (2010) DUFs: families in search of function. Acta Crystallogr F 66:1148–1152
Baumeister W, Lembcke G (1992) Structural features of archaebacterial cell envelopes. J Bioenerg Biomembr 24:567–575
Baumeister W, Barth M, Hegerl R et al (1986) 3-dimensional structure of the regular surface-layer (Hpi Layer) of Deinococcus radiodurans. J Mol Biol 187:241–253
Beveridge TJ (1993) Current trends and future-prospects in prokaryotic envelope research—a microscopists view. J Appl Bacteriol 74:S143–S153
Beveridge TJ, Stewart M, Doyle RJ et al (1985) Unusual stability of the Methanospirillum hungatei sheath. J Bacteriol 162:728–737
Beveridge TJ, Koval SF, Sleytr UB et al (1993) Advances in bacterial paracrystalline surface-layers. Nato Adv Sci Inst Se 252:323–327
Blackford BL, Xu W, Jericho MH et al (1994) Direct observation by scanning-tunneling-microscopy of the 2-dimensional lattice structure of the S-layer sheath of the archaeobacterium Methanospirillum hungatei Gp1. Scanning Microsc 8:507–512
Boot HJ, Kolen CP, Pouwels PH (1995) Identification, cloning, and nucleotide sequence of a silent S-layer protein gene of Lactobacillus acidophilus ATCC 4356 which has extensive similarity with the S-layer protein gene of this species. J Bacteriol 177:7222–7230
Breitwieser A, Kupcu S, Howorka S et al (1996) 2-D protein crystals as an immobilization matrix for producing reaction zones in dipstick-style immunoassays. BioTechniques 21:918–925
Brockl G, Behr M, Fabry S et al (1991) Analysis and nucleotide sequence of the genes encoding the surface-layer glycoproteins of the hyperthermophilic methanogens Methanothermus fervidus and Methanothermus sociabilis. Eur J Biochem 199:147–152
Calabi E, Ward S, Wren B et al (2001) Molecular characterization of the surface layer proteins from Clostridium difficile. Mol Microbiol 40:1187–1199
Calabi E, Calabi F, Phillips AD et al (2002) Binding of Clostridium difficile surface layer proteins to gastrointestinal tissues. Infect Immun 70:5770–5778
Callegari ML, Riboli B, Sanders JW et al (1998) The S-layer gene of Lactobacillus helveticus CNRZ 892: cloning, sequence and heterologous expression. Microbiology 144(Part 3):719–726
Chami M, Bayan N, Peyret JL et al (1997) The S-layer protein of Corynebacterium glutamicum is anchored to the cell wall by its C-terminal hydrophobic domain. Mol Microbiol 23:483–492
Doig P, Emody L, Trust TJ (1992) Binding of laminin and fibronectin by the trypsin-resistant major structural domain of the crystalline virulence surface array protein of Aeromonas salmonicida. J Biol Chem 267:43–49
Dooley JSG, Trust TJ (1988) Surface protein-composition of Aeromonas hydrophila strains virulent for fish—identification of a surface array protein. J Bacteriol 170:499–506
Dooley JSG, Engelhardt H, Baumeister W et al (1989) 3-dimensional structure of an open form of the surface-layer from the fish pathogen Aeromonas salmonicida. J Bacteriol 171:190–197
Dufrene YF (2001) Application of atomic force microscopy to microbial surfaces: from reconstituted cell surface layers to living cells. Micron 32:153–165
Dworkin J, Blaser MJ (1997) Molecular mechanisms of Campylobacter fetus surface layer protein expression. Mol Microbiol 26:433–440
Egelseer E, Schocher I, Sara M et al (1995) The S-layer from Bacillus stearothermophilus DSM 2358 functions as an adhesion site for a high-molecular-weight amylase. J Bacteriol 177:1444–1451
Egelseer EM, Schocher I, Sleytr UB et al (1996) Evidence that an N-terminal S-layer protein fragment triggers the release of a cell-associated high-molecular-weight amylase in Bacillus stearothermophilus ATCC 12980. J Bacteriol 178:5602–5609
Egelseer EM, Leitner K, Jarosch M et al (1998) The S-layer proteins of two Bacillus stearothermophilus wild-type strains are bound via their N-terminal region to a secondary cell wall polymer of identical chemical composition. J Bacteriol 180:1488–1495
Egelseer EM, Danhorn T, Pleschberger M et al (2001) Characterization of an S-layer glycoprotein produced in the course of S-layer variation of Bacillus stearothermophilus ATCC 12980 and sequencing and cloning of the sbsD gene encoding the protein moiety. Arch Microbiol 177:70–80
Eichler J, Adams MWW (2005) Posttranslational protein modification in Archaea. Microbiol Mol Biol R 69:393
Engelhardt H (2007a) Are S-layers exoskeletons? The basic function of protein surface layers revisited. J Struct Biol 160:115–124
Engelhardt H (2007b) Mechanism of osmoprotection by archaeal S-layers: a theoretical study. J Struct Biol 160:190–199
Engelhardt H, Peters J (1998) Structural research on surface layers: a focus on stability, surface layer homology domains, and surface layer cell wall interactions. J Struct Biol 124:276–302
Ethordic A, Egelseer EM, Tesarz M et al (2012) Crystallization of domains involved in self-assembly of the S-layer protein SbsC. Acta Crystallogr F 68:1511–1514
Etienne-Toumelin I, Sirard JC, Duflot E et al (1995) Characterization of the Bacillus anthracis S-layer: cloning and sequencing of the structural gene. J Bacteriol 177:614–620
Fagan RP, Fairweather NF (2014) Biogenesis and functions of bacterial S-layers. Nat Rev Microbiol 12:211–222
Fagan RP, Albesa-Jove D, Qazi O et al (2009) Structural insights into the molecular organization of the S-layer from Clostridium difficile. Mol Microbiol 71:1308–1322
Ferner-Ortner-Bleckmann J, Huber-Gries C, Pavkov T et al (2009) The high-molecular-mass amylase (HMMA) of Geobacillus stearothermophilus ATCC 12,980 interacts with the cell wall components by virtue of three specific binding regions. Mol Microbiol 72:1448–1461
Firtel M, Southam G, Harauz G et al (1994a) The organization of the paracrystalline multilayered spacer-plugs of Methanospirillum hungatei. J Struct Biol 112:160–171
Firtel M, Xu W, Southam G et al (1994b) Tip-induced displacement and imaging of a multilayered bacterial structure by scanning-tunneling-microscopy. Ultramicroscopy 55:113–119
Ford MJ, Nomellini JF, Smit J (2007) S-layer anchoring and localization of an S-layer-associated protease in Caulobacter crescentus. J Bacteriol 189:2226–2237
Francoleon DR, Boontheung P, Yang Y et al (2009) S-layer, surface-accessible, and concanavalin A binding proteins of Methanosarcina acetivorans and Methanosarcina mazei. J Proteome Res 8:1972–1982
Galagan JE, Nusbaum C, Roy A et al (2002) The genome of M. acetivorans reveals extensive metabolic and physiological diversity. Genome Res 12:532–542
Garduno RA, Lee EJY, Kay WW (1992a) S-layer-mediated association of Aeromonas salmonicida with murine macrophages. Infect Immun 60:4373–4382
Garduno RA, Phipps BM, Baumeister W et al (1992b) Novel structural patterns in divalent cation-depleted surface-layers of Aeromonas salmonicida. J Struct Biol 109:184–195
Gilmour R, Messner P, Guffanti AA et al (2000) Two-dimensional gel electrophoresis analyses of pH-dependent protein expression in facultatively alkaliphilic Bacillus pseudofirmus OF4 lead to characterization of an S-layer protein with a role in alkaliphily. J Bacteriol 182:5969–5981
Graham LL, Beveridge TJ (1994) Structural differentiation of the Bacillus subtilis 168 cell-wall. J Bacteriol 176:1413–1421
Gyorvary ES, Stein O, Pum D et al (2003) Self-assembly and recrystallization of bacterial S-layer proteins at silicon supports imaged in real time by atomic force microscopy. J Microsc 212:300–306
Horejs C, Pum D, Sleytr UB et al (2008) Structure prediction of an S-layer protein by the mean force method. J Chem Phys 128:065106
Horejs C, Pum D, Sleytr UB et al (2010) Surface layer protein characterization by small angle x-ray scattering and a fractal mean force concept: from protein structure to nanodisk assemblies. J Chem Phys 133:175102
Horejs C, Mitra MK, Pum D et al (2011) Monte Carlo study of the molecular mechanisms of surface-layer protein self-assembly. J Chem Phys 134:125103
Houwink AL (1953) A macromolecular mono-layer in the cell wall of Spirillum spec. Biochimica et Biophysica Acta 10:360–366
Houwink AL (1956) Flagella, gas vacuoles and cell-wall structure in Halobacterium halobium—an electron microscope study. J Gen Microbiol 15:146–150
Howorka S, Sara M, Wang Y et al (2000) Surface-accessible residues in the monomeric and assembled forms of a bacterial surface layer protein. J Biol Chem 275:37876–37886
Huber C, Ilk N, Runzler D et al (2005) The three S-layer-like homology motifs of the S-layer protein SbpA of Bacillus sphaericus CCM 2177 are not sufficient for binding to the pyruvylated secondary cell wall polymer. Mol Microbiol 55:197–205
Ilk N, Egelseer EM, Sleytr UB (2011) S-layer fusion proteins–construction principles and applications. Curr Opin Biotechnol 22:824–831
Jahn-Schmid B, Messner P, Unger FM et al (1996) Toward selective elicitation of TH1-controlled vaccination responses: vaccine applications of bacterial surface layer proteins. J Biotechnol 44:225–231
Jahn-Schmid B, Siemann U, Zenker A et al (1997) Bet v 1, the major birch pollen allergen, conjugated to crystalline bacterial cell surface proteins, expands allergen-specific T cells of the Th1/Th0 phenotype in vitro by induction of IL-12. Int Immunol 9:1867–1874
Janesch B, Koerdt A, Messner P et al (2013a) The S-layer homology domain-containing protein SlhA from Paenibacillus alvei CCM 2051(T) is important for swarming and biofilm formation. PloS One 8:e76566
Janesch B, Messner P, Schaffer C (2013b) Are the surface layer homology domains essential for cell surface display and glycosylation of the S-layer protein from Paenibacillus alvei CCM 2051T? J Bacteriol 195:565–575
Jarosch M, Egelseer EM, Mattanovich D et al (2000) S-layer gene sbsC of Bacillus stearothermophilus ATCC 12980: molecular characterization and heterologous expression in Escherichia coli. Microbiology 146(Part 2):273–281
Jarosch M, Egelseer EM, Huber C et al (2001) Analysis of the structure-function relationship of the S-layer protein SbsC of Bacillus stearothermophilus ATCC 12980 by producing truncated forms. Microbiology 147:1353–1363
Jing H, Takagi J, Liu JH et al (2002) Archaeal surface layer proteins contain beta propeller, PKD, and beta helix domains and are related to metazoan cell surface proteins. Structure 10:1453–1464
Kadurugamuwa JL, Mayer A, Messner P et al (1998) S-layered Aneurinibacillus and Bacillus spp. are susceptible to the lytic action of Pseudomonas aeruginosa membrane vesicles. J Bacteriol 180:2306–2311
Karrasch S, Hegerl R, Hoh JH et al (1994) Atomic force microscopy produces faithful high-resolution images of protein surfaces in an aqueous environment. Proc Natl Acad Sci U S A 91:836–838
Karrenberg FH, Wildhaber I, Baumeister W (1987) Surface-structure variants in Deinococcus radiodurans. Curr Microbiol 16:15–20
Kern J, Schneewind O (2010) BslA, the S-layer adhesin of B. anthracis, is a virulence factor for anthrax pathogenesis. Mol Microbiol 75:324–332
Kern J, Wilton R, Zhang RG et al (2011) Structure of surface layer homology (SLH) domains from Bacillus anthracis surface array protein. J Biol Chem 286:26042–26049
Kinns H, Badelt-Lichtblau H, Egelseer EM et al (2010) Identifying assembly-inhibiting and assembly-tolerant sites in the SbsB S-layer protein from Geobacillus stearothermophilus. J Mol Biol 395:742–753
Konisky J, Lynn D, Hoppert M et al (1994) Identification of the Methanococcus voltae S-layer structural gene. J Bacteriol 176:1790–1792
Koval SF (1988) Paracrystalline protein surface arrays on bacteria. Can J Microbiol 34:407–414
Koval SF, Hynes SH (1991) Effect of paracrystalline protein surface-layers on predation by Bdellovibrio bacteriovorus. J Bacteriol 173:2244–2249
Koval SF, Bayer ME (1997) Bacterial capsules: no barrier against Bdellovibrio. Microbiology (UK) 143:749–753
Kuen B, Sleytr UB, Lubitz W (1994) Sequence analysis of the sbsA gene encoding the 130-kDa surface-layer protein of Bacillus stearothermophilus strain PV72. Gene 145:115–120
Kupcu Z, Marz L, Messner P et al (1984) Evidence for the glycoprotein nature of the crystalline cell wall surface layer of Bacillus stearothermophilus strain NRS2004/3a. FEBS Lett 173:185–190
Kupcu S, Sara M, Sleytr UB (1995) Liposomes coated with crystalline bacterial cells surface protein (S-layer) as immobilization structures for macromolecules. Biochimica et Biophysica Acta 1235:263–269
Kupcu S, Sleytr UB, Sara M (1996) Two-dimensional paracrystalline glycoprotein S-layers as a novel matrix for the immobilization of human IgG and their use as microparticles in immunoassays. J Immunol Methods 196:73–84
Kupcu S, Lohner K, Mader C et al (1998) Microcalorimetric study on the phase behaviour of S-layer coated liposomes. Mol Membr Biol 15:69–74
Lechner J, Sumper M (1987) The primary structure of a prokaryotic glycoprotein—cloning and sequencing of the cell-surface glycoprotein gene of Halobacteria. J Biol Chem 262:9724–9729
Lechner J, Wieland F, Sumper M (1986) Sulfated dolicholphosphate oligosaccharides are transiently methylated during biosynthesis of Halobacterial glycoproteins. Syst Appl Microbiol 7:286–292
Leibovitz E, Ohayon H, Gounon P et al (1997) Characterization and subcellular localization of the Clostridium thermocellum scaffoldin dockerin binding protein SdbA. J Bacteriol 179:2519–2523
Lejardi A, Lopez AE, Sarasua JR et al (2013) Making novel bio-interfaces through bacterial protein recrystallization on biocompatible polylactide derivative films. J Chem Phys 139:121903
Lighezan L, Georgieva R, Neagu A (2012) A study of the thermal denaturation of the S-layer protein from Lactobacillus salivarius. Phys Scripta 86
Lister TE, Pinhero PJ (2001) In vivo atomic force microscopy of surface proteins on Deinococcus radiodurans. Langmuir 17:2624–2628
Liu SY, Gherardini FC, Matuschek M et al (1996) Cloning, sequencing, and expression of the gene encoding a large S-layer-associated endoxylanase from Thermoanaerobacterium sp strain JW/SL-YS 485 in Escherichia coli. J Bacteriol 178:1539–1547
Luo Y, Frey EA, Pfuetzner RA et al (2000) Crystal structure of enteropathogenic Escherichia coli intimin-receptor complex. Nature 405:1073–1077
Lupas AN, Gruber M (2005) The structure of alpha-helical coiled coils. Adv Protein Chem 70:37–78
Mader C, Kupcu S, Sara M et al (1999) Stabilizing effect of an S-layer on liposomes towards thermal or mechanical stress. Biochimica et Biophysica Acta 1418:106–116
Mader C, Kupcu S, Sleytr UB et al (2000) S-layer-coated liposomes as a versatile system for entrapping and binding target molecules. Biochimica et Biophysica Acta 1463:142–150
Mader C, Huber C, Moll D et al (2004) Interaction of the crystalline bacterial cell surface layer protein SbsB and the secondary cell wall polymer of Geobacillus stearothermophilus PV72 assessed by real-time surface plasmon resonance biosensor technology. J Bacteriol 186:1758–1768
Mayerhofer LE, Macario AJL, Demacario EC (1992) Lamina, a novel multicellular form of Methanosarcina mazei S-6. J Bacteriol 174:309–314
Mescher MF, Strominger JL (1976) Purification and characterization of a prokaryotic glycoprotein from cell-envelope of Halobacterium salinarium. J Biol Chem 251:2005–2014
Mesnage S, Tosi-Couture E, Mock M et al (1997) Molecular characterization of the Bacillus anthracis main S-layer component: evidence that it is the major cell-associated antigen. Mol Microbiol 23:1147–1155
Mesnage S, Fontaine T, Mignot T et al (2000) Bacterial SLH domain proteins are non-covalently anchored to the cell surface via a conserved mechanism involving wall polysaccharide pyruvylation. EMBO J 19:4473–4484
Messner P (1997) Bacterial glycoproteins. Glycoconj J 14:3–11
Messner P, Sleytr UB (1992) Crystalline bacterial cell-surface layers. Adv Microb Physiol 33:213–275
Messner P, Pum D, Sara M et al (1986a) Ultrastructure of the cell envelope of the archaebacteria Thermoproteus tenax and Thermoproteus neutrophilus. J Bacteriol 166:1046–1054
Messner P, Pum D, Sleytr UB (1986b) Characterization of the ultrastructure and the self-assembly of the surface layer of Bacillus stearothermophilus strain NRS 2004/3a. J Ultra Mol Struct R 97:73–88
Messner P, Allmaier G, Schaffer C et al (1997) Biochemistry of S-layers. FEMS Microbiol Rev 20:25–46
Messner P, Steiner K, Zarschler K et al (2008) S-layer nanoglycobiology of bacteria. Carbohydr Res 343:1934–1951
Moll D, Huber C, Schlegel B et al (2002) S-layer-streptavidin fusion proteins as template for nanopatterned molecular arrays. Proc Natl Acad Sci U S A 99:14646–14651
Muller DJ, Baumeister W, Engel A (1996) Conformational change of the hexagonally packed intermediate layer of Deinococcus radiodurans monitored by atomic force microscopy. J Bacteriol 178:3025–3030
Muller DJ, Baumeister W, Engel A (1999) Controlled unzipping of a bacterial surface layer with atomic force microscopy. Proc Natl Acad Sci U S A 96:13170–13174
Murray RGE, Dooley JSG, Whippey PW et al (1988) Structure of an S-layer on a pathogenic strain of Aeromonas hydrophila. J Bacteriol 170:2625–2630
Noonan B, Trust TJ (1995) Molecular analysis of an a-protein secretion mutant of Aeromonas salmonicida reveals a surface layer-specific protein secretion pathway. J Mol Biol 248:316–327
Norville JE, Kelly DF, Knight TF et al (2007) 7 angstrom projection map of the S-layer protein sbpA obtained with trehalose-embedded monolayer crystals. J Struct Biol 160:313–323
Oh YJ, Sekot G, Duman M et al (2013) Characterizing the S-layer structure and anti-S-layer antibody recognition on intact Tannerella forsythia cells by scanning probe microscopy and small angle X-ray scattering. JMR 26:542–549
Ohnesorge F, Heckl WM, Haberle W et al (1992) Scanning force microscopy studies of the S-layers from Bacillus coagulans E38–66, Bacillus sphaericus CCM2177 and of an antibody binding process. Ultramicroscopy 42–44(Part B):1236–1242
Park TJ, Lee SJ, Park JP et al (2011) Characterization of a bacterial self-assembly surface layer protein and its application as an electrical nanobiosensor. J Nanosci Nanotechnol 11:402–407
Pavkov T, Oberer M, Egelseer EM et al (2003) Crystallization and preliminary structure determination of the C-terminal truncated domain of the S-layer protein SbsC. Acta Crystallogr D 59:1466–1468
Pavkov T, Egelseer EM, Tesarz M et al (2008) The structure and binding behavior of the bacterial cell surface layer protein SbsC. Structure 16:1226–1237
Pavkov T, Howorka S, Keller W (2011) The structure of bacterial S-layer proteins. In Howorka S (ed) Progress in molecular biology and translational science, vol 103. Academic Press, Burlington, pp 73–130
Peters J, Peters M, Lottspeich F et al (1987) Nucleotide-sequence analysis of the gene encoding the Deinococcus radiodurans surface protein, derived amino-acid-sequence, and complementary protein chemical studies. J Bacteriol 169:5216–5223
Peters J, Nitsch M, Kuhlmorgen B et al (1995) Tetrabrachion: a filamentous archaebacterial surface protein assembly of unusual structure and extreme stability. J Mol Biol 245:385–401
Peters J, Baumeister W, Lupas A (1996) Hyperthermostable surface layer protein tetrabrachion from the archaebacterium Staphylothermus marinus: evidence for the presence of a right-handed coiled coil derived from the primary structure. J Mol Biol 257:1031–1041
Petersen BO, Sara M, Mader C et al (2008) Structural characterization of the acid-degraded secondary cell wall polymer of Geobacillus stearothermophilus PV72/p2. Carbohydr Res 343:1346–1358
Peyfoon E, Meyer B, Hitchen PG et al (2010) The S-layer glycoprotein of the crenarchaeote Sulfolobus acidocaldarius is glycosylated at multiple sites with chitobiose-linked N-glycans. Archaea
Peyret JL, Bayan N, Joliff G et al (1993) Characterization of the cspB gene encoding PS2, an ordered surface-layer protein in Corynebacterium glutamicum. Mol Microbiol 9:97–109
Posch G, Pabst M, Brecker L et al (2011) Characterization and scope of S-layer protein O-glycosylation in Tannerella forsythia. J Biol Chem 286:38714–38724
Posch G, Andrukhov O, Vinogradov E et al (2013) Structure and immunogenicity of the rough-type lipopolysaccharide from the periodontal pathogen Tannerella forsythia. CVI 20:945–953
Poxton IR, Mccoubrey J, Blair G (2001) The pathogenicity of Clostridium difficile. Clin Microbiol Infect 7:421–427
Pum D, Sara M, Sleytr UB (1989) Structure, surface charge, and self-assembly of the S-layer lattice from Bacillus coagulans E38–66. J Bacteriol 171:5296–5303
Pum D, Messner P, Sleytr UB (1991) Role of the S layer in morphogenesis and cell division of the archaebacterium Methanocorpusculum sinense. J Bacteriol 173:6865–6873
Pum D, Toca-Herrera JL, Sleytr UB (2013) S-layer protein self-assembly. Int J Mol Sci 14:2484–2501
Rachel R, Jakubowski U, Tietz H et al (1986) Projected structure of the surface protein of Deinococcus radiodurans determined to 8 a resolution by cryomicroscopy. Ultramicroscopy 20:305–316
Runzler D, Huber C, Moll D et al (2004) Biophysical characterization of the entire bacterial surface layer protein SbsB and its two distinct functional domains. J Biol Chem 279:5207–5215
Sakakibara J, Nagano K, Murakami Y et al (2007) Loss of adherence ability to human gingival epithelial cells in S-layer protein-deficient mutants of Tannerella forsythensis. Microbiology 153:866–876
Sara M, Sleytr UB (1996) Biotechnology and biomimetic with crystalline bacterial cell surface layers (S-layers). Micron 27:141–156
Sara M, Sleytr UB (2000) S-Layer proteins. J Bacteriol 182:859–868
Sara M, Kalsner I, Sleytr UB (1988) Surface properties from the S-layer of Clostridium thermosaccharolyticum D120–70 and Clostridium thermohydrosulfuricum L111–69. Arch Microbiol 149:527–533
Sara M, Pum D, Sleytr UB (1992) Permeability and charge-dependent adsorption properties of the S-layer lattice from Bacillus coagulans E38–66. J Bacteriol 174:3487–3493
Sara M, Dekitsch C, Mayer HF et al (1998) Influence of the secondary cell wall polymer on the reassembly, recrystallization, and stability properties of the S-layer protein from Bacillus stearothermophilus PV72/p2. J Bacteriol 180:4146–4153
Schaffer C, Messner P (2004) Surface-layer glycoproteins: an example for the diversity of bacterial glycosylation with promising impacts on nanobiotechnology. Glycobiology 14:31R–42R
Schaffer C, Messner P (2005) The structure of secondary cell wall polymers: how Gram-positive bacteria stick their cell walls together. Microbiology 151:643–651
Schaffer C, Kahlig H, Christian R et al (1999) The diacetamidodideoxyuronic-acid-containing glycan chain of Bacillus stearothermophilus NRS 2004/3a represents the secondary cell-wall polymer of wild-type B. stearothermophilus strains. Microbiology 145(Part 7):1575–1583
Schaffer C, Wugeditsch T, Kahlig H et al (2002) The surface layer (S-layer) glycoprotein of Geobacillus stearothermophilus NRS 2004/3a. Analysis of its glycosylation. J Biol Chem 277:6230–6239
Scheuring S, Stahlberg H, Chami M et al (2002) Charting and unzipping the surface layer of Corynebacterium glutamicum with the atomic force microscope. Mol Microbiol 44:675–684
Scholz HC, Riedmann E, Witte A et al (2001) S-layer variation in Bacillus stearothermophilus PV72 is based on DNA rearrangements between the chromosome and the naturally occurring megaplasmids. J Bacteriol 183:1672–1679
Schultzelam S, Beveridge TJ (1994a) Nucleation of celestite and strontianite on a cyanobacterial S-layer. Appl Environ Microbiol 60:447–453
Schultzelam S, Beveridge TJ (1994b) Physicochemical characteristics of the mineral-forming S-layer from the Cyanobacterium synechococcus strain Gl24. Can J Microbiol 40:216–223
Schuster B, Gufler PC, Pum D et al (2004) S-layer proteins as supporting scaffoldings for functional lipid membranes. IEEE T Nanobiosci 3:16–21
Sekot G, Posch G, Messner P et al (2011) Potential of the Tannerella forsythia S-layer to delay the immune response. J Dent Res 90:109–114
Sekot G, Posch G, Oh YJ et al (2012) Analysis of the cell surface layer ultrastructure of the oral pathogen Tannerella forsythia. Arch Microbiol 194:525–539
Sekot G, Schuster D, Messner P et al (2013) Small-angle X-ray scattering for imaging of surface layers on intact bacteria in the native environment. J Bacteriol 195:2408–2414
Shenton W, Pum D, Sleytr UB et al (1997) Synthesis of cadmium sulphide superlattices using self-assembled bacterial S-layers. Nature 389:585–587
Shin SH, Comolli LR, Tscheliessnig R et al (2013) Self-assembly of “S-bilayers”, a step toward expanding the dimensionality of S-layer assemblies. ACS Nano 7:4946–4953
Simon P, Lichte H, Wahl R et al (2004) Electron holography of non-stained bacterial surface layer proteins. Bba-Biomembranes 1663:178–187
Sleytr UB, Thorne KJ (1976) Chemical characterization of the regularly arranged surface layers of Clostridium thermosaccharolyticum and Clostridium thermohydrosulfuricum. J Bacteriol 126:377–383
Sleytr UB, Beveridge TJ (1999) Bacterial S-layers. Trends Microbiol 7:253–260
Sleytr UB, Kocur M, Glauert AM et al (1973) A study by freeze-etching of the fine structure of Micrococcus radiodurans. Arch Mikrobiol 94:77–87
Sleytr UB, Sara M, Kupcu Z et al (1986) Structural and chemical characterization of S-layers of selected strains of Bacillus stearothermophilus and Desulfotomaculum nigrificans. Arch Microbiol 146:19–24
Sleytr UB, Messner P, Pum D et al (1993) Crystalline bacterial cell surface layers. Mol Microbiol 10:911–916
Sleytr UB, Pum D, Sara M (1997) Advances in S-layer nanotechnology and biomimetics. Adv Biophys 34:71–79
Sleytr UB, Messner P, Pum D et al (1999) Crystalline bacterial cell surface layers (S layers): from supramolecular cell structure to biomimetics and nanotechnology. Angew Chem Int Edit 38:1035–1054
Smit J, Engelhardt H, Volker S et al (1992) The S-layer of Caulobacter crescentus: three-dimensional image reconstruction and structure analysis by electron microscopy. J Bacteriol 174:6527–6538
Smith RH, Messner P, Lamontagne LR et al (1993) Induction of T-cell immunity to oligosaccharide antigens immobilized on crystalline bacterial surface layers (S-layers). Vaccine 11:919–924
Sowers KR, Boone JE, Gunsalus RP (1993a) Disaggregation of Methanosarcina spp. and growth as single cells at elevated osmolarity. Appl Environ Microbiol 59:3832–3839
Sowers KR, Thai TT, Gunsalus RP (1993b) Transcriptional regulation of the carbon monoxide dehydrogenase gene (cdhA) in Methanosarcina thermophila. J Biol Chem 268:23172–23178
Steindl C, Schaffer C, Wugeditsch T et al (2002) The first biantennary bacterial secondary cell wall polymer and its influence on S-layer glycoprotein assembly. Biochem J 368:483–494
Steiner K, Pohlentz G, Dreisewerd K et al (2006) New insights into the glycosylation of the surface layer protein SgsE from Geobacillus stearothermophilus NRS 2004/3a. J Bacteriol 188:7914–7921
Stetefeld J, Jenny M, Schulthess T et al (2000) Crystal structure of a naturally occurring parallel right-handed coiled coil tetramer. Nat Struct Biol 7:772–776
Stewart M, Beveridge TJ, Sprott GD (1985) Crystalline order to high resolution in the sheath of Methanospirillum hungatei: a cross-beta structure. J Mol Biol 183:509–515
Stewart M, Beveridge TJ, Trust TJ (1986) 2 Patterns in the Aeromonas salmonicida a-layer may reflect a structural transformation that alters permeability. J Bacteriol 166:120–127
Sumper M, Berg E, Mengele R et al (1990) Primary structure and glycosylation of the S-layer protein of Haloferax volcanii. J Bacteriol 172:7111–7118
Takeoka A, Takumi K, Koga T et al (1991) Purification and characterization of S-layer proteins from Clostridium difficile Gai-0714. J Gen Microbiol 137:261–267
Tang J, Ebner A, Badelt-Lichtblau H et al (2008) Recognition imaging and highly ordered molecular templating of bacterial S-layer nanoarrays containing affinity-tags. Nano Letters 8:4312–4319
Thompson SA, Shedd OL, Ray KC et al (1998) Campylobacter fetus surface layer proteins are transported by a type I secretion system. J Bacteriol 180:6450–6458
Thornley MJ, Glauert AM, Sleytr UB (1973) Isolation of outer membranes with an ordered array of surface subunits from Acinetobacter. J Bacteriol 114:1294–1308
Toca-Herrera JL, Moreno-Flores S, Friedmann J et al (2004) Chemical and thermal denaturation of crystalline bacterial S-layer proteins: an atomic force microscopy study. Microsc Res Tech 65:226–234
Trust TJ, Kostrzynska M, Emody L et al (1993) High-affinity binding of the basement-membrane protein collagen type-IV to the crystalline virulence surface protein array of Aeromonas salmonicida. Mol Microbiol 7:593–600
Tsuboi A, Uchihi R, Tabata R et al (1986) Characterization of the genes-coding for 2 major cell-wall proteins from protein-producing Bacillus brevis 47—complete nucleotide-sequence of the outer wall protein gene. J Bacteriol 168:365–373
Weigert S, Sara M (1995) Surface modification of an ultrafiltration membrane with crystalline-structure and studies on interactions with selected protein molecules. J Membrane Sci 106:147–159
Weigert S, Sara M (1996) Ultrafiltration membranes prepared from crystalline bacterial cell surface layers as model systems for studying the influence of surface properties on protein adsorption. J Membrane Sci 121:185–196
Weiner C, Sara M, Dasgupta G et al (1994a) Affinity cross-flow filtration: purification of IgG with a novel protein a affinity matrix prepared from two-dimensional protein crystals. Biotechnol Bioeng 44:55–65
Weiner C, Sara M, Sleytr UB (1994b) Novel protein a affinity matrix prepared from two-dimensional protein crystals. Biotechnol Bioeng 43:321–330
Wiegrabe W, Nonnenmacher M, Guckenberger R et al (1991) Atomic force microscopy of a hydrated bacterial surface protein. J Microsc (Oxf) 163:79–84
Wugeditsch T, Zachara NE, Puchberger M et al (1999) Structural heterogeneity in the core oligosaccharide of the S-layer glycoprotein from Aneurinibacillus thermoaerophilus DSM 10155. Glycobiology 9:787–795
Zhao GS, Ali E, Sakka M et al (2006) Binding of S-layer homology modules from Clostridium thermocellum SdbA to peptidoglycans. Appl Microbiol Biotechnol 70:464–469
Zuber B, Chami M, Houssin C et al (2008) Direct visualization of the outer membrane of mycobacteria and corynebacteria in their native state. J Bacteriol 190:5672–5680
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Madhurantakam, C., Howorka, S., Remaut, H. (2014). S-layer Structure in Bacteria and Archaea. In: Barton, L., Bazylinski, D., Xu, H. (eds) Nanomicrobiology. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1667-2_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1667-2_2
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1666-5
Online ISBN: 978-1-4939-1667-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)