Abstract
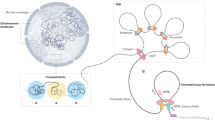
Understanding how eukaryotic gene regulation works implies unraveling the mechanisms used by transcription factors to access DNA information packaged in chromatin. The current view is that different cell types express different parts of the genome because they are equipped with different sets of transcription factors. A few transcription factors are called pioneer factors because they are able to bind to their sites in nucleosomes and to open up chromatin thus enabling access for other transcription factors, which are unable to recognize DNA packaged in nucleosomes. But it is also possible that the way DNA is organized in chromatin differs between cell types and contributes to cell identity by restricting or enhancing access to specific gene cohorts. To unravel these mechanisms we studied the interaction of progesterone receptor with the genome of breast cancer cells and found that it binds preferentially to sites organized in nucleosomes, which contribute to functional interactions leading to gene regulation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kornberg RD, Lorch Y (1999) Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell 98:285–294
Kornberg RD, Lorch Y (2002) Chromatin and transcription: where do we go from here. Curr Opin Genet Dev 12:249–251
Gaffney DJ, McVicker G, Pai AA et al (2012) Controls of nucleosome positioning in the human genome. PLoS Genet 8:e1003036
He X, Chatterjee R, John S et al (2013) Contribution of nucleosome binding preferences and co-occurring DNA sequences to transcription factor binding. BMC Genomics 14:428
Mirny LA (2010) Nucleosome-mediated cooperativity between transcription factors. Proc Natl Acad Sci U S A 107:22534–22539
Beato M, Herrlich P, Schutz G (1995) Steroid hormone receptors: many actors in search of a plot. Cell 83:851–857
Biddie SC, John S, Sabo PJ et al (2011) Transcription factor AP1 potentiates chromatin accessibility and glucocorticoid receptor binding. Mol Cell 43:145–155
Carroll JS, Liu XS, Brodsky AS et al (2005) Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122:33–43
Hurtado A, Holmes KA, Ross-Innes CS et al (2011) FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat Genet 43:27–33
Ballare C, Castellano G, Gaveglia L et al (2013) Nucleosome-driven transcription factor binding and gene regulation. Mol Cell 49:67–79
Strutt H, Paro R (1999) Mapping DNA target sites of chromatin proteins in vivo by formaldehyde crosslinking. Methods Mol Biol 119:455–467
Axel R (1975) Cleavage of DNA in nuclei and chromatin with staphylococcal nuclease. Biochemistry 14:2921–2925
Clark RJ, Felsenfeld G (1971) Structure of chromatin. Nat New Biol 229:101–106
Frank SR, Schroeder M, Fernandez P et al (2001) Binding of c-Myc to chromatin mediates mitogen-induced acetylation of histone H4 and gene activation. Genes Dev 15:2069–2082
Langmead B, Trapnell C, Pop M et al (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/Map format and SAMtools. Bioinformatics 25:2078–2079
Kent WJ, Sugnet CW, Furey TS et al (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Ye T, Krebs AR, Choukrallah MA et al (2011) seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res 39:e35
Althammer S, Gonzalez-Vallinas J, Ballare C et al (2011) Pyicos: a versatile toolkit for the analysis of high-throughput sequencing data. Bioinformatics 27:3333–3340
Nelson CC, Hendy SC, Shukin RJ et al (1999) Determinants of DNA sequence specificity of the androgen, progesterone, and glucocorticoid receptors: evidence for differential steroid receptor response elements. Mol Endocrinol 13:2090–2107
Bailey TL, Williams N, Misleh C et al (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Acknowledgements
We thank Dr Giancarlo Castellano (Hospital Clínic, Barcelona) for the first analysis of the ChIP and MNase-seq data and Laura Gaveglia for the first MNase experiments.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Vicent, G.P., Nacht, A.S., Ballaré, C., Zaurin, R., Soronellas, D., Beato, M. (2014). Progesterone Receptor Interaction with Chromatin. In: Castoria, G., Auricchio, F. (eds) Steroid Receptors. Methods in Molecular Biology, vol 1204. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-1346-6_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1346-6_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-1345-9
Online ISBN: 978-1-4939-1346-6
eBook Packages: Springer Protocols