Abstract
The mitochondrion is the main site for ATP production in the adult heart and comprises up to 40 % of the cardiac myocyte volume. It is now recognized that a complex network of nuclear transcription factors is essential for the coordinated regulation of mitochondrial biogenesis, maturation and function. These transcription factors guide developmental changes in mitochondrial number, structure, and dynamics as well as respond to various physiologic and pathophysiologic cues to meet the energetic needs of the adult heart. The peroxisome proliferator-activated receptor gamma coactivator-1 (PGC-1) orchestrates the actions of many of these transcription factors to maintain a high level of mitochondrial ATP production. There is increasing evidence that during the development of cardiac hypertrophy and in the failing heart, the activity of this network, including PGC-1, is altered. This review summarizes our current understanding of the perturbations in the gene regulatory pathways that occur during the development of heart failure. An appreciation of the role this regulatory circuitry serves in the regulation of cardiac energy metabolism may guide the development of novel therapeutic targets aimed at the metabolic disturbances that presage heart failure.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction: Nuclear Transcription Factors Controlling Cardiac Mitochondrial Biogenesis and Function
1.1 Transcriptional Control of the Mitochondrial Genome
The mitochondrial genome encodes 13 essential protein subunits of the electron transport chain (ETC) along with several rRNAs and tRNAs necessary for the translation of the mitochondrial-encoded transcripts. The human cardiac myocyte is estimated to contain approximately 7,000 mtDNA copies per diploid genome [1], reflective of the high mitochondrial density in the cell. Several nuclear-encoded factors act within the mitochondria to activate transcription of mitochondrial-encoded products. Factors such as the mitochondrial transcription factor A (Tfam), the mitochondrial transcription factor B2 (TFB2M) and others are critical for mtDNA transcription, organization, and maintenance (Reviewed in [2]).
1.2 Nuclear Respiratory Factors (NRFs)
The nuclear respiratory factors 1 and 2 (NRF-1 and NRF-2) are nuclear transcription factors that control many fundamental aspects of mitochondrial number and function. The importance of NRF-1 as a nuclear factor involved in mitochondrial biogenesis was first demonstrated by its regulation of cytochrome c gene expression [3]. Subsequently, it was shown that NRF-1 regulates expression of several proteins that act directly on regulators of the mitochondrial genome including Tfam and TFB2M [4, 5]; providing the first evidence for a direct connection between the transcriptional control of the nuclear and mitochondrial genomes. Gene deletion of NRF-1 results in embryonic lethality at E6.5 with diminished mtDNA levels and respiratory chain activity [6]. NRF-2, the human homolog of the murine GABP, is a polypeptide transcription factor containing an ETS-domain DNA subunit. NRF-1, together with NRF-2, regulates all ten nuclear-encoded cytochrome oxidase subunits [7, 8]. More recently, chromatin immunoprecipitation followed by deep sequencing (ChIP-seq) has confirmed NRF-1 binding sites in promoters of genes encoding components present in all electron transport chain complexes (Table 1) [9].
1.3 Nuclear Receptor Transcription Factors
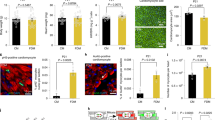
A subset of the nuclear receptor superfamily exerts transcriptional control on cardiac mitochondrial fuel metabolism and respiratory function. Some of these nuclear receptors are ligand-activated and, as such, can respond directly to cellular signals and metabolite levels (substrate availability). One relevant family, the peroxisome proliferator-activated receptors (PPARs), was originally identified as regulators of peroxisomal fatty acid oxidation (FAO) enzymes [10]. The PPARs bind to cognate DNA elements as an obligate heterodimer with the retinoid X receptor (RXR). A variety of FA derivatives have been shown to serve as ligands for the PPARs, yet the endogenous ligands have not been fully delineated. All three isoforms, PPARα, δ(β), and γ are expressed in the cardiac myocyte with overlapping but distinct functions, with PPARα and PPARδ exhibiting the highest level of expression. PPARα target genes comprise many proteins and enzymes involved in FA uptake and mitochondrial FAO (Table 1). Insight into the function of PPARα in heart has been gained by loss- and gain-of-function mouse models. Mice lacking PPARα have reduced cardiac FAO rates [11–13] while overexpression of PPARα in the heart leads to increased uptake, storage and oxidation of fatty acids [14]. As discussed further below, the metabolic phenotype of mice overexpressing PPARα in heart recapitulates many aspects of the insulin resistant diabetic heart. Interestingly, recent evidence suggests that PPARα-activating ligands are released from intracellular triglyceride stores (Fig. 1). Specifically, adipose triglyceride lipase (ATGL) was shown to be necessary for the generation of PPARα ligands in the cardiac myocyte; ATGL knockout mice display profoundly reduced expression of PPARα target genes involved in FAO [15]. These new results also suggest that lipolysis of intracellular triglyceride stores is an important source of long-chain fatty acids for mitochondrial FAO [16]. These collective results suggest a metabolic control mechanism whereby PPARα senses substrate availability to regulate capacity for mitochondrial FAO in heart.
The PGC-1 gene regulatory circuit controls cardiac mitochondrial function and fuel utilization capacity. PGC-1α integrates signals from diverse physiologic stimuli including developmental cues, exercise and β-adrenergic signaling. Coactivation of multiple downstream transcription factors drives the expression of proteins involved in virtually all aspects of mitochondrial energy production, regulating capacity for ATP production. Activity of PGC-1α and this circuit is inhibited during cardiac hypertrophy and heart failure. PGC-1α, PPARγ coactivator-1 alpha; PPARα, peroxisome proliferator-activated receptor alpha; RXR, retinoid X receptor; ERR, estrogen-related receptor; NRF, nuclear respiratory factor; OXPHOS, oxidative phosphorylation; ATGL, adipose triglyceride lipase; AMPK, AMP-activated protein kinase
PPARδ also drives high rates of mitochondrial FAO in the cardiac myocyte as its deletion reduces FAO capacity and leads to cardiac dysfunction [17]. However, unlike PPARα, cardiac overexpression of PPARδ does not lead to lipid accumulation and associated lipotoxicity. In fact, PPARδ appears to protect against myocyte lipid accumulation on a high-fat diet and cardiac dysfunction caused by pressure overload [18, 19]. In addition, PPARδ activates glucose uptake and oxidation at least in part due to its regulation of the insulin-responsive glucose transporter, Glut4 [17, 18]. Thus, PPARδ appears to drive a more balanced energy substrate utilization pattern. Cardiac-specific deletion of PPARδ also leads to a reduction of mitochondrial FAO and glucose oxidation rates resulting from a decrease in expression of enzymes in both pathways [17]. Therefore, PPARα and PPARδ share many similar targets, however these 2 nuclear receptors drive distinct metabolic pathways in the cardiac myocyte.
The estrogen-related receptors (ERRs) form a second group of nuclear receptors critical for maintaining mitochondrial function in the cardiac myocyte. All three ERR isoforms (ERRα, ERRβ, and ERRγ) are expressed in the heart and are known as “orphan” nuclear receptors due to the lack of a known ligand. ERR target genes overlap with that of the PPARs but also regulate genes involved in virtually all aspects of mitochondrial energy production including the TCA cycle, respiratory chain and oxidative phosphorylation (Table 1) [20]. Loss-of-function studies in mice have revealed roles for the ERRs in the heart. ERRα knockout mice display cardiac dysfunction when subjected to pressure-overload related to reduced capacity for maintaining phosphocreatine stores and mitochondrial ATP synthesis rates in response to energetic stress [21]. Likewise, gene deletion of ERRγ results in cardiomyopathy and death immediately following birth [22] related to the loss of the normal postnatal shift to oxidative metabolism and fatty acid utilization. As such, ERRs are required for the establishment and maintenance of cardiac energy production.
1.4 Other Nuclear Transcription Factors
Additional transcription factors have also been shown to regulate mitochondrial pathways and function in heart. A computational approach identified Yin Yang 1 (YY1) binding sites in the promoter regions of many mitochondrial genes regulated by the nutrient sensor mammalian target of rapamycin, mTOR (Table 1) [23]. YY1 deletion in skeletal muscle results in profound defects in mitochondrial structure and energy production [24]. The oncoprotein c-Myc has also been shown to activate expression of genes involved in mitochondrial biogenesis [25, 26]. Although expressed at very low levels in the normal heart, c-Myc is induced by a variety of pathologic stimuli including pressure overload [27]. The major effects of c-Myc expression in the heart are the activation of glucose utilization and downregulation of FAO (Table 1) [28]. Forced expression of c-Myc in the heart also leads to abnormal mitochondrial biogenesis with respiratory change defects leading to cardiac dysfunction [29]. These data are consistent with a role of c-Myc in cardiac myocyte cell-cycle re-entry and activation of the fetal gene program during pathologic hypertrophy.
2 The PPARγ Coactivator 1 (PGC-1) Transcriptional Coregulators: Transducers of Physiological Cues to the Control of Cardiac Mitochondrial Biogenesis and Function
A huge breakthrough in our understanding of how mitochondrial biogenesis and function is regulated in accordance with energy demands came with the discovery of the PGC-1 coregulators. PGC-1α was first discovered in brown adipocytes as an activator of PPARγ [30]. The closely related PGC-1β [31, 32] and more distant relative, PGC-1 related coactivator (PRC) [33] comprise the other members of the family. The PGC-1 coactivators are characterized by an LXXLL motif that mediates interaction with nuclear receptors, an RNA recognition motif (RRM), and a host cell factor-1 (HCF) binding domain. PGC-1α and PGC-1β are expressed in tissues with a high oxidative capacity including the heart. Remarkably, PGC-1α is a highly inducible factor that responds to stimuli such as cold exposure and exercise [34–36]. PGC-1α expression in the heart increases during cardiac development and coincident with the large mitochondrial biogenic response that occurs just before birth [37]. Overexpression of PGC-1α in the cardiac myocyte leads to a robust mitochondrial biogenic response and increased expression of nuclear-encoded mitochondrial genes involved in multiple energy production pathways [37]. This function is orchestrated by PGC-1’s interaction with and activation of PPARα [38], PPARβ/δ [39], ERRα and ERRγ [40–42]. In this manner, PGC-1α integrates multiple physiologic and developmental cues to regulate most aspects of mitochondrial function and energy production (illustrated in Fig. 1).
The importance of PGC-1 coactivators in the control of mitochondrial number and function is reinforced by loss-of-function studies in mice. Mice with loss of either PGC-1α [43, 44] or PGC-1β [45, 46] are viable and fertile with no obvious defect in cardiac energy metabolism or mitochondrial density. In the heart, pressure-overload induced hypertrophy results in accelerated heart failure in PGC-1α knockout mice [47]. A similar phenotype is also observed in PGC-1β knockouts subjected to pressure-overload hypertrophy [48]. In contrast to the single knockouts, deletion of both PGC-1α and PGC-1β results in perinatal lethality within 24 h of birth due to an arrest in mitochondrial biogenesis leading to heart failure [49]. This work was key to defining the important role played by PGC-1α/β in the perinatal cardiac mitochondrial biogenic response. In addition to its role in mitochondrial biogenesis during cardiac development, recent studies indicate that PGC-1α and PGC-1β are necessary for mitochondrial maturation including regulation of the mitochondrial fusion genes mitofusion 1 (Mfn1) and mitofusin 2 (Mfn2) among others during postnatal cardiac development [50, 51]. The regulation of Mfn1 and Mfn2 are at least in part, driven by PGC-1 coactivation of ERRα [50, 52]. Interestingly, inhibition of mitochondrial fusion blocks cardiac myocyte differentiation [53], providing a potential link between PGC-1, mitochondrial dynamics, and cardiac development.
3 Control of Mitochondrial Function by Cellular and Metabolic Signaling
3.1 PGC-1 Responds to Cellular Signals
The PGC-1 coactivators are dynamically regulated at both transcriptional and post-transcriptional levels to integrate a variety of physiologic and pathophysiologic cues to match energy production with cardiac energy demands. Consistent with this role, multiple cellular signaling pathways converge on the PGC-1α gene including calcineurin, calmodulin-dependent kinase (CaMK), cAMP signaling, and AMP-activated protein kinase (AMPK) [54–57]. The activation of PGC-1α expression by cAMP downstream of adrenergic stimulation is mediated through direct regulation by the cAMP response element binding protein (CREB) [56].
PGC-1 activity is also modulated by post-translational modifications including phosphorylation and acetylation. AMPK has been shown to directly phosphorylate and activate PGC-1 [58]. AMPK is an energy sensing kinase that is activated by energy depletion, i.e. higher AMP/ATP ratios. Accordingly, phosphorylation of PGC-1 by AMPK provides a mechanism to boost mitochondrial energy production upon increased physiologic demands such as exercise. In addition to AMPK, the nicotinamide adenine nucleotide (NAD+)-dependent deacetylase, Sirtuin 1 (SIRT1), is another metabolic sensor that regulates PGC-1 activity. AMPK and SIRT1 act cooperatively to activate PGC-1α activity providing a compelling link between mitochondrial biogenesis and metabolic signaling pathways sensing changes in cellular energy status [59, 60]. The effect of PGC-1 acetylation is supported by the observation that PGC-1 acetylation status is increased in muscle in mice fed a high-fat diet [61].
4 Dysregulation of Transcriptional Networks Controlling Mitochondrial Function Relevant to Heart Disease
4.1 Deactivation of Transcriptional Circuits in the Hypertrophied and Failing Heart
Similar to the expression of many structural and contractile proteins, there is a switch to the “fetal gene program” evident in metabolic enzyme gene expression during the development of cardiac hypertrophy and failure, including a decrease in the expression of genes encoding mitochondrial FAO enzymes [62]. There is significant evidence that this fetal metabolic switch in the hypertrophied and failing heart involves alterations in the expression and activity of the transcription factors controlling mitochondrial function and biogenesis. PPARα expression is decreased in human and animal models of pressure overload-induced cardiac hypertrophy and in heart failure [62–65]. Evidence also exists that during cardiac hypertrophy PPARα, in addition to decreased expression, is regulated at the post-transcriptional level by phosphorylation via p38 to impair activity [66]. This is in agreement with the observed metabolic shift from FAO to glucose metabolism that occurs during the transition to cardiac hypertrophy and heart failure [67–69]. ERRα expression has also been shown to be decreased in animal models and human heart failure samples [70–72]. Coordinated repression of both PPARα and ERRα could represent major determinants in the regulation of mitochondrial energy production in the failing heart. As ERRα can directly regulate PPARα [73], it is tempting to speculate that repression of ERRα establishes this vicious cycle.
Given its role as a “master regulator” of mitochondrial biogenesis and function, a logical question is whether dysregulation of PGC-1 is involved in the pathogenesis of the “energy-starved” phenotype of the failing heart. Indeed, changes in fuel selection and mitochondrial function in the hypertrophied and failing heart are likely consequences of dysregulation of the PGC-1 circuit. Consistent with this notion, the expression of PGC-1α has been found to be downregulated in both animal models and human heart failure samples [62, 70, 74]. Similar findings have been reported for PGC-1β [48]. It should be noted however, that not all studies have shown a downregulation in PGC-1 in human heart failure samples [71, 72]. These differences likely reflect different etiologies, timepoints, or other variables that could influence PGC-1 expression secondarily. In addition, the role of altered PGC-1 signaling as an early (causal?) versus late event in the metabolic remodeling of heart failure remains to be determined.
4.2 Dysregulation of Transcriptional Circuits in the Diabetic Heart
The metabolic derangements that occur in the obese, insulin resistant, and diabetic patient appear to be quite distinct from that of hypertensive or ischemic heart disease. Insulin-resistant and diabetic hearts exhibit reduced capacity for glucose utilization, higher rates of mitochondrial FAO. Increased cardiac fatty acid uptake and β-oxidation have been observed in experimental models as well as obese and diabetic patients [75–78]. Interestingly, upregulation of cardiac mitochondrial FAO enzyme expression has been observed in a number of diabetic animal models consistent with an activation of FAO rates [79–81]. There is also increasing evidence that impaired glucose tolerance, even before the onset of diabetes, is associated with cardiac steatosis [82]. In the obese and diabetic patient population, myocardial lipid accumulation has been shown by some to be associated with diastolic and, in some studies, systolic ventricular dysfunction [78, 83, 84]. Collectively, this metabolic inflexibility in the setting of neutral lipid accumulation and resulting cardiac dysfunction has been termed cardiac “lipotoxicity” [85].
The high rates of mitochondrial FAO observed in both experimental models and human patients indicate activation of transcriptional networks controlling these pathways. Indeed, mice that overexpress PPARα exclusively in the heart (MHC-PPARα) exhibit increased FAO associated with a decrease in glucose utilization resulting in an accumulation of triglyceride and a diabetic-like phenotype [14, 86, 87]. These animals exhibit left ventricular hypertrophy and cardiac dysfunction that can be inhibited by deletion of CD36 or the LDL receptor, thus depriving PPARα of activating ligands [88, 89]. Furthermore, the activity of the PGC-1/PPARα circuit is increased in the insulin resistant mouse heart [90]. However, during transition to diabetes in mouse models the activity of PGC-1α appears to fall leading to a proposed vicious cycle of mitochondrial dysfunction and myocyte lipotoxicity [91]. The relevance of this latter observation to humans remains to be determined.
5 Conclusions
The healthy heart has an amazing capacity to generate ATP and shift fuel utilization preferences in response to pathophysiologic stimuli to meet its energy needs. Much of this control occurs at the level of gene regulation through a complex circuit involving the PGC-1 coactivators and nuclear receptors. This circuit becomes constrained during the development of heart failure. The unmet needs in the heart failure arena suggest a prime opportunity to develop therapeutics that modulate mitochondrial function and fuel metabolism by targeting this circuitry. In the long-term, such therapies could, perhaps, be tailored to the etiology of heart failure and the accompanying metabolic derangements.
References
Miller FJ, Rosenfeldt FL, Zhang C et al (2003) Precise determination of mitochondrial DNA copy number in human skeletal and cardiac muscle by a PCR-based assay: lack of change of copy number with age. Nucleic Acids Res 31:e61
Scarpulla RC, Vega RB, Kelly DP (2012) Transcriptional integration of mitochondrial biogenesis. Trends Endocrinol Metab 23:459–466
Evans MJ, Scarpulla RC (1989) Interaction of nuclear factors with multiple sites in the somatic cytochrome c promoter. Characterization of upstream NRF-1, ATF, and intron Sp1 recognition sequences. J Biol Chem 264:14361–14368
Virbasius JV, Scarpulla RC (1994) Activation of the human mitochondrial transcription factor A gene by nuclear respiratory factors: a potential regulatory link between nuclear and mitochondrial gene expression in organelle biogenesis. Proc Natl Acad Sci U S A 91:1309–1313
Gleyzer N, Vercauteren K, Scarpulla RC (2005) Control of mitochondrial transcription specificity factors (TFB1M and TFB2M) by nuclear respiratory factors (NRF-1 and NRF-2) and PGC-1 family coactivators. Mol Cell Biol 25:1354–1366
Huo L, Scarpulla RC (2001) Mitochondrial DNA instability and peri-implantation lethality associated with targeted disruption of nuclear respiratory factor 1 in mice. Mol Cell Biol 21:644–654
Ongwijitwat S, Liang HL, Graboyes EM et al (2006) Nuclear respiratory factor 2 senses changing cellular energy demands and its silencing down-regulates cytochrome oxidase and other target gene mRNAs. Gene 374:39–49
Dhar SS, Ongwijitwat S, Wong-Riley MT (2008) Nuclear respiratory factor 1 regulates all ten nuclear-encoded subunits of cytochrome c oxidase in neurons. J Biol Chem 283:3120–3129
Satoh J, Kawana N, Yamamoto Y (2013) Pathway analysis of ChIP-Seq-based NRF1 target genes suggests a logical hypothesis of their involvement in the pathogenesis of neurodegenerative diseases. Gene Regul Syst Biol 7:139–152
Issemann I, Green S (1990) Activation of a member of the steroid hormone receptor superfamily by peroxisome proliferators. Nature 347:645–650
Leone TC, Weinheimer CJ, Kelly DP (1999) A critical role for the peroxisome proliferator-activated receptor alpha (PPARalpha) in the cellular fasting response: the PPARalpha-null mouse as a model of fatty acid oxidation disorders. Proc Natl Acad Sci U S A 96:7473–7478
Djouadi F, Brandt JM, Weinheimer CJ et al (1999) The role of the peroxisome proliferator-activated receptor alpha (PPAR alpha) in the control of cardiac lipid metabolism. Prostaglandins Leukot Essent Fatty Acids 60:339–343
Watanabe K, Fujii H, Takahashi T et al (2000) Constitutive regulation of cardiac fatty acid metabolism through peroxisome proliferator-activated receptor alpha associated with age-dependent cardiac toxicity. J Biol Chem 275:22293–22299
Finck BN, Han X, Courtois M et al (2003) A critical role for PPARalpha-mediated lipotoxicity in the pathogenesis of diabetic cardiomyopathy: modulation by dietary fat content. Proc Natl Acad Sci U S A 100:1226–1231
Haemmerle G, Moustafa T, Woelkart G et al (2011) ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-alpha and PGC-1. Nat Med 17:1076–1085
Banke NH, Wende AR, Leone TC et al (2010) Preferential oxidation of triacylglyceride-derived fatty acids in heart is augmented by the nuclear receptor PPARalpha. Circ Res 107:233–241
Wang P, Liu J, Li Y et al (2010) Peroxisome proliferator-activated receptor delta is an essential transcriptional regulator for mitochondrial protection and biogenesis in adult heart. Circ Res 106:911–919
Burkart EM, Sambandam N, Han X et al (2007) Nuclear receptors PPARbeta/delta and PPARalpha direct distinct metabolic regulatory programs in the mouse heart. J Clin Invest 117:3930–3939
Liu J, Wang P, Luo J et al (2011) Peroxisome proliferator-activated receptor beta/delta activation in adult hearts facilitates mitochondrial function and cardiac performance under pressure-overload condition. Hypertension 57:223–230
Dufour CR, Wilson BJ, Huss JM et al (2007) Genome-wide orchestration of cardiac functions by the orphan nuclear receptors ERRalpha and gamma. Cell Metab 5:345–356
Huss JM, Imahashi K, Dufour CR et al (2007) The nuclear receptor ERRalpha is required for the bioenergetic and functional adaptation to cardiac pressure overload. Cell Metab 6:25–37
Alaynick WA, Kondo RP, Xie W et al (2007) ERRgamma directs and maintains the transition to oxidative metabolism in the postnatal heart. Cell Metab 6:13–24
Cunningham JT, Rodgers JT, Arlow DH et al (2007) mTOR controls mitochondrial oxidative function through a YY1-PGC-1alpha transcriptional complex. Nature 450:736–740
Blattler SM, Verdeguer F, Liesa M et al (2012) Defective mitochondrial morphology and bioenergetic function in mice lacking the transcription factor Yin Yang 1 in skeletal muscle. Mol Cell Biol 32:3333–3346
Li F, Wang Y, Zeller KI et al (2005) Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol Cell Biol 25:6225–6234
Kim J, Lee JH, Iyer VR (2008) Global identification of Myc target genes reveals its direct role in mitochondrial biogenesis and its E-box usage in vivo. PLoS One 3:e1798
Izumo S, Nadal-Ginard B, Mahdavi V (1988) Protooncogene induction and reprogramming of cardiac gene expression produced by pressure overload. Proc Natl Acad Sci U S A 85:339–343
Ahuja P, Zhao P, Angelis E et al (2010) Myc controls transcriptional regulation of cardiac metabolism and mitochondrial biogenesis in response to pathological stress in mice. J Clin Invest 120:1494–1505
Lee HG, Chen Q, Wolfram JA et al (2009) Cell cycle re-entry and mitochondrial defects in myc-mediated hypertrophic cardiomyopathy and heart failure. PLoS One 4:e7172
Puigserver P, Wu Z, Park CW et al (1998) A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 92:829–839
Kressler D, Schreiber SN, Knutti D et al (2002) The PGC-1-related protein PERC is a selective coactivator of estrogen receptor alpha. J Biol Chem 277:13918–13925
Lin J, Puigserver P, Donovan J et al (2002) Peroxisome proliferator-activated receptor gamma coactivator 1beta (PGC-1beta), a novel PGC-1-related transcription coactivator associated with host cell factor. J Biol Chem 277:1645–1648
Andersson U, Scarpulla RC (2001) PGC-1-related coactivator, a novel, serum-inducible coactivator of nuclear respiratory factor 1-dependent transcription in mammalian cells. Mol Cell Biol 21:3738–3749
Baar K, Wende AR, Jones TE et al (2002) Adaptations of skeletal muscle to exercise: rapid increase in the transcriptional coactivator PGC-1. FASEB J 16:1879–1886
Terada S, Goto M, Kato M et al (2002) Effects of low-intensity prolonged exercise on PGC-1 mRNA expression in rat epitrochlearis muscle. Biochem Biophys Res Commun 296:350–354
Wu Z, Puigserver P, Andersson U et al (1999) Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell 98:115–124
Lehman JJ, Barger PM, Kovacs A et al (2000) Peroxisome proliferator-activated receptor gamma coactivator-1 promotes cardiac mitochondrial biogenesis. J Clin Invest 106:847–856
Vega RB, Huss JM, Kelly DP (2000) The coactivator PGC-1 cooperates with peroxisome proliferator-activated receptor alpha in transcriptional control of nuclear genes encoding mitochondrial fatty acid oxidation enzymes. Mol Cell Biol 20:1868–1876
Wang YX, Lee CH, Tiep S et al (2003) Peroxisome-proliferator-activated receptor delta activates fat metabolism to prevent obesity. Cell 113:159–170
Hentschke M, Susens U, Borgmeyer U (2002) PGC-1 and PERC, coactivators of the estrogen receptor-related receptor gamma. Biochem Biophys Res Commun 299:872–879
Huss JM, Kopp RP, Kelly DP (2002) Peroxisome proliferator-activated receptor coactivator-1alpha (PGC-1alpha) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-alpha and -gamma. Identification of novel leucine-rich interaction motif within PGC-1alpha. J Biol Chem 277:40265–40274
Schreiber SN, Knutti D, Brogli K et al (2003) The transcriptional coactivator PGC-1 regulates the expression and activity of the orphan nuclear receptor estrogen-related receptor alpha (ERRalpha). J Biol Chem 278:9013–9018
Lin J, Wu PH, Tarr PT et al (2004) Defects in adaptive energy metabolism with CNS-linked hyperactivity in PGC-1alpha null mice. Cell 119:121–135
Leone TC, Lehman JJ, Finck BN et al (2005) PGC-1alpha deficiency causes multi-system energy metabolic derangements: muscle dysfunction, abnormal weight control and hepatic steatosis. PLoS Biol 3:e101
Lelliott CJ, Medina-Gomez G, Petrovic N et al (2006) Ablation of PGC-1beta results in defective mitochondrial activity, thermogenesis, hepatic function, and cardiac performance. PLoS Biol 4:e369
Sonoda J, Mehl IR, Chong LW et al (2007) PGC-1beta controls mitochondrial metabolism to modulate circadian activity, adaptive thermogenesis, and hepatic steatosis. Proc Natl Acad Sci U S A 104:5223–5228
Arany Z, Novikov M, Chin S et al (2006) Transverse aortic constriction leads to accelerated heart failure in mice lacking PPAR-gamma coactivator 1alpha. Proc Natl Acad Sci U S A 103:10086–10091
Riehle C, Wende AR, Zaha VG et al (2011) PGC-1beta deficiency accelerates the transition to heart failure in pressure overload hypertrophy. Circ Res 109:783–793
Lai L, Leone TC, Zechner C et al (2008) Transcriptional coactivators PGC-1alpha and PGC-lbeta control overlapping programs required for perinatal maturation of the heart. Genes Dev 22:1948–1961
Martin OJ, Lai L, Soundarapandian MM et al (2014) A role for PGC-1 coactivators in the control of mitochondrial dynamics during postnatal cardiac growth. Circ Res 114(4):626–636
Liesa M, Borda-d'Agua B, Medina-Gomez G et al (2008) Mitochondrial fusion is increased by the nuclear coactivator PGC-1beta. PLoS One 3:e3613
Soriano FX, Liesa M, Bach D et al (2006) Evidence for a mitochondrial regulatory pathway defined by peroxisome proliferator-activated receptor-gamma coactivator-1 alpha, estrogen-related receptor-alpha, and mitofusin 2. Diabetes 55:1783–1791
Kasahara A, Cipolat S, Chen Y et al (2013) Mitochondrial fusion directs cardiomyocyte differentiation via calcineurin and notch signaling. Science 342:734–737
Handschin C, Rhee J, Lin J et al (2003) An autoregulatory loop controls peroxisome proliferator-activated receptor gamma coactivator 1alpha expression in muscle. Proc Natl Acad Sci U S A 100:7111–7116
Schaeffer PJ, Wende AR, Magee CJ et al (2004) Calcineurin and calcium/calmodulin-dependent protein kinase activate distinct metabolic gene regulatory programs in cardiac muscle. J Biol Chem 279:39593–39603
Rohas LM, St-Pierre J, Uldry M et al (2007) A fundamental system of cellular energy homeostasis regulated by PGC-1alpha. Proc Natl Acad Sci U S A 104:7933–7938
Zong H, Ren JM, Young LH et al (2002) AMP kinase is required for mitochondrial biogenesis in skeletal muscle in response to chronic energy deprivation. Proc Natl Acad Sci U S A 99:15983–15987
Jager S, Handschin C, St-Pierre J et al (2007) AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1alpha. Proc Natl Acad Sci U S A 104:12017–12022
Canto C, Gerhart-Hines Z, Feige JN et al (2009) AMPK regulates energy expenditure by modulating NAD+ metabolism and SIRT1 activity. Nature 458:1056–1060
Iwabu M, Yamauchi T, Okada-Iwabu M et al (2010) Adiponectin and AdipoR1 regulate PGC-1alpha and mitochondria by Ca(2+) and AMPK/SIRT1. Nature 464:1313–1319
Coste A, Louet JF, Lagouge M et al (2008) The genetic ablation of SRC-3 protects against obesity and improves insulin sensitivity by reducing the acetylation of PGC-1alpha. Proc Natl Acad Sci U S A 105:17187–17192
Sack MN, Rader TA, Park S et al (1996) Fatty acid oxidation enzyme gene expression is downregulated in the failing heart. Circulation 94:2837–2842
Dewald O, Sharma S, Adrogue J et al (2005) Downregulation of peroxisome proliferator-activated receptor-alpha gene expression in a mouse model of ischemic cardiomyopathy is dependent on reactive oxygen species and prevents lipotoxicity. Circulation 112:407–415
Goikoetxea MJ, Beaumont J, Gonzalez A et al (2006) Altered cardiac expression of peroxisome proliferator-activated receptor-isoforms in patients with hypertensive heart disease. Cardiovasc Res 69:899–907
Karbowska J, Kochan Z, Smolenski RT (2003) Peroxisome proliferator-activated receptor alpha is downregulated in the failing human heart. Cell Mol Biol Lett 8:49–53
Barger PM, Brandt JM, Leone TC et al (2000) Deactivation of peroxisome proliferator-activated receptor-alpha during cardiac hypertrophic growth. J Clin Invest 105:1723–1730
Bishop SP, Altschuld RA (1970) Increased glycolytic metabolism in cardiac hypertrophy and congestive failure. Am J Physiol 218:153–159
Nascimben L, Ingwall JS, Lorell BH et al (2004) Mechanisms for increased glycolysis in the hypertrophied rat heart. Hypertension 44:662–667
Osorio JC, Stanley WC, Linke A et al (2002) Impaired myocardial fatty acid oxidation and reduced protein expression of retinoid X receptor-alpha in pacing-induced heart failure. Circulation 106:606–612
Sihag S, Cresci S, Li AY et al (2009) PGC-1alpha and ERRalpha target gene downregulation is a signature of the failing human heart. J Mol Cell Cardiol 46:201–212
Hu X, Xu X, Lu Z et al (2011) AMP activated protein kinase-alpha2 regulates expression of estrogen-related receptor-alpha, a metabolic transcription factor related to heart failure development. Hypertension 58:696–703
Karamanlidis G, Nascimben L, Couper GS et al (2010) Defective DNA replication impairs mitochondrial biogenesis in human failing hearts. Circ Res 106:1541–1548
Huss JM, Torra IP, Staels B et al (2004) Estrogen-related receptor alpha directs peroxisome proliferator-activated receptor alpha signaling in the transcriptional control of energy metabolism in cardiac and skeletal muscle. Mol Cell Biol 24:9079–9091
Garnier A, Zoll J, Fortin D et al (2009) Control by circulating factors of mitochondrial function and transcription cascade in heart failure: a role for endothelin-1 and angiotensin II. Circ Heart Fail 2:342–350
Belke DD, Larsen TS, Gibbs EM et al (2000) Altered metabolism causes cardiac dysfunction in perfused hearts from diabetic (db/db) mice. Am J Physiol Endocrinol Metab 279:E1104–E1113
Peterson LR, Herrero P, McGill J et al (2008) Fatty acids and insulin modulate myocardial substrate metabolism in humans with type 1 diabetes. Diabetes 57:32–40
Peterson LR, Soto PF, Herrero P et al (2008) Impact of gender on the myocardial metabolic response to obesity. JACC Cardiovasc Imaging 1:424–433
Rijzewijk LJ, van der Meer RW, Lamb HJ et al (2009) Altered myocardial substrate metabolism and decreased diastolic function in nonischemic human diabetic cardiomyopathy: studies with cardiac positron emission tomography and magnetic resonance imaging. J Am Coll Cardiol 54:1524–1532
Hamblin M, Friedman DB, Hill S et al (2007) Alterations in the diabetic myocardial proteome coupled with increased myocardial oxidative stress underlies diabetic cardiomyopathy. J Mol Cell Cardiol 42:884–895
Jullig M, Hickey AJ, Middleditch MJ et al (2007) Characterization of proteomic changes in cardiac mitochondria in streptozotocin-diabetic rats using iTRAQ isobaric tags. Proteomics Clin Appl 1:565–576
Bugger H, Chen D, Riehle C et al (2009) Tissue-specific remodeling of the mitochondrial proteome in type 1 diabetic akita mice. Diabetes 58:1986–1997
McGavock JM, Lingvay I, Zib I et al (2007) Cardiac steatosis in diabetes mellitus: a 1H-magnetic resonance spectroscopy study. Circulation 116:1170–1175
Korosoglou G, Humpert PM, Ahrens J et al (2012) Left ventricular diastolic function in type 2 diabetes mellitus is associated with myocardial triglyceride content but not with impaired myocardial perfusion reserve. J Magn Reson Imaging 35:804–811
Ng AC, Delgado V, Bertini M et al (2010) Myocardial steatosis and biventricular strain and strain rate imaging in patients with type 2 diabetes mellitus. Circulation 122:2538–2544
Unger RH, Scherer PE (2010) Gluttony, sloth and the metabolic syndrome: a roadmap to lipotoxicity. Trends Endocrinol Metab 21:345–352
Finck BN, Lehman JJ, Leone TC et al (2002) The cardiac phenotype induced by PPARalpha overexpression mimics that caused by diabetes mellitus. J Clin Invest 109:121–130
Park SY, Cho YR, Finck BN et al (2005) Cardiac-specific overexpression of peroxisome proliferator-activated receptor-alpha causes insulin resistance in heart and liver. Diabetes 54:2514–2524
Yang J, Sambandam N, Han X et al (2007) CD36 deficiency rescues lipotoxic cardiomyopathy. Circ Res 100:1208–1217
Duncan JG, Bharadwaj KG, Fong JL et al (2010) Rescue of cardiomyopathy in peroxisome proliferator-activated receptor-alpha transgenic mice by deletion of lipoprotein lipase identifies sources of cardiac lipids and peroxisome proliferator-activated receptor-alpha activators. Circulation 121:426–435
Duncan JG, Fong JL, Medeiros DM et al (2007) Insulin-resistant heart exhibits a mitochondrial biogenic response driven by the peroxisome proliferator-activated receptor-alpha/PGC-1alpha gene regulatory pathway. Circulation 115:909–917
Mitra R, Nogee DP, Zechner JF et al (2012) The transcriptional coactivators, PGC-1alpha and beta, cooperate to maintain cardiac mitochondrial function during the early stages of insulin resistance. J Mol Cell Cardiol 52:701–710
Acknowledgments
This work was supported by NIH grants (R01 DK045416, R01 HL058493, R01 HL101189 [D.P.K.]). We thank Lorenzo Thomas for help with manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Vega, R.B., Leone, T.C., Kelly, D.P. (2014). Transcriptional Control of Mitochondrial Biogenesis and Maturation. In: Lopaschuk, G., Dhalla, N. (eds) Cardiac Energy Metabolism in Health and Disease. Advances in Biochemistry in Health and Disease, vol 11. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1227-8_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1227-8_6
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1226-1
Online ISBN: 978-1-4939-1227-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)