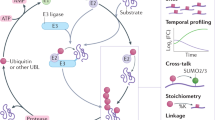
Abstract
Ubiquitylation is a posttranslational modification in which ubiquitin, a conserved polypeptide, is conjugated to targeted proteins. The human genome encodes for over a thousand proteins that can either mediate ubiquitylation, remove the modification, or bind to modified proteins. Hence, the ubiquitin system is expected to affect a large portion of the proteome. Since its discovery, our understanding of the ubiquitin system has come a long way. Great efforts made by several prominent members of the scientific community have paid off, with leaps and bounds being made in areas such as the discovery of ubiquitylated proteins and ubiquitylation sites. Boosting the success of classical proteomic approaches, the recent introduction of a new method (by which ubiquitylated peptides are captured by antibodies) enabled the identification of a large number of ubiquitylation sites. In this chapter, we will review the different proteomic strategies that have been established in order to uncover which proteins are ubiquitylated in the cell, and then further discuss the novel biological insights revealed by these systems-wide studies.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Ubiquitin
- Proteasome
- Mass spectrometry
- Proteomics
- Posttranslational modifications
- Ubiquitin remnant tail (diGly, GlyGly, GG)
1 Introduction: The Ubiquitin System and Ubiquitylation
The proteome is far more complex than the genome from which it is derived due to alternative splicing and extensive protein posttranslational modifications (PTMs) . Importantly, PTMs fulfill key roles by controlling protein-protein interactions, protein localization, enzymatic activities, and protein turnover. To date, ubiquitylation (i.e., modification by ubiquitin conjugation) is one of the most abundantly identified PTM after phosphorylation, of which there are more than 50,000 reported protein modification sites (www.phosphosite.org) [1]. The remarkable progress in the enrichment and proteomic techniques that we will review in this chapter have enabled great advancements in the identification of novel ubiquitylation sites, which now offers a unique opportunity to better understand the ubiquitin system and its impact on the proteome .
Ubiquitylation stands apart from other PTMs in that a small protein (ubiquitin), instead of small functional groups (like phosphate and acetyl), is used as the modifier. Ubiquitin was first discovered in 1975 [2] and was named after its ubiquitous expression in eukaryotes. Ubiquitin is a 76 amino acid, highly structured protein with a molecular weight of about 8.5 kDa. It is covalently attached to proteins through the formation of an isopeptide bond between the carboxyl group of its last glycine residue and, typically, the epsilon amino group of a target lysine residue. There are also other similar protein modifiers, called ubiquitin-like proteins (Ubls) , such as small ubiquitin-like modifier (SUMO), interferon-stimulated gene of 15 kDa (ISG15), and neural-precursor-cell-expressed and developmentally down-regulated gene (NEDD8). These Ubls have both a sequence and a structural homology to ubiquitin and are attached by similar conjugation mechanisms. However, proteomic analyses of these modifications will not be reviewed in this chapter.
Three classes of enzymes are required for the ubiquitylation cascade that leads to substrate modification, namely an ubiquitin-activating enzyme (E1), an ubiquitin-conjugating enzyme (E2), and an ubiquitin ligase (E3). Prior to substrate conjugation, ubiquitin first needs to be activated by the E1 enzyme through an adenosine triphosphate (ATP)-dependent reaction, in which a thioester linkage is formed between the C-terminus of ubiquitin and a cysteine residue of the E1 [3, 4]. The E1 then mediates the transfer of ubiquitin to a cysteine residue of an E2 enzyme via a trans-thioesterification reaction [5, 6]. Finally, an E3 ligase will recruit the target substrate for the last ligation step, either by facilitating the direct transfer of the ubiquitin molecule from the E2 to the associated substrate, or by transiently accepting the ubiquitin from the E2 prior to transferring it to the substrate [7]. Ubiquitylation mostly occurs on lysine residues of substrate proteins. However, in some cases the N-terminus of a protein is conjugated instead of a lysine [8–10]. Other residues, like cysteine, threonine, and serine, have also been reported as possible ubiquitylation sites [11–13] .
Ubiquitylation is a highly dynamic process that relies on a complex and modular network of proteins which comprise the ubiquitin system. There are more than 600 E3 ligases encoded in the human genome that are assisted by about 40 E2s [14]. To maintain balance of the system, ubiquitylation can also be reversed by a class of enzymes called de-ubiquitylating enzymes (DUBs), of which around 80 have so far been identified in humans. DUBs are involved in maintaining ubiquitin homeostasis by processing newly synthesized ubiquitin and recycling used ubiquitin as well as controlling or modulating the fate of ubiquitylated proteins [15].
Ubiquitylation is a versatile PTM due to its built-in ability to generate linkages of diverse architecture, which can each be used in different signalling pathways [16]. Substrate proteins can be modified by a single ubiquitin on a single lysine (monoubiquitylation) or multiple lysine residues (multi-monoubiquitylation). Alternatively, lysine residues within ubiquitin itself can be used to covalently attach subsequent ubiquitin molecules, forming multimeric chains conjugated to a single lysine residue on the targeted protein (polyubiquitylation). The seven lysine residues (K6, 11, 27, 29, 33, 48, and 63) and the amino-terminus (M1) of ubiquitin can be used to generate ubiquitin chains that are either homogenous (one linkage type throughout), mixed (with different linkage types), or branched (arising when two ubiquitins are conjugated to separate lysines on the same ubiquitin moiety, thus creating a branching point). To make matters even more complicated, ubiquitin chains can also be built upon other Ubl modifications. For example, the RNF4 E3 ligase specifically extends the SUMO modification with a poly-ubiquitin chain [17]. To recognize and distinguish between different ubiquitin modifications, over 20 different families of ubiquitin-binding domains (UBDs) have evolved, which can be specific either to mono-ubiquitin or to one type of poly-ubiquitin chain [18–20] .
Due to high proteomic penetrance, ubiquitylation affects most cellular pathways in some way. A major function of ubiquitylation is to target substrates for proteasome degradation. In addition (or in tandem), a plethora of different processes are regulated by ubiquitylation, including endocytosis, selective macro-autophagy, cell cycle control, inflammation and nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) activation, deoxyribonucleic acid (DNA) repair, transcription, antigen processing, viral infection, and ribosome and peroxisome biogenesis [16, 21, 22]. While our general understanding of the enzymatic mechanisms for ubiquitylation has advanced impressively thanks to detailed biochemical analyses, a more comprehensive view of the relationships among the different components of the ubiquitin system is still lacking. Therefore, there is an increasing need to integrate more systems-wide approaches, such as mass spectrometry-based proteomics , which would offer a broader understanding of this intricate system.
2 Proteomic Approaches for Studying Ubiquitylation
To date, systems-wide analyses of ubiquitylated proteins (hereafter referred to as the ubiquitome) are mainly based on three major types of proteomic approaches: mass spectrometry analysis, in vitro protein/peptide arrays, and the newly arising bioinformatics methods. More and more studies combine two or, indeed, all three of these methods. In this chapter, we will mainly focus on the study of the ubiquitome using mass spectrometry and give a brief overview of the other two approaches.
2.1 The Systems-Wide Study of the Ubiquitome by Mass Spectrometry
Most mass spectrometry-based protein analyses use a bottom up approach in which proteins are digested into peptides, which are then separated by liquid chromatography prior to their analysis by the mass spectrometer . Using tandem mass spectrometry (MS/MS), peptides are further fragmented into second order mass spectra, which act as “fingerprints” that are subsequently searched against a database to obtain a potential sequence match. The peptides can, in turn, be used to identify the proteins from which they are derived. Mass spectrometry can thus be used to identify the protein composition of the analyzed biological sample, and, in some cases, also to quantify differences between two or more samples [23].
Analysis of PTMs is well suited for mass spectrometry, as in most cases modified peptides can be characterized by a difference in mass (compared to their unmodified counterparts) that can be detected by the instrument. In the case of ubiquitylation, a characteristic amino acid “tail” is left on the lysine residue of the substrate peptide, which was first used to detect in vivo ubiquitylation sites by mass spectrometry in 1993 [24]. Different proteases have been used to generate peptides with remnant ubiquitin tails for mass spectrometry analysis [24]. However, due to its robust catalytic activity and cleavage site specificity, trypsin is now the most commonly used enzyme (among many other applications) for proteomic studies of ubiquitylation. Trypsin is a serine protease that cuts peptide chains on the carboxyl side of lysine and arginine. Because the carboxy-terminal end of ubiquitin has the amino acid sequence Arg-Gly-Gly, tryptic digestion of ubiquitylated peptides will generate a “tail” consisting of the last two glycine residues of ubiquitin. This di-glycine “tail” (hereafter referred to as the remnant, or diGly tail) adds a signature mass shift of + 114 Da on the modified lysine residues. At the same time, trypsin is unable to cleave ubiquitylated lysines, and thus modified peptides also contain a missed-cleavage site (Fig. 14.1) . One potential problem is that the same mass shift of 114 Da can also occur due to the alkylation of lysine residues during sample preparation when using iodoacetamide (commonly used to protect reduced cysteine residues prior to mass spectrometry) [25]. This is favored even more when using high concentrations of alkylating reagents and heat [26]. Therefore, sample alkylation is commonly performed at no more than room temperature, and iodoacetamide is substituted by chloroacetamide, which generally does not cross-react as readily.
The diGly signature. Trypsin digestion of ubiquitylated proteins leaves a distinctive di-glycine (diGly) signature attached to the modified lysine residues owing to the presence within ubiquitin of an arginine residue directly preceding the two carboxyl-terminal glycines. The diGly-containing peptide will also include a missed-cleavage site, since ubiquitylation blocks trypsin digestion next to the modified lysine
By the early 2000s, most of the effector-enzymes (E1s, E2s, E3s, DUBs) in the ubiquitin system had been uncovered [27]. However, only a small fraction of their substrates were characterized. To fill in the gap—and taking advantage of major progresses in mass spectrometry instrumentation—systems-wide proteomic approaches were developed. Given the transient, and mostly low-abundant nature of ubiquitylation in the cell, conjugated proteins and their ubiquitylation sites are difficult to detect by mass spectrometry analysis without any pre-enrichment. Therefore, the depth and coverage of the ubiquitome is tightly dependent on the enrichment method. In this section, we will present the hallmark systems-wide approaches developed to study the ubiquitome, including the most current methods, with an emphasis on the enrichment techniques .
2.1.1 Enrichment of Ubiquitylated Proteins Using Tagged Ubiquitin
Taking advantage of yeast genetics, Steven Gygi and colleagues were among the first to intracellularly express tagged ubiquitin, enabling them to enrich for ubiquitin conjugates for large-scale analysis of the ubiquitome [28]. The authors used a yeast strain in which an ectopic 6xHis-myc-ubiquitin was expressed instead of wild-type ubiquitin to purify ubiquitin conjugates under denaturing conditions using nickel-affinity chromatography (Fig. 14.2a). In this landmark study, 1075 proteins were identified by mass spectrometry , including 110 ubiquitylation sites (i.e., diGly-containing peptides) in 72 proteins. Because a certain level of contaminating proteins still co-purified along with the ubiquitylated proteins, even under denaturing conditions, the identification of ubiquitylation sites using the characteristic remnant ubiquitin tail was key in this study. One concern which could be raised regarding this method—and several other approaches described below—is that it is based on the ectopic expression of a tagged ubiquitin, which could result in the disruption of ubiquitin levels and/or hindrance of its amino terminus, thereby potentially differentially influencing the ubiquitylation rates in distinct pathways or substrates (depending on which ubiquitylating enzymes are affected).
Workflow diagram for three different basic enrichment methods. The main advantages and disadvantages of each method are shown below each diagram. a Tandem histidine tagged-ubiquitin-based enrichment of ubiquitylated proteins. b UBD-based (Ubiquitin-binding domain-based) enrichment of ubiquitylated proteins. c Anti-diGly antibody-based enrichment of ubiquitylated peptides
Analysis of ubiquitin conjugates is not constrained to single cell model organisms. In order to pull down ubiquitylated proteins from a specific tissue of a whole animal, Ugo Mayor and colleagues used the GAL4/UAS tissue-targeted expression system in Drosophila melanogaster [29]. In this study, ubiquitin tagged with the BirA recognition sequence was over-expressed solely in the nervous system, together with the Escherichia colibirA gene that encodes a biotin ligase, to biotinylate tagged ubiquitin. Consequently, 48 novel neuronal-specific ubiquitylation substrates were identified in this pioneering proteomic study conducted in a multi-cellular organism.
To further reduce the levels of non-specifically bound proteins (binding to the tag or to the affinity column) during the enrichment of tagged ubiquitin, which is notoriously problematic with the histidine-tag system, different approaches have subsequently been used. For instance, Peter Kaiser’s group developed a tandem affinity tag approach by employing a two-step purification, using 6xhistine and biotin tags [30]. In the second step, the high affinity between biotin and streptavidin allows the enrichment of targets under very stringent conditions, such as 2 % sodium dodecyl sulfate (SDS) and 8 M urea. Over 150 ubiquitylated proteins were identified under these conditions. This approach was then applied to mammalian tissue cultures to identify over 600 ubiquitylated conjugates in HeLa cells [31]. In our lab, we instead utilized an 8xhistidine tag that enabled us to use up to 0.5 % SDS for washing in a single-step purification scheme (Fig. 14.2a) [32]. We performed several control experiments in which we mixed differentially labeled cells (using either 14N or 15N stable isotopes) that expressed (or did not express) the tagged ubiquitin, and we showed that, under these conditions, most purified proteins were, indeed, specifically conjugated to the tagged ubiquitin. We identified on average 200 ubiquitylated proteins within a 4-h mass spectrometry analysis. However, only a small number of ubiquitylated peptides containing the diGly remnant tail were identified in both of the above-mentioned studies (mostly derived from ubiquitin and only a few other proteins). It is not entirely clear why this is the case. One possibility is that the sample fractionation conditions used in these two studies were not optimal for peptides containing the ubiquitin remnant tail . By contrast, strong cation exchange was used by Stanley Fields and colleagues in order to further fractionate ubiquitylated peptides after nickel chromatography (also using a 8xhistidine tagged ubiquitin), as the amino group of the remnant ubiquitin tail adds an additional positive charge at low pH in comparison to other peptides [33]. In their study, 870 ubiquitylation sites were identified among 438 proteins. The number of analyzed fractions may also influence the output. After tagging ubiquitin with a tandem tag (streptavidin and hemagglutinin, HA) and separating the purified proteins through gel electrophoresis, Danielsen and colleagues identified over 700 ubiquitylation sites after analyzing 20 gel fractions [34]. Overall, due to its relative simplicity, the purification of proteins conjugated to histidine-tagged ubiquitin (as well as to other tags) remains widely used, but precautions should be taken in order to avoid contaminating materials.
2.1.2 Enrichment of Ubiquitylated Proteins Using Ubiquitin-binding Domains (UBDs)
Because of their ability to bind to ubiquitin, UBDs can be employed to enrich for ubiquitylated conjugates . Since some UBDs specifically bind to distinct types of ubiquitin chains, these can be further exploited to enrich for a subset of ubiquitin conjugates. For instance, the yeast proteasome adaptor proteins Rad23 and Dsk2 have ubiquitin-associated domains (UBAs) which have a higher affinity for poly-ubiquitin chains over free ubiquitin and monoubiquitylated proteins (Fig. 14.2b) [35] .
In 2005, Raymond Deshaies’ group was the first to use UBDs to identify ubiquitylated proteins by mass spectrometry by a two-step affinity purification method [36]. The aim of this study was to identify proteins that require the non-essential Rpn10 proteasome receptor for degradation in yeast cells. To enrich for poly-ubiquitylated proteins targeted to the proteasome, recombinant proteins (Rad23 and Dsk2) that bind to proteasome substrates [37] were used for a first affinity purification under native conditions followed by a second purification step under denaturing conditions, using 6xhistine tagged ubiquitin expressed in the cells. One hundred and twenty-seven proteins were identified as candidate substrates of the proteasome in wild type cells, and an additional 50 or so proteins were only identified in the absence of Rpn10, indicating that they likely require this factor for degradation. This approach was further developed using isotope labeling to more unequivocally identify proteasome substrates in the cell [38]. Other proteomic studies used proteasome adaptor proteins to enrich for ubiquitylated conjugates. For instance, using this approach, Ron Kopito and colleagues found that the overexpression of a mutated fragment of huntingtin (which has been linked to Huntington’s disease) led to the accumulation of several poly-ubiquitin chain types in tissue culture cells [39] .
The group of Manuel Rodriguez developed tandem ubiquitin-binding entities (TUBEs) in order to enrich for ubiquitylated proteins [40]. Notably, this approach relies solely on endogenous ubiquitin to identify conjugated proteins. TUBEs were developed by fusing several UBAs together to increase affinity for proteins conjugated to poly-ubiquitin chains (Fig. 14.2b). TUBEs can also inhibit the activity of DUBs and of the proteasome, preserving ubiquitin chains during the native purification process. Using this approach, Rodriguez and colleagues identified a total of 643 proteins in two biological replicates from human breast adenocarcinoma cells treated with the DNA damage-inducing agent Adriamycin [41]. The ability to enrich for ubiquitin conjugates without ectopic expression of a tagged ubiquitin has great potential, especially for animal models.
One major issue is that many other proteins may interact with ubiquitylated proteins or the recombinant UBD-containing proteins under native conditions. While a high salt concentration was used (2 M NaCl) in the first study mentioned above, a second step purification was required to further enrich for ubiquitylated proteins [36]. It is actually challenging to directly circumvent this issue, since the ubiquitin-UBD interaction is mainly mediated by hydrophobic interactions [19], thus the usage of anionic detergents or chaotropic reagents would be detrimental. Therefore, UBDs may be better suited in the future to analyze subsets of the ubiquitome (taking advantage of the specificity of the recombinant UBD for a particular chain type) in combination with a second purification step .
2.1.3 Enrichment of Ubiquitylated Peptides Using α-diGly Antibodies
A major breakthrough was the introduction of antibodies that directly bind to ubiquitylated peptides. After the first 110 ubiquitylation sites identified by Gygi and colleagues [28], the uncovering of new ubiquitylation sites had largely stalled. One issue was that in that particular analysis in-depth study required 5 days of mass spectrometer instrument time, which is difficult to secure with equipment that is typically oversubscribed. By contrast, identification of phosphorylation sites benefited from tremendous activity starting in 2006, after TiO2 beads began to be widely used to enrich for phosphorylated peptides [42–44]. Fortunately, the ubiquitin field was not idle, and monoclonal antibodies were soon developed to specifically enrich for ubiquitylated peptides.
In the antibody-based approach, ubiquitylated peptides of low abundance are greatly enriched prior to identification by mass spectrometry using antibodies that recognize the ubiquitin remnant tail left on trypsin-cleaved peptides. The laboratory of Samie Jaffrey was the first to publish an antibody-based approach to enrich for diGly peptides in 2010 (Fig. 14.2c) [45]. In this study, 374 ubiquitylation sites on 236 proteins were identified from HEK293 cells. To generate the antibodies, the authors synthesized a lysine-rich protein antigen (histone) containing multiple K-ε-GG that was then injected into mice. In the following year, both Steven Gygi’s and Chuna Ram Choudhary’s groups successfully and independently conducted notable large scale studies: around 19,000 ubiquitylation sites in ~ 5000 human proteins and ~ 11,000 sites in ~ 4200 proteins were mapped, respectively [46, 47]. Several additional studies followed up using the same approach, including analyses of the ubiquitome in rat brain and murine tissues [48, 49]. In addition, several modifications have been made in order to increase the yield of this method [50]. One reason the diGly-antibody approach is so potent is that trypsin digestion essentially abolishes most other protein-protein interactions and, combined with the high-affinity of the antibody, modified peptides are effectively enriched, despite their low abundance in the cell. Other major advantages are that no extra experimental controls are required to distinguish ubiquitylated sites from the rest of the identified peptides in the sample (the detection of the + 114 Da mark is sufficient), it does not rely on ectopic expression of a tagged ubiquitin, it is applicable to all eukaryotic organisms or tissues and it is commercially available in the form of a kit.
While the antibody-based approach is very potent and already widely used, it also has a few shortcomings. Since the NEDD8 and ISG15 Ubls share the same carboxyl-terminal sequence (RGG) with ubiquitin, proteins conjugated to these Ubls will also generate indistinguishable remnant tails after tryptic digestion. Therefore, some ubiquitylated peptides may be mis-assigned. Fortunately, levels of ISG15 are usually undetectable in cells, unless stimulated by interferon (IFN)-α/β [51]. Furthermore, Gygi’s group determined that more than 95 % of the sites which they identified were conjugated to ubiquitin and not NEDD8 (which is mainly conjugated to cullins). The monoclonal antibodies against the ubiquitin remnant may also introduce a bias for some sites, since Choudhary’s group found that these antibodies have a slight sequence preference [49]. Our lab also noted that, using this approach, proteins ubiquitylated at multiple sites may be less prevalent compared to proteins conjugated at a single lysine, as peptide ion intensities would be lower (and thereby possibly not detected by the mass spectrometer) in the former case [52]. Another consideration is that the information regarding the chain linkage on a particular conjugation site (for poly-ubiquitylation) is lost when using this approach. In addition, atypical sites (N-terminal, for instance) are also not selected. Nevertheless, the antibody-based approach has been the key to the recent advancements and has now been adopted by many groups around the world.
2.1.4 An Alternative to the Antibody-Based Approach
The price of commercially available antibodies can be an obstacle, prompting researchers to look for more cost-accessible alternatives. One approach which has shown promise in preliminary trials makes use of the fact that diGly-containing peptides have two N-terminal primary amines (one from the peptide N-terminus and the second from the diGly remnant) that can be modified and exploited further for enrichment. Primary amines can be fluorinated using perfluorinated compounds, while the ε-amino groups of lysines are protected by guanidination [53, 54]. The doubly labeled (and thus containing a diGly signature) peptides have a significantly higher affinity than the singly labeled ones for a matrix which retains fluorinated compounds, thus allowing the separation of the two species by eluting with different concentrations of organic solvent [55]. Fluorous affinity tag enrichment was successfully used to isolate diGly containing peptides from a tryptic digest of pure poly-ubiquitin chains [56]. Later studies have employed reversed-phase chromatography to enrich peptides with large fluorinated moieties attached to cysteine residues [57, 58], but this approach has not yet been used for diGly enrichment. The challenge is now to apply this method to the isolation of diGly-containing peptides from complex samples of biological origin.
2.1.5 Systems-Wide Analysis of Ubiquitin Linkages Using Selective Reaction Monitoring (SRM)
In addition to identifying substrate ubiquitylation sites, it is also important to determine the modification type (i.e., mono- vs. poly-ubiquitylation) . One particularly effective approach utilizes selective reaction monitoring (SRM) to determine, for instance, which chain linkages are present in a cell extract, or synthesized by a given E3 ligase in an in vitro experiment. In SRM, the mass spectrometry instrument is set up to specifically monitor one or several preselected peptide(s), greatly improving both the sensitivity and identification rate from complex biological samples [59–63]. In combination with SRM, artificial peptides (of known concentration) can be added in for absolute quantification (AQUA) [64]. The synthetic peptides are identical to the endogenous peptides of interest, but carry a specific isotopic mark (typically “heavy” lysine or arginine residues, giving a + 8 or + 10 Da mark, respectively). The AQUA peptides co-elute with their counterparts within the sample, and their intensities can be compared to calculate the concentrations (or the absolute quantification) of the peptides of interest. To quantify chain type, typically three peptides are required for each of the seven lysine residues of ubiquitin: one longer peptide, which includes the ubiquitylated lysine with a remnant diGly tail , and two peptides corresponding to the N- and C-terminal parts of the former, on either side of the (in this case non-ubiquitylated) lysine. In this way, a quantification of the relative amounts of modified and unmodified peptide is possible (Fig. 14.3) .
The absolute quantification (AQUA) peptide-SRM (selective reaction monitoring) approach to the quantification of ubiquitin chain linkages. a A mixture of different ubiquitin chains obtained from a biological sample is digested along with known amounts of absolute quantification (AQUA) peptides. The AQUA peptides are isotopically labeled (13C/15N) to distinguish them from their endogenous counterparts. Three peptides are monitored for each linkage type: the diGly-containing peptide and the N- and C-terminal parts of the former that occur by tryptic digestion of an non-ubiquitylated parent peptide. b Tandem mass spectrometry through which only specific charge-to-mass (m/z) ratios are monitored—termed selected reaction monitoring (SRM)—is used in order to obtain a ratio between the endogenous peptides and the artificial standards. By using AQUA peptides that cover both the modified (diGly containing) and the unmodified states, an absolute quantification of the amount of a particular linkage, as well as the comparison between different linkage types, is possible
The AQUA approach was pioneered in the workgroup of Steven Gygi [65] and has since then been successfully used to quantify ubiquitin chain linkages under a variety of conditions and model systems. For instance, it was used to determine which ubiquitin chains were conjugated in vitro onto the cell cycle-regulated protein cyclin B1 [66]. The method was used in numerous studies to quantify in vivo ubiquitin chain linkages in yeast cells [67], in cultured mammalian cells [68–70] and in mice [71]. Notably, AQUA and related methods have helped to explore the role of ubiquitin chain linkages whose biological role were not well understood (see below in Sect. 3.2). These methods have been adopted and further developed by numerous groups. However, implementation of this approach on each proteomic platform and instrument still required dedicated researchers to perform those types of experiments .
Kopito’s group developed an interesting, closely related method that employed spiked-in proteins for Protein Standard Absolute Quantification (PSAQ) [70]. Compared to AQUA, this method comes with a twist, namely that the spiked “standards” are differentially labeled ubiquitins instead of peptides. This allows the monitoring of a different pool of proteins within a sample, namely free ubiquitin, monoubiquitylated and poly-ubiquitylated proteins. Using this approach, the authors were able to show that the majority of conjugated proteins in the cell are, surprisingly, monoubiquitylated and not poly-ubiquitylated .
2.1.6 Systems-Wide Analysis of the Ubiquitome Using Approaches that are Not Based on Mass Spectrometry
Protein array technology can also be used to perform proteomic analysis of the ubiquitin system. One major task is to delineate the substrate specificity of the different enzymes involved in ubiquitylation. This is often difficult to tackle in vivo, because of the diversity of the ubiquitin system and the transient nature of ubiquitylation (due to de-ubiquitylation). The recent development of protein arrays containing purified recombinant proteins from a variety of organisms, ranging from yeast to human, allows researchers to take an in vitro approach in a systems-wide manner. So far, this strategy has been applied to the study of substrates of ligases such as Rsp5 in yeast [72, 73], Nedd4 − 1 and Nedd4 − 2 and the anaphase-promoting complex (APC) in humans [74, 75], and a panel of human DUBs [76]. Its cell-free nature endows the in vitro protein microarray analysis with a number of advantages, but it also has the weakness of decoupling the ubiquitylation reactions from a cellular context (e.g., localization, physiological concentration). In a recent study, Marc Kirschner and colleagues used protein arrays incubated with a cell extract to identify which proteins are conjugated to ubiquitin and other probed Ubls (e.g., SUMO, NEDD8, and ISG15) [77]. Surprisingly, almost entirely distinct subsets of proteins were targeted by each conjugation system. Apart from proteins, peptide arrays have also been used to identify substrate candidates, as many E3s rely on distinct domains to recruit their targets, like the ligand of Numb protein-X (LNX) E3 ligase that contains a PDZ domain [78]. Additional genome-wide approaches have also been developed that do not rely on protein arrays. For instance, Stephen Elledge and colleagues have used a green fluorescent protein (GFP)-fusion library in tissue culture cells to identify substrates of cullin-based E3 enzymes, which form a major ligase family [79].
Another approach to deciphering the ubiquitome is based on bioinformatic site prediction. The first algorithm, called UbiPred, was developed by Ho’s group in 2008. It used 531 physicochemical properties and a limited training set of 157 ubiquitylation sites and 3676 putative non-ubiquitylation sites from 105 proteins for building the prediction algorithm [80]. Recent advances in experimental investigation have helped to develop several additional ubiquitylation site prediction programs, such as UbPred, weighted passive nearest neighbor algorithm (WPNNA), and composition of k-spaced amino acid pairs (CKSAAP)_UbSite, Ubiprober [81–84]. These prediction algorithms have an overall reasonable accuracy. However, due to the fact that there is no universal target sequence for ubiquitylation sites, their usage may be limited. It may be more appropriate to use these approaches to identify candidate substrates for a given ubiquitin ligase. For instance, several algorithms can be used to predict substrates of the APC E3 ligase based on the presence of specific motifs [85, 86]. In one case, a combination of both mass spectrometry and bioinformatics analyses was used to narrow down a list of candidate substrates for the Fbw7 cullin-based E3 ligase [87]. More general computational approaches have increasingly been used to probe the ubiquitome in the past few years, as researchers began to obtain more and more data. We will highlight several of these studies in the next section.
3 Contributions of Systems-Wide Proteomic Approaches to Gaining Insights into the Ubiquitin System
The diverse systems-wide approaches which we have described so far have led to a significantly better understanding of the ubiquitin system, often by providing unique information that could not have been obtained by other means. We will review several key contributions in the last section of this chapter.
3.1 Ubiquitylation Sites
As mentioned earlier, there is no specific motif that directs ubiquitylation. However, several residues are more frequently present (or absent) at certain positions near to the ubiquitylation sites. The absence of a universal ubiquitylation motif can be explained by the fact that there is a large variety of different E3 ligases, each of which are likely to recognize their target in a distinctive way. In addition, E3 ligases do not directly bind to the ubiquitylation sites, but typically to motifs that are adjacent to the conjugated lysine residues. Because the lysine residue directs the nucleophilic attack that leads to its conjugation, nearby residues may chemically influence the reaction. Indeed, Kim and colleagues found that several acidic residues were enriched while basic residues were instead depleted in the vicinity of ubiquitylation sites [46]. Cysteine residues were also less prevalent, since they would likely compete as ubiquitin acceptor residues. Ubiquitylation sites are also slightly more conserved than unmodified sites and, interestingly, often appeared earlier in evolution (i.e., prior to the adoption of ubiquitylation) [88]. The latter observation indicates that the ubiquitin system likely accommodates preexisting sites rather than triggering the appearance of new sites. Intriguingly, ubiquitylation sites were also often found in ordered regions in contrast to phosphorylation sites that are more prevalent in regions predicted to be disordered (i.e., without a specific secondary structure) [89]. It will be important to reexamine some of these observations in the near future as more data is generated.
3.2 Ubiquitin Chains and the Ubiquitin Code
Ubiquitylation supports numerous cellular responses or functions, and the formation of different chain types has long been thought to play a major role in directing conjugates to their final intracellular fates . The targeting of short-lived proteins for degradation is a major function of the ubiquitin system [90], and the K48-linked poly-ubiquitin chain was the first chain type associated with the turnover of these proteins [91]. In contrast, K63-linked ubiquitin chains have distinct cellular functions, like DNA repair and endocytosis [92, 93]. These first landmark studies were key to laying down the now well-accepted hypothesis that chain linkage types determine, at least partially, the fate of the substrates. However, major questions remain unanswered: What is the function of the other chain types (apart from K48 and K63)? and How do chain linkage amounts fluctuate in the cell, especially in response to stress? As these questions are difficult to investigate solely by using conventional approaches (e.g., ubiquitin mutants that cannot form certain chain types), the development of new proteomic approaches will potentially open up new horizons .
Several chain types participate in proteasome degradation. Gygi’s group demonstrated for the very first time that all seven lysines in ubiquitin contribute to the assembly of poly-ubiquitin chains in vivo [28]. Subsequently, it was also found that linear ubiquitin chains (linked via the ubiquitin amino terminus; M1) were present in mammalian cells [94]. Using a SRM-based method, Peng and co-workers first determined the presence of each chain linkage type in yeast cells and found that K11 chains were almost as prevalent as K48 chains [67]. To assess the role of each chain type in proteasomal degradation, the authors then inhibited the proteasome and found out that all chain types rapidly accumulated in the cell, except for K63 chains, and K27 and K33 were affected to a lesser extent. To confirm the role of K11 in proteolysis, they also showed that K11-linked chains were important for the endoplasmic reticulum-associated degradation (ERAD) pathway, which targets misfolded endoplasmic reticulum (ER) proteins [67]. K11 linkages were also found to be important for the proteasomal degradation of cell cycle-regulated proteins in an independent study [95]. However, this linkage is less prevalent in mammalian tissue culture cells [69]. Indeed, in higher eukaryotes, K48 linkages make up about 80 % of the poly-ubiquitin chains targeted to the proteasome, while K29, K11 and possibly K6 were found (in this order) in decreasing concentrations [69]. Nonetheless, K11 chains further accumulate in brain tissues from patients suffering from Alzheimer’s disease (as do K48 and K63 chains, albeit to a lesser extent), thus confirming that these chain types play a prevalent role in the cell. Furthermore, the function of K11 linkages may not be limited to proteasomal degradation. For instance, the major histocompatibility complex class I (MHC I) was found conjugated to both K11- and K63-linked ubiquitin chains that are involved in the endocytosis of MHC I [96, 97]. In another study, the presence of K33-linked chains was confirmed by mass spectrometry on T cell receptor-zeta, another membrane protein that is regulated by endocytosis [98]. This is intriguing, because endocytosis was originally thought to be mediated by mono- and K63-linked poly-ubiquitylation. While these studies greatly expanded our knowledge, one question that remains is whether more complicated chain architectures (i.e., branched chains) are also present in the cell. New techniques will have to be developed to answer this question, as this information is lost with current sample preparation methods .
3.3 Protein Quality Control and the Ubiquitin Proteasome System
A major function of the ubiquitin proteasome system is to eliminate misfolded and damaged proteins from the cell with the assistance of several protein quality control pathways [99] . One striking observation is that the majority of proteins targeted for degradation are newly synthesized [46]. It has already been shown that a large fraction (up to 30 %) of newly translated proteins in tissue culture cells is rapidly degraded by the proteasome, presumably because these proteins were not properly folded [100]. A recent study from Jon Huibregtse’s group confirmed that about 12 % of nascent polypeptides are ubiquitylated [101]. The ubiquitylation of these nascent polypeptides occurs on both stalled and actively translating ribosome complexes. To further characterize these conjugates, an elegant approach was used, in which tandem affinity purification of ubiquitylated nascent polypepides was achieved from purified polysomes by using both flag-tagged ubiquitin and biotinylated puromycin, which forms a covalent bond with the carboxyl terminus of nascent polypeptides prior to releasing them from ribosomes. Using mass spectrometry , the authors found that the majority of these ubiquitylated nascent polypeptides corresponded to cytosolic proteins [101] .
The impairment of the ubiquitin proteasome system has been associated to many aggregation-related disorders, especially neurodegenerative diseases [102, 103], which typically feature ubiquitin-containing aggregates [104]. We found that inhibition of the proteasome can cause the formation of large ubiquitin-containing aggresomes (which are amorphous structures mainly composed of misfolded proteins) even in the absence of disease specific proteins (e.g., mutated cystic fibrosis transmembrane conductance regulator, CFTR, involved in cystic fibrosis) [105]. To further characterize these ubiquitin-containing protein aggregates induced by proteasome inhibition, we combined a biochemical isolation method with quantitative proteomic analysis using stable isotope labeling with amino acids in cell culture (SILAC) [106]. We identified more than 500 proteins in these aggregates, including the p62/sequestosome, several E3s and DUBs, and chaperones. One possibility is that, in addition to those involved in the aggregation process, many of these proteins are aggregation-prone and would normally be efficiently eliminated by the proteasome .
The different protein quality control pathways form an intricate network and keep the proteome in check. While many studies at the time relied on the assessment of a few model substrates, we instead focused on the effect of the heat-stress response on the ubiquitinome in a systems-wide manner. Heat shock that causes protein misfolding has long been known to cause an increase in poly-ubiquitylation in the cell [107]. However, we only recently found that proteins ubiquitylated after heat shock were cytosolic and in part targeted by the Hul5 ubiquitin ligase [32]. By using bioinformatics tools to further analyze our proteomic datasets, we found that several features were shared by the heat-induced ubiquitylated proteins [52]. Notably, these proteins are longer on average and contain fewer hydrophobic residues. Interestingly, intrinsically disordered proteins (that contain large regions predicted not to fold in a specific secondary structure) are also prominently identified among proteins ubiquitylated upon heat shock [52]. We also found that these proteins while generally not essential and less abundant in the cell, contain more predicted interaction motifs in their disordered regions. Presumably, these proteins are ubiquitylated upon heat shock due to the loss of interactions with their binding partners. It is not clear whether this is triggered by a specific protein quality control pathway, or whether these disordered proteins may themselves also be important for the heat-shock response. In addition to heat-shock response, the study of the ubiquitome may be particularly interesting when assessing client proteins of a given chaperone protein. In absence of the chaperone’s activity, its client proteins should misfold and at least a portion of them will be ubiquitylated for proteasome targeting .
3.4 The Quest to Find Ubiquitin Ligase Substrates
One of the major challenges to the study of the ubiquitin proteasome system is to assign substrates to ubiquitin ligases . Several proteomic approaches have been developed in which individual E3s—or components of E3s—are pulled down to identify their substrates, a method that was notably championed by the group of Michele Pagano [108–114]. In each case, follow-up studies are required to verify that the interacting proteins are indeed E3 targets. To further enrich for substrates, inactive E3 ligases were also employed [115]. In another case, the E3 ligase was re-engineered to modify its specificity and conjugate NEDD8 instead of ubiquitin [116]. Alternatively, the substrates of a specific E3s could be identified by analyzing which proteins are no longer ubiquitylated upon the inhibition or deletion of the investigated ligase. For instance, Kim and colleagues monitored changes in the ubiquitome to identify substrates of cullin-RING ubiquitin ligases by comparing cells with and without treatment with the MLN4924 inhibitor that blocks NEDD8 conjugation (neddylation is required to maintain cullin-RING ligases in an active state) [46]. They consequently identified over 4000 ubiquitylation sites that were dependent on cullin-RING ligase activity .
However, cullin-RING ligases form the largest family of E3s. Therefore, the identified proteins may be targeted by any of the 350 or so E3s. Our lab instead focused on identifying candidate substrates of a single ubiquitin ligase, the yeast HECT ligase Hul5. By comparing wild-type and deletion mutant yeast cells, we discovered over 90 putatively misfolded substrates of Hul5, and thereby revealed that this ubiquitin ligase plays an important role in cytosolic protein quality control [32].
More recently, the Parkin-dependent ubiquitome was analyzed under different stress conditions [117]. The Parkin ubiquitin ligase has been linked to Parkinson’s disease (PD) and several Parkin mutations are associated to a familial and early-onset form of PD. In this study, ubiquitylated peptides were enriched using the anti-diGly antibodies prior to quantitative mass spectrometric analysis to compare cells that either did or did not express Parkin under mitochondrial depolarization conditions. The study revealed that Parkin expression dramatically altered the ubiquitylation status of the mitochondrial proteome [117], consistent with its role in mitophagy [118]. These studies illustrate the great potential of analyses of the ubiquitome in linking components of the ubiquitin machinery to their functional roles and biological substrates .
3.5 Chasing Lost Ubiquitylation Events and DUB Functions
DUBs that cleave off the C-terminal carboxyl group of ubiquitin from the ε-amino group of lysine side chains or the α-amino group of conjugated proteins are essential to the ubiquitin system [15]. There are over 80 genes in the human genome predicted to have de-ubiquitylating activity. Because of their role in processing ubiquitin precursors and recycling ubiquitin, DUBs play a key role in maintaining ubiquitin homeostasis. In addition, many DUBs display specificity both for particular ubiquitin chain types, as well as to certain substrates. Altered DUB activity has been linked to several diseases, such as the Machado-Josephdisease (MJD), microcephaly-capillary malformation (MIC-CAP) syndrome, and coronary artery disease(CAD), as well as numerous forms of cancer [119–121]. Therefore, DUBs are considered to have great therapeutic potential as drug targets [122] and the identification of the substrate repertoires and functions of DUBs is urgently needed.
Mass spectrometry has been applied to the study of DUBs in a systems-wide manner. For instance, Wade Harper and colleagues expressed and pulled down 75 human DUBs to identify their interacting partners by mass spectrometry [123]. A similar study was performed with the 20 DUBs of Saccharomyces pombe [124]. More recently, Gardner’s group used in vivo cross-linking prior to co-immunoprecipitation of the yeast DUB Ubp10 in order identify transiently interacting proteins, including ribonucleic acid (RNA) polymerase I, which was later confirmed to be a substrate of Ubp10 [125]. Alternatively, the change in proteome composition was analyzed by quantitative mass spectrometry to identify potential DUB substrates accumulating upon deletion of each of the 20 yeast DUBs [126].
Another approach consists in trapping DUBs using so-called suicide substrates, in which a thiol-reactive group is placed at the C-terminus of ubiquitin to ligate it irreversibly to the DUB’s active site cysteine. Kessler and colleagues used this method to identify DUBs and their interacting proteins by mass spectrometry from mouse tissue [127]. This method, together with quantitative mass spectrometry , was also used to characterize chemical DUB inhibitors to gauge their selectivity, as inhibition of a specific DUB results in less binding to the activity-based probe [128]. One of the tested compounds, P22077, was selective to Ubiquitin-specific-processing protease 7 (USP7), thereby causing a lower rate of association of USP7 to the suicide substrate. Using TUBEs and mass spectrometry, Kessler and colleagues then identified potential substrates of USP7 upon its inhibition [128]. However, researchers have not yet taken full advantage of newly developed proteomic approaches to study DUBs and deubiquitylation in vivo. In the near future, combining SRM and diGly approaches to examine ubiquitin chain linkages and ubiquitylated proteins upon the inhibition or down-regulation of a specific DUB (or, conversely, its over-expression) will help greatly with building a comprehensive knowledge base of DUBs.
3.6 Cross-Talk Between Ubiquitylation and Other PTMs
Multiple PTMs may occur on the same protein and the cross-talk between different functional modulators may be important for protein activity, function, and localization. Histone tails are a well-known example: Being particularly lysine-rich amino acid sequences, they are subjected to a variety of modifications, such as acetylation, methylation and ubiquitylation (on lysine), methylation of arginine, and phosphorylation of serine and tyrosine residues. Different combinations of these PTMs generate the “epigenetic code” and have distinct functions in regulating chromatin organization and accessibility. Thus, the modifications are set in a highly controlled manner [129]. Similarly, the selectivity of the ubiquitin ligases for their substrates is also tightly regulated. In many cases, substrates first need to be modified to trigger their recognition by an E3 ligase and phosphorylation plays a prominent role in this process.
The group of Steven Carr recently established a method (called serial enrichments of different posttranslational modifications, SEPTM) for the serial enrichment of PTMs, such as phosphorylation, ubiquitylation, and acetylation, from the same sample [130]. Phosphorylated peptides were first isolated using the IMAC phosphor-enrichment method followed by the diGly antibody-based method for ubiquitylation enrichment. The same sample was then subjected to the enrichment of acetylated peptides, also using an antibody-based approach. SEPTM enabled the identification of more than 20,000 phosphorylation, 15,000 ubiquitylation, and 3000 acetylation sites from around 8000 proteins and provides a unique opportunity to study the cross-talk between these three different PTMs. Using quantitative mass spectrometry, the authors analyzed changes incurred by proteasome inhibition and showed that the number of phosphorylation sites, as well as the number of ubiquitylation sites, were highly increased. These results confirmed that phosphorylation plays a major role in regulating proteolysis. Similar data were obtained in an independent study from Villen’s group [131]. Here the authors first enriched ubiquitylated proteins using His-tagged ubiquitin followed by the isolation of diGly-containing peptides and phosphorylated peptides. Using this approach, they identified a total of 466 proteins that were ubiquitylated and phosphorylated. While the non-ubiquitylated forms of these proteins were also found to be phosphorylated, proteins that are both ubiquitylated and phosphorylated, and accumulate upon proteasome degradation, have, on average, shorter half-lives, indicating that phosphorylation is likely involved in the regulation of their turnover. By contrast, the acetylome analyzed in the former study was not significantly affected by proteasome inhibition. However, around 400 of the acetylation sites were also found to be ubiquitylated, indicating that cells contain distinctly modified populations of the same proteins. Therefore, cross-talk can be cooperative (multiple PTMs on the same molecule) or competitive (modifying the same site). It will be interesting to further integrate additional PTMs to identify which other modifications may impact ubiquitylation positively or negatively.
4 Concluding Remarks
In summary, systems-wide analyses of the ubiquitome have uniquely contributed to and shaped our understanding of the ubiquitin system. Within a few years, we gained tremendous insights into the ubiquitome. A broad panel of different methods is now available to the scientific community. The further integration of these methods will undoubtedly play an important role in deciphering disease mechanisms linked to ubiquitylation, and potentially make great contributions towards future therapeutic development.
Abbreviations
- AQUA:
-
Absolute quantification (of protein)
- diGly tail:
-
Two glycine from the C terminus of ubiquitin covalently attached to target’s lysine
- DUB:
-
De-ubiquitylating enzyme
- E1:
-
Ubiquitin-activating enzyme
- E2:
-
Ubiquitin-conjugating enzyme
- E3:
-
Ubiquitin ligase
- ERAD:
-
Endoplasmic reticulum-associated degradation
- ISG15:
-
Interferon-stimulated gene of 15 kDa
- MHC I:
-
Histocompatibility complex class I
- MS/MS:
-
Tandem mass spectrometry
- NEDD8:
-
Neural-precursor-cell-expressed and developmentally down-regulated gene 8
- PSAQ:
-
Protein standard absolute quantification
- PTM:
-
Posttranslational modification
- SEPTM:
-
Serial enrichments of different posttranslational modifications
- SILAC:
-
Stable isotope labeling with amino acids in cell culture
- SUMO:
-
Small ubiquitin-like modifier
- TUBEs:
-
Tandem ubiquitin-binding entities
- UBA:
-
Ubiquitin-associated domain
- UBD:
-
Ubiquitin-binding domain
- Ubl:
-
Ubiquitin-like protein
References
Hornbeck PV, Kornhauser JM, Tkachev S et al (2012) PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res 40(Database issue):D261–D270
Goldstein G, Scheid M, Hammerling U et al (1975) Isolation of a polypeptide that has lymphocyte-differentiating properties and is probably represented universally in living cells. Proc Natl Acad Sci U S A 72(1):11–15
Pickart CM (2001) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Schulman BA, Harper JW (2009) Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nat Rev Mol Cell Biol 10(5):319–331
van Wijk SJ, Timmers HT (2010) The family of ubiquitin-conjugating enzymes (E2s): deciding between life and death of proteins. FASEB J 24(4):981–993
Ye Y, Rape M (2009) Building ubiquitin chains: E2 enzymes at work. Nat Rev Mol Cell Biol 10(11):755–764
Metzger MB, Hristova VA, Weissman AM (2012) HECT and RING finger families of E3 ubiquitin ligases at a glance. J Cell Sci 125(Pt 3):531–537
Breitschopf K, Bengal E, Ziv T et al (1998) A novel site for ubiquitination: the N-terminal residue, and not internal lysines of MyoD, is essential for conjugation and degradation of the protein. EMBO J 17(20):5964–5973
Bloom J, Amador V, Bartolini F et al (2003) Proteasome-mediated degradation of p21 via N-terminal ubiquitinylation. Cell 115(1):71–82
Scaglione KM, Basrur V, Ashraf NS et al (2013) The ubiquitin-conjugating enzyme (E2) Ube2w ubiquitinates the N terminus of substrates. J Biol Chem 288(26):18784–18788
Cadwell K, Coscoy L (2005) Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309(5731):127–130
Wang X, Herr RA, Chua WJ et al (2007) Ubiquitination of serine, threonine, or lysine residues on the cytoplasmic tail can induce ERAD of MHC-I by viral E3 ligase mK3. J Cell Biol 177(4):613–624
Shimizu Y, Okuda-Shimizu Y, Hendershot LM (2010) Ubiquitylation of an ERAD substrate occurs on multiple types of amino acids. Mol Cell 40(6):917–926
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434
Komander D, Clague MJ, Urbe S (2009) Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol 10(8):550–563
Komander D, Rape M (2012) The ubiquitin code. Annu Rev Biochem 81:203–229
Tatham MH, Geoffroy MC, Shen L et al. (2008) RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol 10(5):538–546
Dikic I, Wakatsuki S, Walters KJ (2009) Ubiquitin-binding domains—from structures to functions. Nat Rev Mol Cell Biol 10(10):659–671
Winget JM, Mayor T (2010) The diversity of ubiquitin recognition: hot spots and varied specificity. Mol Cell 38(5):627–635
Husnjak K, Dikic I (2012) Ubiquitin-binding proteins: decoders of ubiquitin-mediated cellular functions. Annu Rev Biochem 81:291–322
Mukhopadhyay D, Riezman H (2007) Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science 315(5809):201–205
Grabbe C, Husnjak K, Dikic I (2011) The spatial and temporal organization of ubiquitin networks. Nat Rev Mol Cell Biol 12(5):295–307
Armirotti A, Damonte G (2010) Achievements and perspectives of top-down proteomics. Proteomics 10(20):3566–3576
Morishima-Kawashima M, Hasegawa M, Takio K et al (1993) Ubiquitin is conjugated with amino-terminally processed tau in paired helical filaments. Neuron 10(6):1151–1160
Nielsen ML, Vermeulen M, Bonaldi T et al (2008) Iodoacetamide-induced artifact mimics ubiquitination in mass spectrometry. Nat Methods 5(6):459–460
Kaiser P, Mayor T (2011) Gold for ubiquitin in Vancouver: first conference on proteomics of protein degradation and ubiquitin pathways held June 6–8, 2010 in Vancouver, University of British Columbia, organized by Lan Huang, Thibault Mayor, and Peipei Ping. Mol Cell Proteomics 10(5):R110.003863
Semple CA, RIKEN GER Group, GSL Members (2003) The comparative proteomics of ubiquitination in mouse. Genome Res 13(6B):1389–1394
Peng J, Schwartz D, Elias JE et al (2003) A proteomics approach to understanding protein ubiquitination. Nat Biotechnol 21(8):921–926
Franco M, Seyfried NT, Brand AH et al (2011) A novel strategy to isolate ubiquitin conjugates reveals wide role for ubiquitination during neural development. Mol Cell Proteomics 10(5):M110.002188
Tagwerker C, Flick K, Cui M et al (2006) A tandem affinity tag for two-step purification under fully denaturing conditions: application in ubiquitin profiling and protein complex identification combined with in vivocross-linking. Mol Cell Proteomics 5(4):737–748
Meierhofer D, Wang X, Huang L et al (2008) Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J Proteome Res 7(10):4566–4576
Fang NN, Ng AH, Measday V et al (2011) Hul5 HECT ubiquitin ligase plays a major role in the ubiquitylation and turnover of cytosolic misfolded proteins. Nat Cell Biol 13(11):1344–1352
Starita LM, Lo RS, Eng JK et al (2012) Sites of ubiquitin attachment in Saccharomyces cerevisiae. Proteomics 12(2):236–240
Danielsen JM, Sylvestersen KB, Bekker-Jensen S et al (2011) Mass spectrometric analysis of lysine ubiquitylation reveals promiscuity at site level. Mol Cell Proteomics 10(3):M110.003590
Lowe ED, Hasan N, Trempe JF et al (2006) Structures of the Dsk2 UBL and UBA domains and their complex. Acta Crystallogr D Biol Crystallogr 62(Pt 2):177–188
Mayor T, Lipford JR, Graumann J et al (2005) Analysis of polyubiquitin conjugates reveals that the Rpn10 substrate receptor contributes to the turnover of multiple proteasome targets. Mol Cell Proteomics 4(6):741–751
Verma R, Oania R, Graumann J et al (2004) Multiubiquitin chain receptors define a layer of substrate selectivity in the ubiquitin-proteasome system. Cell 118(1):99–110
Mayor T, Graumann J, Bryan J et al (2007) Quantitative profiling of ubiquitylated proteins reveals proteasome substrates and the substrate repertoire influenced by the Rpn10 receptor pathway. Mol Cell Proteomics 6(11):1885–1895
Bennett EJ, Shaler TA, Woodman B et al (2007) Global changes to the ubiquitin system in Huntington’s disease. Nature 448(7154):704–708
Hjerpe R, Aillet F, Lopitz-Otsoa F et al (2009) Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep 10(11):1250–1258
Lopitz-Otsoa F, Rodriguez-Suarez E, Aillet F et al (2012) Integrative analysis of the ubiquitin proteome isolated using tandem ubiquitin binding entities (TUBEs). J Proteomics 75(10):2998–3014
Olsen JV, Blagoev B, Gnad F et al (2006) Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127(3):635–648
Villen J, Beausoleil SA, Gerber SA et al (2007) Large-scale phosphorylation analysis of mouse liver. Proc Natl Acad Sci U S A 104(5):1488–1493
Bodenmiller B, Mueller LN, Mueller M et al (2007) Reproducible isolation of distinct, overlapping segments of the phosphoproteome. Nat Methods 4(3):231–237
Xu G, Paige JS, Jaffrey SR (2010) Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat Biotechnol 28(8):868–873
Kim W, Bennett EJ, Huttlin EL et al (2011) Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol Cell 44(2):325–340
Wagner SA, Beli P, Weinert BT et al (2011) A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol Cell Proteomics 10(10):M111.013284
Na CH, Jones DR, Yang Y et al (2012) Synaptic protein ubiquitination in rat brain revealed by antibody-based ubiquitome analysis. J Proteome Res 11(9):4722–4732
Wagner SA, Beli P, Weinert BT et al (2012) Proteomic analyses reveal divergent ubiquitylation site patterns in murine tissues. Mol Cell Proteomics 11(12):1578–1585
Udeshi ND, Svinkina T, Mertins P et al (2013) Refined preparation and use of anti-diglycine remnant (K-epsilon-GG) antibody enables routine quantification of 10,000s of ubiquitination sites in single proteomics experiments. Mol Cell Proteomics 12(3):825–831
Skaug B, Chen ZJ (2010) Emerging role of ISG15 in antiviral immunity. Cell 143(2):187–190
Ng AH, Fang NN, Comyn SA et al (2013) System-wide analysis reveals intrinsically disordered proteins are prone to ubiquitylation after misfolding stress. Mol Cell Proteomics 12(9):2456–2467
Hale JE, Butler JP, Knierman MD et al (2000) Increased sensitivity of tryptic peptide detection by MALDI-TOF mass spectrometry is achieved by conversion of lysine to homoarginine. Anal Biochem 287(1):110–117
Peters EC, Horn DM, Tully DC et al (2001) A novel multifunctional labeling reagent for enhanced protein characterization with mass spectrometry. Rapid Commun Mass Spectrom 15(24):2387–2392
Nakamura H, Linclau B, Curran DP (2001) Fluorous triphasic reactions: transportative deprotection of fluorous silyl ethers with concomitant purification. J Am Chem Soc 123(41):10119–10120
Brittain SM, Ficarro SB, Brock A et al (2005) Enrichment and analysis of peptide subsets using fluorous affinity tags and mass spectrometry. Nat Biotechnol 23(4):463–468
Go EP, Uritboonthai W, Apon JV et al (2007) Selective metabolite and peptide capture/mass detection using fluorous affinity tags. J Proteome Res 6(4):1492–1499
Ying W, Perlman DH, Li L et al (2009) Highly efficient and selective enrichment of peptide subsets combining fluorous chemistry with reversed-phase chromatography. Rapid Commun Mass Spectrom 23(24):4019–4030
Baty JD, Robinson PR (1977) Single and multiple ion recording techniques for the analysis of diphenylhydantoin and its major metabolite in plasma. Biomed.Mass Spectrom 4(1):36–41
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422(6928):198–207
Domon B, Aebersold R (2006) Mass spectrometry and protein analysis. Science 312(5771):212–217
Gallien S, Duriez E, Domon B (2011) Selected reaction monitoring applied to proteomics. J Mass Spectrom 46(3):298–312
Picotti P, Aebersold R (2012) Selected reaction monitoring-based proteomics: workflows, potential, pitfalls and future directions. Nat Methods 9(6):555–566
Gerber SA, Rush J, Stemman O et al (2003) Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc Natl Acad Sci U S A 100(12):6940–6945
Kirkpatrick DS, Gerber SA, Gygi SP (2005) The absolute quantification strategy: a general procedure for the quantification of proteins and post-translational modifications. Methods 35(3):265–273
Kirkpatrick DS, Hathaway NA, Hanna J et al (2006) Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nat Cell Biol 8(7):700–710
Xu P, Duong DM, Seyfried NT et al (2009) Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 137(1):133–145
Mirzaei H, Rogers RS, Grimes B et al (2010) Characterizing the connectivity of poly-ubiquitin chains by selected reaction monitoring mass spectrometry. Mol Biosyst 6(10):2004–2014
Dammer EB, Na CH, Xu P et al (2011) Polyubiquitin linkage profiles in three models of proteolytic stress suggest the etiology of Alzheimer disease. J Biol Chem 286(12):10457–10465
Kaiser SE, Riley BE, Shaler TA et al (2011) Protein standard absolute quantification (PSAQ) method for the measurement of cellular ubiquitin pools. Nat Methods 8(8):691–696
Bedford L, Layfield R, Mayer RJ et al (2011) Diverse polyubiquitin chains accumulate following 26 S proteasomal dysfunction in mammalian neurones. Neurosci Lett 491(1):44–47
Gupta R, Kus B, Fladd C et al (2007) Ubiquitination screen using protein microarrays for comprehensive identification of Rsp5 substrates in yeast. Mol Syst Biol 3:116
Lu JY, Lin YY, Qian J et al (2008) Functional dissection of a HECT ubiquitin E3 ligase. Mol Cell Proteomics 7(1):35–45
Persaud A, Alberts P, Amsen EM et al (2009) Comparison of substrate specificity of the ubiquitin ligases Nedd4 and Nedd4-2 using proteome arrays. Mol Syst Biol 5:333
Merbl Y, Kirschner MW (2009) Large-scale detection of ubiquitination substrates using cell extracts and protein microarrays. Proc Natl Acad Sci U S A 106(8):2543–2548
Loch CM, Strickler JE (2012) A microarray of ubiquitylated proteins for profiling deubiquitylase activity reveals the critical roles of both chain and substrate. Biochim Biophys Acta 1823(11):2069–2078
Merbl Y, Refour P, Patel H et al (2013) Profiling of ubiquitin-like modifications reveals features of mitotic control. Cell 152(5):1160–1172
Guo Z, Song E, Ma S et al (2012) Proteomics strategy to identify substrates of LNX, a PDZ domain-containing E3 ubiquitin ligase. J Proteome Res 11(10):4847–4862
Emanuele MJ, Elia AE, Xu Q et al (2011) Global identification of modular cullin-RING ligase substrates. Cell 147(2):459–474
Tung CW, Ho SY (2008) Computational identification of ubiquitylation sites from protein sequences. BMC Bioinformatics 9:310. doi:10.1186/1471-2105-9-310
Radivojac P, Vacic V, Haynes C et al (2010) Identification, analysis, and prediction of protein ubiquitination sites. Proteins 78(2):365–380
Chen Z, Chen YZ, Wang XF et al (2011) Prediction of ubiquitination sites by using the composition of k-spaced amino acid pairs. PLoS ONE 6(7):e22930
Feng KY, Huang T, Feng KR et al (2013) Using WPNNA classifier in ubiquitination site prediction based on hybrid features. Protein Pept Lett 20(3):318–323
Chen X, Qiu JD, Shi SP et al (2013) Incorporating key position and amino acid residue features to identify general and species-specific ubiquitin conjugation sites. Bioinformatics 29(13):1614–1622
Michael S, Trave G, Ramu C et al (2008) Discovery of candidate KEN-box motifs using cell cycle keyword enrichment combined with native disorder prediction and motif conservation. Bioinformatics 24(4):453–457
Liu Z, Yuan F, Ren J et al (2012) GPS-ARM: computational analysis of the APC/C recognition motif by predicting D-boxes and KEN-boxes. PLoS ONE 7(3):e34370
Arabi A, Ullah K, Branca RM et al (2012) Proteomic screen reveals Fbw7 as a modulator of the NF-kappaB pathway. Nat Commun 3:976
Hagai T, Toth-Petroczy A, Azia A et al (2012) The origins and evolution of ubiquitination sites. Mol Biosyst 8(7):1865–1877
Hagai T, Azia A, Toth-Petroczy A et al (2011) Intrinsic disorder in ubiquitination substrates. J Mol Biol 412(3):319–324
Hershko A, Ciechanover A, Heller H et al (1980) Proposed role of ATP in protein breakdown: conjugation of protein with multiple chains of the polypeptide of ATP-dependent proteolysis. Proc Natl Acad Sci U S A 77(4):1783–1786
Chau V, Tobias JW, Bachmair A et al (1989) A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243(4898):1576–1583
Spence J, Sadis S, Haas AL et al (1995) A ubiquitin mutant with specific defects in DNA repair and multiubiquitination. Mol Cell Biol 15(3):1265–1273
Galan JM, Haguenauer-Tsapis R (1997) Ubiquitin lys63 is involved in ubiquitination of a yeast plasma membrane protein. EMBO J 16(19):5847–5854
Tokunaga F, Sakata S, Saeki Y et al (2009) Involvement of linear polyubiquitylation of NEMO in NF-kappaB activation. Nat Cell Biol 11(2):123–132
Jin L, Williamson A, Banerjee S et al (2008) Mechanism of ubiquitin-chain formation by the human anaphase-promoting complex. Cell 133(4):653–665
Boname JM, Thomas M, Stagg HR et al (2010) Efficient internalization of MHC I requires lysine-11 and lysine-63 mixed linkage polyubiquitin chains. Traffic 11(2):210–220
Goto E, Yamanaka Y, Ishikawa A et al (2010) Contribution of lysine 11-linked ubiquitination to MIR2-mediated major histocompatibility complex class I internalization. J Biol Chem 285(46):35311–35319
Huang H, Jeon MS, Liao L et al (2010) K33-linked polyubiquitination of T cell receptor-zeta regulates proteolysis-independent T cell signaling. Immunity 33(1):60–70
Comyn SA, Chan GT, Mayor T (2013) False start: Cotranslational protein ubiquitination and cytosolic protein quality control. J Proteomics
Schubert U, Anton LC, Gibbs J et al (2000) Rapid degradation of a large fraction of newly synthesized proteins by proteasomes. Nature 404(6779):770–774
Wang F, Durfee LA, Huibregtse JM (2013) A cotranslational ubiquitination pathway for quality control of misfolded proteins. Mol Cell 50(3):368–378
Lecker SH, Goldberg AL, Mitch WE (2006) Protein degradation by the ubiquitin-proteasome pathway in normal and disease states. J Am Soc Nephrol 17(7):1807–1819
Ross CA, Poirier MA (2004) Protein aggregation and neurodegenerative disease. Nat Med (10 Suppl):S10–S17
Alves-Rodrigues A, Gregori L, Figueiredo-Pereira ME (1998) Ubiquitin, cellular inclusions and their role in neurodegeneration. Trends Neurosci 21(12):516–520
Gies E, Wilde I, Winget JM et al (2010) Niclosamide prevents the formation of large ubiquitin-containing aggregates caused by proteasome inhibition. PLoS ONE 5(12):e14410
Wilde IB, Brack M, Winget JM et al (2011) Proteomic characterization of aggregating proteins after the inhibition of the ubiquitin proteasome system. J Proteome Res 10(3):1062–1072
Carlson N, Rogers S, Rechsteiner M (1987) Microinjection of ubiquitin: changes in protein degradation in HeLa cells subjected to heat-shock. J Cell Biol 104(3):547–555
Bennett EJ, Rush J, Gygi SP et al (2010) Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell 143(6):951–965
Busino L, Millman SE, Scotto L et al. (2012) Fbxw7alpha- and GSK3-mediated degradation of p100 is a pro-survival mechanism in multiple myeloma. Nat Cell Biol 14(4):375–385
Guardavaccaro D, Frescas D, Dorrello NV et al (2008) Control of chromosome stability by the beta-TrCP-REST-Mad2 axis. Nature 452(7185):365–369
Peschiaroli A, Dorrello NV, Guardavaccaro D et al (2006) SCFbetaTrCP-mediated degradation of claspin regulates recovery from the DNA replication checkpoint response. Mol Cell 23(3):319–329
Dorrello NV, Peschiaroli A, Guardavaccaro D et al (2006) S6K1- and betaTRCP-mediated degradation of PDCD4 promotes protein translation and cell growth. Science 314(5798):467–471
Liu B, Zheng Y, Wang TD et al (2012) Proteomic identification of common SCF ubiquitin ligase FBXO6-interacting glycoproteins in three kinds of cells. J Proteome Res 11(3):1773–1781
Teixeira FR, Yokoo S, Gartner CA et al (2010) Identification of FBXO25-interacting proteins using an integrated proteomics approach. Proteomics 10(15):2746–2757
Yumimoto K, Matsumoto M, Oyamada K et al. (2012) Comprehensive identification of substrates for F-box proteins by differential proteomics analysis. J Proteome Res 11(6):3175–3185
Zhuang M, Guan S, Wang H et al (2013) Substrates of IAP ubiquitin ligases identified with a designed orthogonal E3 ligase, the NEDDylator. Mol Cell 49(2):273–282
Sarraf SA, Raman M, Guarani-Pereira V et al (2013) Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496(7445):372–376
Narendra D, Tanaka A, Suen DF et al. (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183(5):795–803
Luise C, Capra M, Donzelli M et al (2011) An atlas of altered expression of deubiquitinating enzymes in human cancer. PLoS ONE 6(1):e15891
McDonell LM, Mirzaa GM, Alcantara D et al. (2013) Mutations in STAMBP, encoding a deubiquitinating enzyme, cause microcephaly-capillary malformation syndrome. Nat Genet 45(5):556–562
Cilenti L, Balakrishnan MP, Wang XL et al. (2011) Regulation of Abro1/KIAA0157 during myocardial infarction and cell death reveals a novel cardioprotective mechanism for Lys63-specific deubiquitination. J Mol Cell Cardiol 50(4):652–661
Fraile JM, Quesada V, Rodriguez D et al (2012) Deubiquitinases in cancer: new functions and therapeutic options. Oncogene 31(19):2373–2388
Sowa ME, Bennett EJ, Gygi SP et al (2009) Defining the human deubiquitinating enzyme interaction landscape. Cell 138(2):389–403
Kouranti I, McLean JR, Feoktistova A et al (2010) A global census of fission yeast deubiquitinating enzyme localization and interaction networks reveals distinct compartmentalization profiles and overlapping functions in endocytosis and polarity. PLoS Biol 8(9). doi:10.1371/journal.pbio.1000471
Richardson LA, Reed BJ, Charette JM et al (2012) A conserved deubiquitinating enzyme controls cell growth by regulating RNA polymerase I stability. Cell Rep 2(2):372–385
Poulsen JW, Madsen CT, Young C et al (2012) Comprehensive profiling of proteome changes upon sequential deletion of deubiquitylating enzymes. J Proteomics 75(13):3886–3897
Borodovsky A, Ovaa H, Kolli N et al (2002) Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chem Biol 9(10):1149–1159
Altun M, Kramer HB, Willems LI et al (2011) Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chem Biol 18(11):1401–1412
Lennartsson A, Ekwall K (2009) Histone modification patterns and epigenetic codes. Biochim Biophys Acta 1790(9):863–868
Mertins P, Qiao JW, Patel J et al (2013) Integrated proteomic analysis of post-translational modifications by serial enrichment. Nat Methods 10(7):634–637
Swaney DL, Beltrao P, Starita L et al (2013) Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation. Nat Methods 10(7):676–682
Acknowledgments
We thank Sophie Comyn and Gerard Chan, and all other members of the Mayor lab for discussion and suggestions. This work is supported by grants from the Natural Sciences and Engineering Research Council of Canada (NSERC) and the Canadian Institutes of Health Research (CIHR). TM is supported by New Investigator Career Awards from CIHR and the Michael Smith Foundation for Health Research.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Fang, N., Albu, R., Mayor, T. (2014). Systems-Wide Analysis of Protein Ubiquitylation: We Finally Have the Tiger by the Tail. In: Houry, W. (eds) The Molecular Chaperones Interaction Networks in Protein Folding and Degradation. Interactomics and Systems Biology, vol 1. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1130-1_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1130-1_14
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1129-5
Online ISBN: 978-1-4939-1130-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)