Abstract
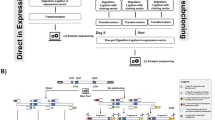
Although proteins can be artificially improved by random insertion and deletion mutagenesis methods, these procedures are technically difficult. Here we describe a simple method called random insertional–deletional strand exchange mutagenesis (RAISE). This method is based on gene shuffling and can be used to introduce a wide variety of insertions, deletions, and substitutions. RAISE involves three steps: DNA fragmentation, attachment of a random short sequence, and reconstruction. This yields unique mutants and can be a powerful technique for protein engineering.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Antikainen NM, Martin SF (2005) Altering protein specificity: techniques and applications. Bioorg Med Chem 13:2701–2716
Otten LG, Quax WJ (2005) Directed evolution: selecting today’s biocatalysts. Biomol Eng 22:1–9
Robertson DE, Steer BA (2004) Recent progress in biocatalyst discovery and optimization. Curr Opin Chem Biol 8:141–149
Powell KA, Ramer SW, del Cardayre SB et al (2001) Directed evolution and biocatalysis. Angew Chem Int Ed Engl 40:3948–3959
Brakmann S (2001) Discovery of superior enzymes by directed molecular evolution. Chembiochem 2:865–871
Farinas ET, Bulter T, Arnold FH (2001) Directed enzyme evolution. Curr Opin Biotechnol 12:545–551
Goldsmith M, Tawfik DS (2013) Enzyme engineering by targeted libraries. Methods Enzymol 523:257–283
Goldsmith M, Tawfik DS (2012) Directed enzyme evolution: beyond the low-hanging fruit. Curr Opin Struct Biol 22:406–412
Wang M, Si T, Zhao H (2012) Biocatalyst development by directed evolution. Bioresour Technol 115:117–125
Neylon C (2004) Chemical and biochemical strategies for the randomization of protein encoding DNA sequences: library construction methods for directed evolution. Nucleic Acids Res 32:1448–1459
Leung DW, Chen E, Goeddel DV (1989) A method for random mutagenesis of a defined DNA segment using a modified polymerase chain reaction. Technique 1:11–15
Shortle D, Sondek J (1995) The emerging role of insertions and deletions in protein engineering. Curr Opin Biotechnol 6:387–393
Jones DD (2005) Triplet nucleotide removal at random positions in a target gene: the tolerance of TEM-1 b-lactamase to an amino acid deletion. Nucleic Acids Res 33:e80
Baldwin AJ, Busse K, Simm AM et al (2008) Expanded molecular diversity generation during directed evolution by trinucleotide exchange (TriNEx). Nucleic Acids Res 36:e77
Murakami H, Hohsaka T, Sisido M (2002) Random insertion and deletion of arbitrary number of bases for codon-based random mutation of DNAs. Nat Biotechnol 20:76–81
Pikkemaat MG, Janssen DB (2002) Generating segmental mutations in haloalkane dehalogenase: a novel part in the directed evolution toolbox. Nucleic Acids Res 30:e35
Hayes F, Hallet B (2000) Pentapeptide scanning mutagenesis: encouraging old proteins to execute unusual tricks. Trends Microbiol 8:571–577
Kim D, Rhee Y, Rhodes D et al (2005) Directed evolution and identification of control regions of ColE1 plasmid replication origins using only nucleotide deletions. J Mol Biol 351:763–775
Fujii R, Kitaoka M, Hayashi K (2006) RAISE: a simple and novel method of generating random insertion and deletion mutations. Nucleic Acids Res 34:e30
Stemmer WPC (1994) Rapid evolution of a protein in vitro by DNA shuffling. Nature 370:389–391
Lewin B (1994) Gene. Oxford University Press, Oxford
Lewis SM (1994) The mechanism of V(D)J joining: lessons from molecular, immunological, and comparative analyses. In: Dixon FJ (ed) Advances in immunology, vol 56. Academic, San Diego, pp 27–150
Komori T, Okada A, Stewart V et al (1993) Lack of N regions in antigen receptor variable region genes of TdT-deficient lymphocytes. Science 261:1171–1175
Pascarella S, Argos P (1992) Analysis of insertions/deletions in protein structures. J Mol Biol 224:461–471
Lorimer IAJ, Pastan I (1995) Random recombination of antibody single chain Fv sequences after fragmentation with DNaseI in the presence of Mn2+. Nucleic Acids Res 23:3067–3068
Aharoni A, Griffiths AD, Tawfik DS (2005) High-throughput screens and selections of enzyme-encoding genes. Curr Opin Chem Biol 9:210–216
Goddard JP, Reymond JL (2004) Recent advances in enzyme assays. Trends Biotechnol 22:363–370
Goddard JP, Reymond JL (2004) Enzyme assays for high-throughput screening. Curr Opin Biotechnol 15:314–322
Schmidt M, Bornscheuer UT (2005) High-throughput assays for lipases and esterases. Biomol Eng 22:51–56
Acknowledgments
This study was supported in part by a grant from the Program for Promotion of Basic Research Activities for Innovative Biosciences (PROBRAIN).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Fujii, R., Kitaoka, M., Hayashi, K. (2014). Random Insertional–Deletional Strand Exchange Mutagenesis (RAISE): A Simple Method for Generating Random Insertion and Deletion Mutations. In: Gillam, E., Copp, J., Ackerley, D. (eds) Directed Evolution Library Creation. Methods in Molecular Biology, vol 1179. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1053-3_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1053-3_10
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1052-6
Online ISBN: 978-1-4939-1053-3
eBook Packages: Springer Protocols