Abstract
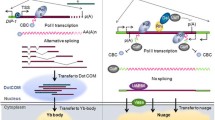
Endogenous siRNAs (endo-siRNAs) are well documented and characterized in C. elegans and Drosophila. Endo-siRNAs can also be found in vertebrates; however, their biology is much less clear. They are thought to be produced by Dicer and to contribute to transposon silencing. Because of their generally low abundance and their similarity with miRNAs and products of physiological RNA turn-over, endo-siRNAs are difficult to investigate. Here, we report a system, oocytes from Xenopus laevis, that allows for the generation and analysis of endo-siRNAs from double-stranded RNA precursors.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Okamura K, Lai EC (2008) Endogenous small interfering RNAs in animals. Nat Rev Mol Cell Biol 9:673–678
Chapman EJ, Carrington JC (2007) Specialization and evolution of endogenous small RNA pathways. Nat Rev Genet 8:884–896
Wang Q, Carmichael GG (2004) Effects of length and location on the cellular response to double-stranded RNA. Microbiol Mol Biol Rev 68:432–452
Tam OH, Aravin AA, Stein P, Girard A, Murchison EP, Cheloufi S, Hodges E, Anger M, Sachidanandam R, Schultz RM, Hannon GJ (2008) Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453:534–538
Carlile M, Swan D, Jackson K, Preston-Fayers K, Ballester B, Flicek P, Werner A (2009) Strand selective generation of endo-siRNAs from the Na/phosphate transporter gene Slc34a1 in murine tissues. Nucleic Acids Res 37:2274–2282
Watanabe T, Totoki Y, Toyoda A, Kaneda M, Kuramochi-Miyagawa S, Obata Y, Chiba H, Kohara Y, Kono T, Nakano T, Surani MA, Sakaki Y, Sasaki H (2008) Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453:539–543
Soumillon M, Necsulea A, Weier M, Brawand D, Zhang X, Gu H, Barthes P, Kokkinaki M, Nef S, Gnirke A, Dym M, de Massy B, Mikkelsen TS, Kaessmann H (2013) Cellular source and mechanisms of high transcriptome complexity in the Mammalian testis. Cell Rep 3:2179–2190
Carlile M, Nalbant P, Preston-Fayers K, McHaffie GS, Werner A (2008) Processing of naturally occurring sense/antisense transcripts of the vertebrate Slc34a gene into short RNAs. Physiol Genomics 34:95–100
Saccomanno L, Bass BL (1994) The cytoplasm of Xenopus oocytes contains a factor that protects double-stranded RNA from adenosine-to-inosine modification. Mol Cell Biol 14:5425–5432
Markovich D (2008) Expression cloning and radiotracer uptakes in Xenopus laevis oocytes. Nat Protoc 3:1975–1980
Tammaro P, Shimomura K, Proks P (2008) Xenopus oocytes as a heterologous expression system for studying ion channels with the patch-clamp technique. Methods Mol Biol 491:127–139
Acknowledgment
This work has been supported by the Dunhill Medical Trust and Newcastle University.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Alnumeir, S., Werner, A. (2014). Generation of Endo-siRNAs in Xenopus laevis Oocytes. In: Werner, A. (eds) Animal Endo-SiRNAs. Methods in Molecular Biology, vol 1173. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-0931-5_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-0931-5_3
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-0930-8
Online ISBN: 978-1-4939-0931-5
eBook Packages: Springer Protocols