Abstract
Human immunodeficiency virus 1 (HIV-1) currently infects over 34 million people across the world and remains one of the greatest global public health burdens despite the development of highly active antiretroviral therapy. An effective prophylactic vaccine is therefore greatly needed but remains elusive due to HIV-1’s ability to evade the immune system. The isolation of potent, broadly neutralizing antibodies (bnAb) from infected individuals suggests a vaccine might in principle be possible. Structural characterization of such antibodies in complex with component pieces of the heavily glycosylated envelope glycoprotein gp160 (Env) is providing information critical for the design of more effective immunogens. Recently, a number of glycan-dependent bnAbs have been isolated that are very potent. Four of these bnAbs have been structurally characterized in complex with their glycosylated antigens. In this chapter, we describe the different binding modes of these glycan-dependent antibodies and how they define novel sites of vulnerability that can be used for design of a new generation of glycosylated immunogens.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 A Pressing Need for a Protective Vaccine
According to the latest estimates by the World Health Organization, human immunodeficiency virus 1 (HIV-1) infects 2.4 million people and kills 1.8 million people globally every year (WHO 2011). Without a protective vaccine, HIV-1 will long remain a critical global public health burden. Vaccines can be designed primarily to elicit antibody responses (e.g., subunit vaccines), cytotoxic T-cell responses (e.g., DNA vaccines), or both (chemically inactivated or live-attenuated viruses). For most vaccines against viral pathogens, neutralizing antibodies are the primary correlate of immune protection (Plotkin 2010) and, in simian HIV (SHIV) models, passive transfer of broadly neutralizing antibodies (bnAb) confers protection against viral challenge (Mascola et al. 1999, 2000; Baba et al. 2000). These results suggest it may be possible to develop a protective Env vaccine that elicits bnAb responses.
5.2 HIV-1 Envelope Glycoprotein and Sites of Vulnerability
HIV-1 has only one surface-exposed viral protein, the homotrimeric envelope glycoprotein gp160 (Env), which mediates cell entry, and therefore represents the only target for the humoral immune response. Unfortunately for vaccine development, Env is the most genetically diverse gene within an already highly diverse HIV-1 genome, thereby representing an effective and efficient evolutionary mechanism to escape from the adaptive immune responses. Nevertheless, a structural and functional understanding of the Env protein and its recognition by bnAbs has uncovered some exciting new opportunities for vaccine design.
During viral maturation, Env is activated by furin cleavage (Hallenberger et al. 1992) into a receptor-binding domain, gp120, and a fusion-mediating, transmembrane domain, gp41, which is a type-1 viral fusion protein (Colman and Lawrence 2003). The activated Env components remain non-covalently associated in a metastable trimer of heterodimers poised to undergo the extensive conformational rearrangements required for membrane fusion (Freed 2001). Assembly of a functional Env is not efficient: dimers, malformed trimers, and higher-order aggregates that cannot mediate viral fusion are common (Moore et al. 2006; Crooks et al. 2011). Furthermore, once assembled, Env is prone to shedding its gp120 components, resulting in nonfunctional Env (Schneider et al. 1986), thus diverting the immune response away from functional trimeric Env and making structural characterization of a stable trimer much more difficult.
Despite these challenges, crystal structures have been determined for peptide fragments, gp120 core monomers, and scaffolded gp120 and gp41 loops from a variety of viral strains, bound to antibodies and/or receptors (some examples are Kwong et al. 1998, 2000a; Calarese et al. 2003; Cardoso et al. 2005; Huang et al. 2005, 2012; Zhou et al. 2007, 2010; McLellan et al. 2011; Pejchal et al. 2011; Kong et al. 2013). These high-resolution structures can then be used to fit into the electron microscopy (EM) reconstructions of Env and Env complexes to give at least a low resolution picture of the Env architecture and interaction with antibodies or receptor (Fig. 5.1) (Zhu et al. 2006, 2008; Liu et al. 2008; Harris et al. 2011; McLellan et al. 2011; Pejchal et al. 2011; Tran et al. 2012; Kong et al. 2013; Julien et al. 2013a). The gp120 glycoprotein has an overall globular fold with five highly variable loops (V1–V5) extending from its core (Kwong et al. 1998). The core can be divided into two structural regions: an inner domain consisting of a bundle formed by two helices, 2 strands, and a 5-stranded β-sandwich, and an outer domain consisting of stacked double β-barrels consisting of six and seven strands. The axes of the outer barrel and inner bundle are approximately parallel. The outer domain faces away from the trimer, and is decorated by most of the 16–32 Asparagine (N)-linked glycans found on gp120. The N-linked glycans are coded by sequons, Asn-x-Thr or Asn-x-Ser, where x can be any amino acid except proline (reviewed in Schwarz and Aebi 2011). While no current high resolution gp120 structures include the V1–V2 region, crystal structures of two scaffolded V1–V2 loop regions show they adopt a surprisingly compact, β-sheet core with smaller than anticipated actual V1 and V2 loops (McLellan et al. 2011). The V3 loop, on the other hand, appears to be quite flexible along its length in several different crystal structures and adopts an extended, β-hairpin structure (Stanfield et al. 2004, 2006; Huang et al. 2005). During viral entry, the CD4 receptor binds to a conserved region on the gp120 core consisting primarily of the outer domain as well as a portion of the inner domain followed by binding to a coreceptor (typically chemokine receptors CCR5 or CXCR4). Together, these binding events are thought to initiate conformational rearrangements whereby gp41 mediates fusion of the host and viral membranes (reviewed in Knipe and Howley 2007).
Combining structural information from crystallography and microscopy. Docking of the crystal structure of Fab PGT 128 in complex with gp120 outer domain into a negative stain electron micrograph reconstruction of Fab PGT 128 bound to gp120 trimer. The top view is looking down on the threefold axis of the trimer, while the bottom view is from the side. The EM envelope is shown in blue, with the Fab structure shown in blue and white ribbons, and the outer domain in red. Glycans are shown in yellow. This figure was previously published in Pejchal et al. (2011)
Epitopes on Env for bnAbs tend to cluster around particular regions or “sites of vulnerability” important for the viral functions described above. The best-characterized site of vulnerability is the CD4 receptor-binding site, which is targeted by some of the most potent and broadly neutralizing antibodies isolated to date (Wu et al. 2010). Due to its important viral function, the CD4-binding site must be exposed and relatively conserved in order to engage the host cell receptor for viral entry. However, this region is recessed within the trimer and encircled by N-linked glycans; only antibodies that approach the region from the “correct” angle of approach can bind and neutralize, a constraint that presents a formidable challenge for vaccine design. A second site of vulnerability is the membrane proximal region (MPER) on gp41 where antibody binding may interfere with viral membrane fusion. Unlike the CD4-binding site, this epitope region contains continuous linear sequences of amino acids, as shown by several antibody-MPER peptide complex structures (Ofek et al. 2004; Cardoso et al. 2005, 2007; Julien et al. 2008; Montero et al. 2008; Huang et al. 2012); however, whether additional parts of gp120 or gp41 are included in these overlapping epitopes is unclear. The MPER is also somewhat recessed, in this case between the membrane and the rest of gp41 and gp120, and thus its proximity to the membrane may also raise issues of membrane recognition and autoimmunity (Haynes et al. 2005; Liao et al. 2011; Chang et al. 2012).
Originally thought to be immunologically silent, the large surfaces on Env covered with N-linked glycans have recently been shown to contain epitopes that are targeted by bnAbs from human patients and non-human primates (Calarese et al. 2003; Walker et al. 2009, 2011a, b; Gray et al. 2011; McLellan et al. 2011; Mouquet et al. 2011; Pejchal et al. 2011). A number of these glycan-recognizing bnAbs have been isolated and are just as potent and broad as the best bnAbs against glycan-free sites. Recently, several of these antibodies have been structurally characterized by X-ray crystallography and EM (Calarese et al. 2003; McLellan et al. 2011; Pejchal et al. 2011; Kong et al. 2013; Julien et al. 2013a, b).
5.3 Anti-glycan Antibodies
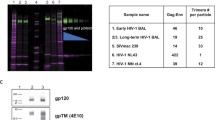
In 1998, the Katinger group isolated three of the four most potent and broadly neutralizing antibodies known at that time, human bnAbs 2F5, 4E10 and 2G12 (Kunert et al. 1998). 2G12 was somewhat of a mystery, as mutations designed to remove N-linked glycans on recombinant gp120 abolished 2G12 binding but, as the glycans were self, they would not normally have been considered immunogenic. In 2003, the crystal structure of 2G12 revealed that it had a remarkable tertiary structure, where the variable heavy chain (VH) domains of the two Fab fragments in a single IgG were domain swapped, and that it did, indeed, bind carbohydrate, as vividly illustrated in complexes of 2G12 with several different high mannose sugars, including GlcNAc2Man9 (Calarese et al. 2003). These findings launched a multitude of studies into antibody recognition of carbohydrate, and many creative attempts to create a carbohydrate-based vaccine (for review of this subject see Kong et al. 2012 and Chap. 6). Then, in 2009, a concerted effort to discover new broadly neutralizing antibodies in the serum of elite neutralizing patients led to the discovery of many more glycan-dependent antibodies (Walker et al. 2009, 2011a; Corti et al. 2010; Wu et al. 2010; Bonsignori et al. 2011; Mouquet et al. 2011). In these panels, carbohydrate-binding antibodies were represented in members of the PGT 121, PGT 128, PGT 135, PGT 141 and PG9/PG16 families. The PGT 121, 128 and 135 families of antibodies recognize an epitope involving a glycan at N332 at the base of the V3 loop, while the PG9/PG16 and PGT 141 families recognize an epitope involving the N160 glycan in the V1/V2 region. Remarkably, all of these new antibodies utilize unusual structural features to target their glycan epitopes, including post-translational modifications, long complementarity determining region (CDR) H3 loops, and novel insertions and deletions in their CDRs. These antibodies are all highly potent and broadly neutralizing and, when compared in the same neutralization assay against a cross-clade panel of 162 pseudoviruses (Walker et al. 2011a), they have extremely low median IC50 values (in μg/mL) of 0.02 for PGT 128, 0.17 for PGT 135, and 0.23 for PG9 compared to 2.38 for 2G12, while retaining considerable breadth at an IC50 of less than 50 μg/mL (33 % neutralization of the panel by PGT 135 and 32 % by 2G12, 72 % by PGT 128 and 77 % by PG9).
5.3.1 2G12
The 2G12 IgG has a unique domain-swapped structure, where the VH domain from each of the two Fab fragments in the IgG swap with the neighboring Fab, resulting in an antibody with two side-by-side and tightly intertwined Fab fragments (Calarese et al. 2003) (Fig. 5.2a). The overall shape of the resultant IgG is linear, rather than the common Y-shape seen for typical IgG molecules. The Fab fragments pack tightly against each other via a novel VH–V′H interface. Several unusual mutations in 2G12 are critical for promoting the domain swap, including a Pro at position H113 in the elbow region, an Ala at position H14, Glu at H75, as well as an Ile at H19 in the VH–V′H interface (Huber et al. 2010). While wild-type 2G12 is heavily somatically mutated (38/16 somatic mutations at the amino acid level in the heavy/light chains), introducing as few as 5–7 of the wild-type mutations into germline 2G12 heavy chain is sufficient to induce dimerization of the Fabs (Huber et al. 2010). Structures of 2G12 in complex with high-mannose glycans show that the 2G12 antigen-binding site interacts with the terminal Manα1,2Man residues from the D1 arm of GlcNAc2Man9 (Calarese et al. 2003, 2005). In the crystal structure with GlcNAc2Man9, the D2 arm from a crystallographic symmetry-related complex also binds into a pocket formed at the VH–VH′ interface, suggesting that 2G12 may interact with anywhere from 2 to 4 high mannose sugars on the viral Env. Several studies (Trkola et al. 1996; Sanders et al. 2002; Scanlan et al. 2002) have implicated sugars at positions N262, N295, N332, N339, N386, N392, and N448 as being important for 2G12 binding. Thus, 2G12 has capitalized on its unusual domain-swapping to create a multivalent binding site for multiple glycan moieties with no apparent protein–protein interactions. The interactions in the two primary combining sites bury a total of 804 Å2 on the dimeric Fab surface and 791 Å2 of carbohydrate, with 77 % of the Fab interactions coming from the heavy chain, while the secondary binding sites are capable of burying another 700 Å2 of Fab surface area (Table 5.1). A single mutation (Ile to Arg at H19) is sufficient to abrogate the domain exchange in recombinantly produced 2G12, and monomeric 2G12 Fab can bind free Manα1,2Man (Doores et al. 2010a). However, the monomeric Fab does not bind the high mannose cluster on the Env trimer to any detectable level, perhaps because the Fab arms of the conventional IgG cannot be brought close enough together to span the necessary glycans.
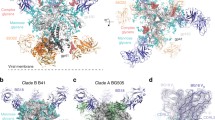
Broadly neutralizing anti-HIV antibodies that recognize carbohydrate epitopes. The four Fabs, 2G12, PG9, PGT 135 and PGT 128, are all shown with their light chains in cyan and heavy chains in dark blue (and dark green for 2G12). The H3 CDR loop is colored red, and gp120 constructs are colored light green. The glycans are shown as ball-and-stick models with yellow and red carbon and oxygen atoms, respectively. (a) 2G12 uses a domain swap of its variable heavy chain domains to create an interlocked dimer of Fabs. This domain swap results in four potential binding sites for carbohydrates, two in the normal antigen-binding sites, and two more at the VH–VH′ interfaces. (b) PG9 binds to a scaffolded V1/V2 construct with its very large hammerhead CDR H3 loop, interacting with two glycans attached to Asn160 and Asn156 in the CAP45 strain. In a related structure with the ZM109 strain, a different but spatially close glycan (at position 173) binds in place of that at 156. (c) PGT 135 binding to the gp120 core, primarily contacting glycans at N332 and N392. (d) PGT 128 is shown binding to a gp120 outer domain construct. The glycans contacted are from positions N332 and N301
2G12 has performed well as regards to both safety and efficacy in early phase clinical trials in combination with 2F5 and 4E10, when all are produced in CHO cells and used as passive immunotherapy during strategic highly active antiretroviral therapy (HAART) interruption periods in patients (Joos et al. 2006; Manrique et al. 2007; Mehandru et al. 2007; Vcelar et al. 2007; Huber et al. 2008). The high cost of the IgG produced in CHO cells has led to testing of more cost-effective ways to mass produce 2G12 and other therapeutic antibodies, and 2G12 produced in plants and formulated as a vaginal microbicide have recently been tested in clinical trials for safety (Fox 2011).
5.3.2 PGT 128
PGT 128 is a member of the PGT family of antibodies recently isolated through the efforts of the International AIDS Vaccine Initiative Protocol G, where a group of elite neutralizers was screened for broadly neutralizing sera (Walker et al. 2009, 2011a). Crystal structures of PGT 128 have been determined to high resolution in complex with GlcNAc2Man9, and to medium resolution in complex with a glycosylated gp120 outer domain construct (Figs. 5.2d and 5.3) (Pejchal et al. 2011). In the complex with free GlcNAc2Man9, the Fab binds to one glycan, interacting with the D1 and D3 arms, while in a complex with gp120 outer domain, the Fab binds to two glycans, and also interacts with part of the gp120 V3 base. The primary glycan-binding site is occupied by the N332 glycan, while a secondary binding site is filled by the N301 glycan, with ordered density present for the GlcNAc2Man5 component of the glycan (Fig. 5.3). The tip of the long H3 CDR (21 residues, between and including H93 to H102) penetrates through the glycan shield to contact V3 residues 323–325 via a β-sheet type interaction. The PGT 128 Fab has other unusual features in its CDR loops, including an unusually long (six amino acids) insertion in H2, and a short L1 (deletion of residues L28 and L29), as well as a disulfide bond linking CDRs H1 and H2. The interaction buries a considerable 1,076 Å2 of surface on the Fab (86 % from the heavy chain) and 1,053 Å2 of surface on the gp120 (Table 5.1).
Fab PGT 128 bound to a gp120 core construct. The heavy chain (dark blue) and light chain (cyan) of PGT 128 Fab (Pejchal et al. 2011) are shown bound to gp120 outer domain (light green). PGT 128 makes extensive contacts with glycans at N301 (dark green) and N332 (yellow) shown as ball-and-stick models surrounded by their electron density. The terminal arms of the glycans are labeled (D1, D2 or D3). In other isolates, this Fab may alternately contact the glycan at N295 (pink, there was no ordered density for this glycan in the PGT 128 crystal structure). The distances between the glycans measured from C1 of the first GlcNAc of each glycan (or ND1 of Asn 295) are shown in a schematic on the right. Also highlighted are the backbone contacts between the CDR H3 loop of PGT 128 (red) and a strand of the V3 loop on gp120, with dashes indicating approximate positions of hydrogen bonds
5.3.3 PG9/PG16
PG9 and PG16 are structurally very similar antibodies derived from the same germline gene family and recognize glycans in the V1/V2 region of gp120 (Pancera et al. 2010; Pejchal et al. 2010). Thus far, all crystal structures of gp120 core or outer domain constructs have included only the base of V1/V2. However, Kwong and colleagues (McLellan et al. 2011) managed to co-crystallize PG9 using a scaffolded V1/V2, in which a V1/V2 peptide had its base constrained in a β-hairpin conformation by a scaffold protein (Figs. 5.2b and 5.4). Structures have been determined for PG9 in complex with V1/V2 scaffolds from two different HIV-1 strains, and both show that PG9 interacts with two glycans from V1/V2, but one of these glycans can be attached to different, closely spaced amino acids depending on the glycan positions of the isolates in question. The scaffolded V1/V2 domain folds into a four-stranded β-sheet Greek key structure. In the scaffolded CAP45 V1/V2, the antibody binds to glycans at N160 and N156, while in the scaffolded ZM109 V1/V2 the contacted glycans are N160 and N173. Glycans at N156 and N173 are in close spatial proximity so that they can substitute for each other in this binding interaction.
PG9 bound to a scaffolded V1/V2 domain. The heavy chain (dark blue) and light chain (cyan) of PG9 is shown bound to the V1/V2 domain of CAP 45 gp120 (light green) (McLellan et al. 2011). Contacts are made to glycans at N160 and N156, which are colored yellow and green, and shown as ball-and-stick models surrounded by their electron density. The terminal arms of the glycans that interact with the Fab are labeled (D1, D2 or D3). They are spaced about 14 Å apart measured from C1 of the first GlcNAc, as shown in the schematic above. Sulfated tyrosines from the CDR H3 loop residues H100G and H100H are shown. Extensive backbone contacts between the CDR H3 loop and V1/V2 domain are also shown with dashes indicating approximate positions of hydrogen bonds
Using a strategy similar to that of PGT 128, the PG9 bisects space between the two interacting glycans with its CDRs, capitalizing on an extremely long, hammerhead-shaped CDR H3 (Fig. 5.4). As a result, the heavy chain dominates the interaction contributing about 80 % of the buried surface (Table 5.1). The CDR H3 is unusual not only in its length (30 residues between and including H93 and H102), but that it contains sulfated tyrosine residues that are important for interaction with a cationic site on the V1/V2 of Env. A negative stain EM reconstruction at 18 Å resolution for PG9 bound to a trimeric Env construct (Julien et al. 2013b) shows only a single PG9 Fab can bind at the apex of the trimer. Modeling of high-resolution crystal structure information into the EM reconstruction suggests that PG9 binds slightly off-center to two of the three gp120 subunits in the trimer. These results also suggest that, in addition to the glycans at N156 and N160 contacted by PG9 in the crystal structure with a scaffolded V1/V2 domain, PG9 may make secondary interactions with a glycan at N160 on a neighboring gp120 subunit. PGT 145 is also thought to recognize the V1/V2 region, and has a very long (33-residue) extended β-hairpin CDR H3 with sulfated tyrosines (McLellan et al. 2011).
5.3.4 PGT 135
PGT 135 is another potent and broadly neutralizing antibody that includes glycans as a major part of its epitope on gp120. The structure of PGT 135 has recently been determined as a multicomponent complex that included the gp120 core (Kong et al. 2013) (Figs. 5.2c and 5.5), as well as antibody 17b and the D1–D2 domains of CD4 that were essential for crystallization. PGT 135 interacts with several glycans, primarily with those at N332 and N392, and with the protein surface between the glycans. Additional interactions with glycans at N295 and N386 are important for binding and neutralization in an isolate-dependent manner. Not to be outdone by the other anti-glycan antibodies, PGT 135 also has unusual features, namely a rare five amino-acid insertion in CDR H1 that enables that CDR to reach through the glycan canopy to encounter the gp120 protein surface near N386. PGT 135 is also unusual in that, as shown by EM, it can adopt multiple degenerate angles of approach to bind what are most likely glycosylation variants (glycoforms) of gp120 (Fig. 5.6). Unlike the other anti-glycan antibodies, PGT 135 uses its light chain extensively, contributing 485 Å2 out of a total of 1,355 Å2 of the surface area buried on the Fab that equates to more than double the surface area buried by PG9, PGT 128, and 2G12 light chains (Table 5.1). Similar to PGT 128 and PG9, PGT 135 uses an extended CDR H3 (20 residues between and including H93 and H102) to nestle into and breach the glycan shield.
PGT 135 bound to gp120 core. The heavy chain (dark blue) and light chain (cyan) of PGT 135 Fab (Kong et al. 2013) is shown interacting with gp120 (light green). Most of the interaction is mediated by the extended CDR H3 (red) and the CDR H1 (blue) loops. This Fab interacts mainly with glycans at N332 (yellow) and N392 (green), but also contacts the glycan at N386 (orange). These glycans are shown as ball-and-stick models surrounded by their electron density, and the terminal arms of the glycans that interact with the Fab are labeled (D1, D2 or D3). The spacing of these glycans, including the non-contacting but critical glycan at N295 (pink), are measured from C1 of the first GlcNAc and shown in the schematic on the right
Flexibility in PGT 135 recognition of gp140 trimer. An overlay of the 2D class averages for PGT 135 Fab-BG505 SOSIP.664 gp140 trimer (blue outline) and the first eigenvector (orange) of the 2D principal component analysis of the PGT 135-trimer images. The positions of glycans N386 and N332 are indicated with red circles. This analysis of negative stain EM reconstructions shows that PGT 135 can bind to trimer with some flexibility. This variance in the binding mode for PGT 135 may be due to a distribution of different approach angles, either because of slight shifts between the interacting glycans or interaction with different glycoforms. This figure was previously published in Kong et al. (2013)
5.4 Overlapping Epitopes of Glycan-Binding bnAbs
A major discovery from these structural and functional studies is that glycan-binding bnAbs have epitopes that cluster and center around two sites: (1) a highly solvent-exposed surface between the base of the V3 loop and the base of the V4 loop centered around glycan N332; and (2) a portion of the V2 loop containing glycans N160 and N156 (Fig. 5.7). Structural studies of these bnAbs have pinpointed the chemical composition and conformational features of N-linked glycans that are recognized. Understanding why these particular glycans are targeted by broadly neutralizing responses compared to all of the other gp120 glycans and the functional consequences is critical to designing immunogens aimed to re-elicit such glycan-dependent bnAbs.
Schematic drawing of the gp120 region containing the highly conserved glycan sites involved in broadly neutralizing antibody recognition. The topology and connectivity of the underlying protein surface is depicted by arrows (β-strands) and cylinders (α-helices) as similarly presented in a previous publication (Kong et al. 2013). Regions on the protein that are contacted by PGT 128 and PGT 135 are colored red. The N-linked glycosylation sites are shown as red circles with their amino acid position and sequence conservation labeled. Sequence conservation was calculated from 3054 aligned HIV-1 Env sequences from the HIV Database from the Los Alamos National Laboratories (http://www.hiv.lanl.gov/). Note that while N334 is 20 % conserved, an N-linked site at either N332 or N334 together is 92 % conserved
5.5 Nature of the N-Linked Glycans on gp120
N-linked glycosylation of Env is carried out by the host cell glycosylation machinery, so that the glycan sugars are chemically identical to those found on human proteins and should appear as self to our immune systems. The glycosylation process starts during protein synthesis and continues in the endoplasmic reticulum, where glycans are initially linked to asparagines as a GlcNAc2Man9Glu3 (for review, see Schwarz and Aebi 2011). In mammalian systems, sugars are trimmed from that glycan to a smaller form (GlcNAc2Man5) as the protein moves towards the Golgi. There, further processing takes place to delete and add back sugars to create the final glycoforms that can be classified as either high-mannose, complex, or hybrid, depending on their composition. The final glycan composition can also vary, depending on the type of cell producing the virus, and depending on the physical accessibility of the glycan to processing enzymes. It is clear that N-linked glycans play a vital role for the virus because nearly 50 % of Env’s molecular weight is carbohydrate. Early structural studies of gp120 did not focus on the N-linked glycans because their heterogeneity hampered structural characterization and, therefore, they were routinely enzymatically cleaved or removed by mutagenesis. However, by modeling glycans onto Env structures determined by crystallography and EM, it appears that, within the context of the trimer, the N-linked glycans effectively cover most of the underlying protein surface that would otherwise be solvent-exposed (Kwong et al. 2000b; Binley et al. 2010; Kong et al. 2010).
Using glycans as a steric shield provides a cost-effective countermeasure to antibody recognition because the addition of just a few N-linked glycans can sterically protect a large surface area. A single N-linked glycan is not an effective protective barrier because antibodies can usually still access the protein surface and largely avoid the glycan. However, a cluster of three or more closely spaced glycans would effectively prevent antibody binding to the protein surface below. The 15–32 glycans on gp120 cluster in this manner, raising the possibility that changes at one glycan position may affect the composition and physical arrangement of the entire shield. For example, it was found that removal of glycans near the CD4-binding site of the gp120 core protein abolished antibody binding to the V3 loop, a region over 20 Å away from the glycan deletion (Wei et al. 2003). However, densely packed N-linked glycans not only block binding by antibodies but also glycan-processing enzymes in the Golgi, resulting in regions with relatively homogenous glycoforms that are not typically found on host glycoproteins. Indeed, studies of glycans on gp120 reveal predominantly high mannose types (Geyer et al. 1988), especially in a region of the outer domain of gp120 (Wyatt et al. 1998; Scanlan et al. 2002; Doores et al. 2010b), and therefore represent a weakness in the glycan shield because the clusters of high-mannose glycans are both unusual (hence regarded as nonself by the immune system) and conserved. The N332 site of vulnerability discussed above is centered on this homogeneous patch, and the N160 site appears to be particularly conserved as a Man5 glycan (McLellan et al. 2011).
Despite this weakness, anti-HIV-1-neutralizing antibodies that target glycan are not easily elicited. In fact, similar to anti-CD4-binding site antibodies (Liao et al. 2013), it takes over 2 years for infected patients to develop sera targeting these sites, by which time the virus is so established that sterilizing immunity is effectively impossible (Gray et al. 2011). Clearly, when designing a vaccine, it is impossible to completely reproduce the presentation of glycans on Env because of (1) natural variation in the N-linked sequon positions, (2) heterogeneity in the glycan composition, and (3) conformational flexibility. Understanding how bnAbs tolerate these different types of heterogeneity is, therefore, critical for vaccine design.
5.6 N-Linked Glycan Sequon Variation
All N-linked glycosylation sequons on gp120 are relatively conserved across different isolates and clades (Scanlan et al. 2002; Zhang et al. 2004) (Table 5.2). However, despite overall sequon conservation, N-linked glycan sites on gp120 migrate over the course of infection (Wei et al. 2003), and these sequon changes can significantly impact the antigenic structure of gp120, allowing for escape from the antibody response (Moore et al. 2012, 2013). However, glycans cannot occupy or cover critical parts of Env, for example, the receptor-binding site or monomer–monomer interfaces making up the trimer.
One type of glycan shifting in Env involves Nx(S/T/N)x/(S/T) motifs, where a mutation altering the central S/T/N position in this motif can easily shift a glycan by two residues along the protein chain. For example, a mutation from NxTxT to NxNxT moves the glycan attachment site by two amino-acid positions. This single amino-acid change can be created by a single nucleotide change, as two of the four codons for Thr and the two codons each for Asn and Ser are neighbors on the codon table, with ACT/ACC (Thr) and AGT/AGC (Ser) being readily convertible to AAT/AAC (Asn).
A recent study has shown the N332 site of vulnerability is subject to this kind of sequon evolution, particularly at the N332 glycan (Moore et al. 2012). It was observed that, in some patients infected with HIV-1 clade C viruses, the N332 site is absent and instead a glycan is present at N334. Over the course of infection, the N334 glycan is lost but shifts to the N332 site. As broadly neutralizing responses targeting the N332 glycan are developed, the sequon shifts back again to N334, resulting in virus escape from neutralization. It should be noted that N332 and N334 glycans cannot be present at the same time because the presence of N at position 334 would eliminate the NxS/T site of N332. Analysis of Env sequences from the Los Alamos database reveals that oscillating glycans between positions 334 and 332 may be quite common as N332 occurs in 73 % of non-redundant sequences, while N334 occurs in 20 % of non-redundant sequences. Together, 93 % of all HIV-1 viruses have a glycan in this vicinity, suggesting that it is somehow critical for the virus to shield the protein surface at this location. Notwithstanding, PGT 128, a broadly neutralizing antibody against the N332 site, can bind to some viruses with either N332 or N334 glycans (Walker et al. 2011a). The crystal structure of PGT 128 in complex with an outer domain construct of gp120 shows that the antibody contacts mostly the N332 and N301 glycans, and a portion of the V3 loop nonspecifically through backbone contacts. Thus, it might seem quite reasonable that PGT 128 can tolerate this sequon shifting because the N332 and N334 positions are very close to each other and the protein–protein interaction is between flexible loops that could likely accommodate some small variation in the binding mode. However, PGT 128 cannot bind to every isolate with an N334 glycan (Moore et al. 2012) and PGT 128’s tolerance to the glycan shifting may depend on neighboring amino-acid composition and glycosylation.
Similarly, PGT 128 can bind to and neutralize some isolates that do not have the N332 glycan, relying instead on the N295 glycan (Walker et al. 2011a). Although these glycan sites are ~8 Å apart, they are nearly equidistant from N301 (19 Å for N332 and 22 Å for N295), suggesting that either PGT 128 or the glycans can slightly alter their positions to enable recognition. However, 69 % of isolates have both N332 and N301 glycans, matching the 70 % neutralization breadth of PGT 128 without needing any contribution from the N295 glycan, which is 59 % conserved. The ability to bind both N332 and N295 glycans may also compensate for leaky sequon usage, since not all glycosylation sites are 100 % glycosylated.
2G12 is another bnAb that recognizes glycans around the N332 site of vulnerability with considerable flexibility. From mutational studies, the 2G12-binding site includes the glycans at N295, N332, N339, and N386 (Trkola et al. 1996; Scanlan et al. 2002). From these studies, it was observed that 2G12 could also bind N334 in lieu of N332 and N397 in lieu of N339 (Trkola et al. 1996). This promiscuity is not unexpected because 2G12 contacts the tips of glycans, which are flexible enough to compensate for the few angstroms of translational changes due to sequon shifting. In fact, 2G12 can bind glycan tips presented in a variety of contexts: (1) synthetic mannosides on gold particles (Marradi et al. 2011); (2) glycoconjugates made of polyamidoamine (Kabanova et al. 2010); (3) oligosaccharides scaffolded on DNA (Gorska et al. 2009); (4) synthetic glycans conjugated on lysines on bovine serum albumin (Astronomo et al. 2008); (5) oligomannose dendrons (Wang et al. 2008); (6) glycoproteins from Saccharomyces cerevisae engineered to incorporate high mannose glycans (Luallen et al. 2008, 2009; Dunlop et al. 2010; Agrawal-Gamse et al. 2011); and (7) lipo-oligosaccharides from soil bacteria (Clark et al. 2012). Furthermore, when gp120 is expressed in the presence of kifunensine to yield homogeneous GlcNAc2Man9 type glycans, it can bind to 2G12 in a 1:2 ratio (Scanlan et al. 2007) as a secondary high-mannose-binding site is apparently created.
PGT 135, a bnAb that binds to glycans at N386, N392, and N332, also exhibits a degree of promiscuity. From the crystal structure, it is clear that the PGT 135 epitope is centered on and between these glycans, involving the entire faces of N332 and N392 glycans as well as a substantial portion of the protein surface. Therefore, it is a little unexpected to find that, for some HIV-1 isolates, PGT 135 requires the presence of N295, a glycan that is 10 Å away and blocked from PGT 135 contact by the N332 glycan. This scenario raises the possibility that the presence of N295 in some isolates impacts the conformation and/or chemical composition of the glycans that PGT 135 directly binds.
It is remarkable how HIV-1 adopts an evolving glycan shield to block the antibody response but perhaps it is even more surprising that the immune system can elicit bnAbs to overcome this strategy through flexible glycan recognition. Even though the N-linked glycans are relatively conserved in Env, flexibility in their recognition is important because no individual glycan is completely conserved across all clades, sequon usage is not 100 %, and each glycan is relatively dispensable in terms of affecting protein folding (Ohgimoto et al. 1998), thus allowing HIV-1 to tolerate glycan-deletion and glycan-addition mutations. However, knowing that promiscuity is common among glycan-recognizing antibodies does not translate easily to immunogen design. Thus, further structural insights are likely going to be needed on how the same bnAb can recognize different glycans through subtly varying its binding mode and the exact epitope it recognizes.
5.7 Variation in N-Linked Glycan Composition
Unlike protein sequence, the composition of an N-linked glycan is not dictated by a stable chemical transcript, but rather by solvent accessibility and the availability of glycosidases that modify the glycan in the Golgi, parameters that are stochastic in nature. Consequently, it is interesting that antibodies against glycosylated epitopes can neutralize so broadly and potently. Structural studies of bnAbs PGT 135, PGT 128, 2G12, and PG9 have revealed some of the ways these antibodies overcome the heterogeneity. It is important to note that glycan–protein interactions typically have dissociation constants in the 10−3–10−6 M range, which are much weaker than affinity-matured antibody–antigen interactions with dissociation constants typically 10−8–10−12 M (Liang et al. 2007). This discrepancy holds true even although both types of interfaces are mediated by similar van der Waals and hydrophilic interactions. The low affinity of proteins for glycans is perhaps due to the highly soluble nature of glycans, which prefer interactions with water rather than with protein (Bundle and Young 1992). Anti-glycan bnAbs exhibit similar low affinities for glycans; for example, on a neoglycolipid microarray, PGT 128, which is observed to interact with GlcNAc2Man8 and GlcNAc2Man5 containing glycans at N332 and N301, respectively, in the crystal structure, binds well to GlcNAc2Man8–9 on the array, but not to smaller glycans (Pejchal et al. 2011). 2G12 has substantially lower avidity towards these glycans on the array, and PGT 135 has even lower avidity still (Kong et al. 2013). Finally, Fab PG9 and Fab PG16 affinities for individual glycans are so low that NMR saturation experiments were necessary to confirm dissociation constants in the millimolar range (McLellan et al. 2011). Nonetheless, these antibodies can achieve avidity with dissociation constants of ~10−9 M towards the glycoprotein by binding to multiple glycans and/or protein surface components, where each individual interaction can still have relatively low affinity. Thus, minor variations in glycan composition may slightly modify the affinity for already weak individual interactions without abrogating overall binding. However, studies have shown that certain glycoforms do completely abolish bnAb binding: (1) PGT 135 and PG9 (Doores et al. 2010b), but not 2G12 or PGT 128, lose binding if gp120 is expressed in the presence of kifunensine, which results in predominantly GlcNAc2Man9 glycans; and (2) PGT 128 and 2G12, but not PGT 135, lose binding if gp120 is expressed in the presence of N-butyldeoxynojirimycin (NB-DNJ) and kifunensine, which can lead to either blockage of the tip of the D1 arm with 1–3 glucose residues or shortening of the D1 arm by removal of all Manα1,2Man-linked residues (Roth et al. 2003; Doores and Burton 2010). Thus, it is very important for immunogen design to know exactly which glycoforms can or cannot be tolerated and which are present on each virus.
For example, structural and biophysical studies have confirmed that 2G12 binds to the terminal mannose extensions of N-linked glycans with its two primary and two secondary binding sites. Crystal structures suggest that 2G12 binds to the D1 arm of N295 and N392/N386 glycans at the primary binding sites, and to the D2 and D3 arms of N339 and N332 glycans at the secondary binding sites (Calarese et al. 2003, 2005). This result suggests that 2G12 can tolerate variation in the D2 and D3 arms on N295 and N392/N386 glycans, and in the D1 arms of the N339 and N332 glycans. Lack of one or two terminal mannoses that directly bind to 2G12 may also be tolerated since the antibody would still maintain interactions with 2–3 remaining glycans. The full extent of 2G12 tolerance remains to be tested in a synthetic glycopeptide context where glycan compositions can be chemically controlled.
In contrast to 2G12, which binds to the glycan tips, PGT 128, PGT 135, and PG9 use extended CDR loops to reach the underlying protein surface while making contacts across the lateral faces of the glycans. Substantial surface area on each glycan is buried by these antibodies on the order of 300–500 Å2, a size comparable to those of some entire antibody epitopes. However, the size of the buried surface does not correlate with the low affinity for individual glycans. One advantage to making contacts across the glycan face is increased dependence on the more protein-proximal glycan moieties that are generally invariant across glycoforms. For example, in the crystal structure, PGT 128 interacts with a glycan at position N301 that displays enough electron density to be minimally consistent with a GlcNAc2Man5. However, higher oligomannose or even hybrid type glycans containing this set of glycan monomers would also be consistent with the density. Site-specific analysis of gp120 glycans suggests that this site may in fact be complex or hybrid type glycan (Leonard et al. 1990; Go et al. 2008, 2009, 2011). Thus, by making contacts only to the highly conserved moieties within the GlcNAc2Man5 core, PGT 128 is able to accommodate substantial glycoform variation. Similarly, PG9 recognizes GlcNAc2Man4 at position N156 in a manner that also allows comparable glycoform variation. More extremely, PGT 135 contacts the glycan at N386 only from the GlcNAc stem to the β-mannose, potentially allowing the antibody to tolerate all N-linked glycan types.
One risk in raising antibodies only against the conserved lateral face of N-linked glycans is that the antibodies may interact closely with the distal ends of the glycan monomers, preventing the antibodies from binding glycans that have further extensions from the conserved core. This sort of interaction is observed for PG9 binding to the N160 GlcNAc2Man5 glycan. In the crystal structure, further extensions of D1 and D2 arms of the N160 glycan would impede binding of the PG9 antibody. For example, a GlcNAc2Man7–9 at N160 would be sterically impossible within the binding site, explaining the lack of PG9 activity when the antigen is grown in kifunensine. However, site-specific glycan analysis suggests that the N160 site is predominantly GlcNAc2Man5, explaining in part the broad neutralization by PG9. Another example is the interaction between PGT 135 and N392 glycan. In the crystal structure, there is visible electron density for a GlcNAc2Man8 glycan at this position with complete D1 and D3 arms. PGT 135 binds across the glycan’s D1 arm and buries the D2 arm. A full D2 arm as in GlcNAc2Man9 would result in a severe clash with PGT 135, preventing the antibody from binding. Thus, small variation in high mannose glycoforms can have significant impact on bnAb activity.
From these structural studies, it is clear that site-specific characterization of N-linked glycans within the bnAb epitope is as critical to vaccine design as the exact mapping of sites of vulnerability across protein surfaces.
5.8 Conformational Variation of Glycans
It has long been supposed that structures of N-linked glycans are difficult to characterize by X-ray crystallography due to their exceptional conformational flexibility. Electron density maps of glycans tend to be disordered except at the protein-proximal GlcNAc unless the glycan is stabilized by crystal contacts or by ligand binding. It is also challenging to characterize glycan structure by NMR due to the similar chemical environment of protons and the paucity of protons between connected glycan monomers that are less than 5 Å, the detection limit for Nuclear Overhauser effects. Structures from molecular dynamics simulations using NMR constraints tend to be less flexible, but it has been argued that the constraints are from time-averaged data lacking the resolution for movement (Woods and Tessier 2010). Nonetheless, analyses of available glycan structures show strong preferences for defined phi and psi torsion angles with minimal energy cost (Petrescu et al. 1999; Lutteke et al. 2006; Lutteke 2009). When attached to a protein, glycan conformation may be further constrained, especially when stabilized by neighboring aromatic residues that interact with the protein-proximal GlcNAc (Petrescu et al. 2004; Culyba et al. 2011; Price et al. 2012). This type of stabilization may be occurring for glycan N332 on HIV-1 gp120, which has a histidine or tyrosine at position 330 in 96 % of isolates. In general, glycans on gp120 may also be stabilized by being densely packed within the glycan shield.
The initial orientation of a whole glycan may not be critical to antibody recognition if the antibody only recognizes a contiguous region on a single glycan and, thus, is not constrained by contacts to other parts of the antigen to approach the glycan from a particular angle. However, in the case of some bnAbs that bind across the lateral faces of multiple glycans and protein surfaces, the initial orientation of the whole glycan and its degree of flexibility may be important because the angle of approach to the glycan would be constrained by those other contacts. For example, PGT 135 buries a large protein surface between the N332 and N392 glycans on gp120, which constrains the antibody to recognize a particular orientation of glycans N332 and N392 and approach from a particular angle. Remarkably, PGT 135 may be able to tolerate small differences in the glycan orientations by adopting degenerate angles of approach as detected by EM studies (Kong et al. 2013). PG9 is similarly constrained to bind the N160 glycan from one side, but PGT 128 may be less constrained because it binds to a potentially flexible portion of gp120. In fact, the protein–protein and protein–glycan N301 interactions do not restrict PGT 128 from binding N295 when N332 is unavailable. However, the N332/N295 glycan necessary for PGT 128 binding must be in the proper conformation so that the correct glycan face is accessible to PGT 128. Thus, it is clear that for these antibodies, conformations of the glycans play an important role in their recognition, and perhaps in antibody elicitation.
The only way to elucidate glycan conformation and antibody binding relationships is through structural characterization at relatively high resolution. Unfortunately, there are only four crystal structures of glycan-binding bnAbs to HIV-1. However, three of these structures, 2G12, PGT 135, and PGT 128, focus on the N332 site of vulnerability, allowing for some degree of comparative analysis. All three structures have the N332 glycan, and both the 2G12 and the PGT 135 structures have the N392 glycan. A caveat for the comparison is that the PGT 135 structure contains a glycosylated gp120 core while the PGT 128 and 2G12 structures contain different outer domain fragments with different levels of glycosylation. Nevertheless, the N332 glycan in all three structures remains relatively unchanged in terms of overall orientation and conformation. In contrast, the N392 glycan conformations in the PGT 135 and 2G12 structures are significantly different, with the terminal mannose arms pointing in almost opposite directions. The conformational stability of the N332 glycan may explain why it is commonly targeted in broadly neutralizing sera, especially if the antibodies in the sera are predominantly binding across one of its faces along with other elements on gp120.
The biophysical properties of a glycan face are determined by the corresponding properties of the individual glycan monomers along that face. Although N-linked glycans are highly polar and soluble, each glycan monomer sugar has both a polar and a hydrophobic face that is dictated by the direction that the hydroxyls are pointing. This asymmetry corresponds to the ring pucker of the chair conformation, and can be defined by whether the carbons around the ring are arranged counter clockwise (hydrophobic face) or clockwise (polar face). The individual glycan monomers that make up the N332 glycan are similarly aligned across the crystal structures and determine the properties of the intact glycan faces being recognized by different bnAbs. PGT 135 appears to recognize a more hydrophobic face of the N332 glycan, with 62 % of the buried surface area on the glycan consisting of hydrophobic ring faces. This matches the 64 % of the buried surface on PGT 135 that is contributed by hydrophobic residues. PGT 128 recognizes a more polar side of the glycan, with only 31 % of the buried surface on the glycan consisting of hydrophobic ring faces. High-resolution structures of other N332-directed bnAbs are thus needed to fully define the relevant faces of glycans that can be recognized.
N-linked glycans are large protein adducts, potentially extending beyond 20 Å, making them comparable to small protein loops. Clearly it is insufficient to simply define which glycans an antibody binds to, or to think of a glycan as comparable to a single amino acid. The structural studies described above highlight the importance of defining the particular orientation and the different faces of an N-linked glycan that are being recognized.
5.9 Conclusion
Over the past decade, structural studies of bnAbs bound to Env have revealed critical targets for vaccine design. Some of targets like the CD4-binding site and the MPER are recessed or partially occluded within Env trimer, severely limiting the angle of approach antibodies can take to interact with them. The glycosylated epitopes of glycan-binding bnAbs are largely free from such constraints because they are highly accessible in the trimer and crystal structures clearly show that a wide range of approach angles is allowed to the Env trimer surface (Fig. 5.1). However, unlike protein-binding bnAbs, glycan-binding bnAbs have epitopes that encompass the conformational and chemical features of “self” glycans, but are packaged on the virus in a nonself way, and often exhibit multiple levels of heterogeneity. High-resolution structural studies have revealed the types of glycoforms that can be accommodated by the bnAbs. They have also broadly outlined requirements for specific conformations and overall orientations of the targeted glycans. Despite all these advances, further structural studies are required to fully define the spectrum of glycosylated targets on HIV-1 Env and biochemical studies to define the exact nature of the glycans and their heterogeneity at each individual glycan position in different strains and clades, as well as for Env produced in different cell lines.
References
Agrawal-Gamse C, Luallen RJ, Liu B et al (2011) Yeast-elicited cross-reactive antibodies to HIV Env glycans efficiently neutralize virions expressing exclusively high-mannose N-linked glycans. J Virol 85:470–480
Astronomo RD, Lee HK, Scanlan CN et al (2008) A glycoconjugate antigen based on the recognition motif of a broadly neutralizing human immunodeficiency virus antibody, 2G12, is immunogenic but elicits antibodies unable to bind to the self glycans of gp120. J Virol 82:6359–6368
Baba TW, Liska V, Hofmann-Lehmann R et al (2000) Human neutralizing monoclonal antibodies of the IgG1 subtype protect against mucosal simian-human immunodeficiency virus infection. Nat Med 6:200–206
Binley JM, Ban YE, Crooks ET et al (2010) Role of complex carbohydrates in human immunodeficiency virus type 1 infection and resistance to antibody neutralization. J Virol 84:5637–5655
Bonsignori M, Hwang KK, Chen X et al (2011) Analysis of a clonal lineage of HIV-1 envelope V2/V3 conformational epitope-specific broadly neutralizing antibodies and their inferred unmutated common ancestors. J Virol 85:9998–10009
Bundle DR, Young NM (1992) Carbohydrate-protein interactions in antibodies and lectins. Curr Opin Struct Biol 2:666–673
Calarese DA, Scanlan CN, Zwick MB et al (2003) Antibody domain exchange is an immunological solution to carbohydrate cluster recognition. Science 300:2065–2071
Calarese DA, Lee HK, Huang CY et al (2005) Dissection of the carbohydrate specificity of the broadly neutralizing anti-HIV-1 antibody 2G12. Proc Natl Acad Sci U S A 102:13372–13377
Cardoso RM, Zwick MB, Stanfield RL et al (2005) Broadly neutralizing anti-HIV antibody 4E10 recognizes a helical conformation of a highly conserved fusion-associated motif in gp41. Immunity 22:163–173
Cardoso RM, Brunel FM, Ferguson S et al (2007) Structural basis of enhanced binding of extended and helically constrained peptide epitopes of the broadly neutralizing HIV-1 antibody 4E10. J Mol Biol 365:1533–1544
Chang H, Biswas S, Tallarico AS et al (2012) Human B-cell ontogeny in humanized NOD/SCID gammac(null) mice generates a diverse yet auto/poly- and HIV-1-reactive antibody repertoire. Genes Immun 13:399–410
Clark BE, Auyeung K, Fregolino E et al (2012) A bacterial lipooligosaccharide that naturally mimics the epitope of the HIV-neutralizing antibody 2G12 as a template for vaccine design. Chem Biol 19:254–263
Colman PM, Lawrence MC (2003) The structural biology of type I viral membrane fusion. Nat Rev Mol Cell Biol 4:309–319
Connolly ML (1993) The molecular surface package. J Mol Graph 11:139–141
Corti D, Langedijk JP, Hinz A et al (2010) Analysis of memory B cell responses and isolation of novel monoclonal antibodies with neutralizing breadth from HIV-1-infected individuals. PLoS One 5:e8805
Crooks ET, Tong T, Osawa K et al (2011) Enzyme digests eliminate nonfunctional Env from HIV-1 particle surfaces, leaving native Env trimers intact and viral infectivity unaffected. J Virol 85:5825–5839
Culyba EK, Price JL, Hanson SR et al (2011) Protein native-state stabilization by placing aromatic side chains in N-glycosylated reverse turns. Science 331:571–575
Doores KJ, Burton DR (2010) Variable loop glycan dependency of the broad and potent HIV-1-neutralizing antibodies PG9 and PG16. J Virol 84:10510–10521
Doores KJ, Bonomelli C, Harvey DJ et al (2010a) Envelope glycans of immunodeficiency virions are almost entirely oligomannose antigens. Proc Natl Acad Sci U S A 107:13800–13805
Doores KJ, Fulton Z, Huber M et al (2010b) Antibody 2G12 recognizes di-mannose equivalently in domain- and nondomain-exchanged forms but only binds the HIV-1 glycan shield if domain exchanged. J Virol 84:10690–10699
Dunlop DC, Bonomelli C, Mansab F et al (2010) Polysaccharide mimicry of the epitope of the broadly neutralizing anti-HIV antibody, 2G12, induces enhanced antibody responses to self oligomannose glycans. Glycobiology 20:812–823
Fox JL (2011) HIV drugs made in tobacco. Nat Biotechnol 29:852
Freed EO (2001) HIV-1 replication. Somat Cell Mol Genet 26:13–33
Geyer H, Holschbach C, Hunsmann G et al (1988) Carbohydrates of human immunodeficiency virus. Structures of oligosaccharides linked to the envelope glycoprotein 120. J Biol Chem 263:11760–11767
Go EP, Irungu J, Zhang Y et al (2008) Glycosylation site-specific analysis of HIV envelope proteins (JR-FL and CON-S) reveals major differences in glycosylation site occupancy, glycoform profiles, and antigenic epitopes’ accessibility. J Proteome Res 7:1660–1674
Go EP, Chang Q, Liao HX et al (2009) Glycosylation site-specific analysis of clade C HIV-1 envelope proteins. J Proteome Res 8:4231–4242
Go EP, Hewawasam G, Liao HX et al (2011) Characterization of glycosylation profiles of HIV-1 transmitted/founder envelopes by mass spectrometry. J Virol 85:8270–8284
Gorska K, Huang KT, Chaloin O et al (2009) DNA-templated homo- and heterodimerization of peptide nucleic acid encoded oligosaccharides that mimick the carbohydrate epitope of HIV. Ang Chem Int Ed Engl 48:7695–7700
Gray ES, Madiga MC, Hermanus T et al (2011) The neutralization breadth of HIV-1 develops incrementally over four years and is associated with CD4+ T cell decline and high viral load during acute infection. J Virol 85:4828–4840
Hallenberger S, Bosch V, Angliker H et al (1992) Inhibition of furin-mediated cleavage activation of HIV-1 glycoprotein gp160. Nature 360:358–361
Harris A, Borgnia MJ, Shi D et al (2011) Trimeric HIV-1 glycoprotein gp140 immunogens and native HIV-1 envelope glycoproteins display the same closed and open quaternary molecular architectures. Proc Natl Acad Sci U S A 108:11440–11445
Haynes BF, Fleming J, St Clair EW et al (2005) Cardiolipin polyspecific autoreactivity in two broadly neutralizing HIV-1 antibodies. Science 308:1906–1908
Huang CC, Tang M, Zhang MY et al (2005) Structure of a V3-containing HIV-1 gp120 core. Science 310:1025–1028
Huang J, Ofek G, Laub L et al (2012) Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491:406–412
Huber M, von Wyl V, Ammann CG et al (2008) Potent human immunodeficiency virus-neutralizing and complement lysis activities of antibodies are not obligatorily linked. J Virol 82:3834–3842
Huber M, Le KM, Doores KJ et al (2010) Very few substitutions in a germ line antibody are required to initiate significant domain exchange. J Virol 84:10700–10707
Joos B, Trkola A, Kuster H et al (2006) Long-term multiple-dose pharmacokinetics of human monoclonal antibodies (MAbs) against human immunodeficiency virus type 1 envelope gp120 (MAb 2G12) and gp41 (MAbs 4E10 and 2F5). Antimicrob Agents Chemother 50:1773–1779
Julien JP, Bryson S, Nieva JL et al (2008) Structural details of HIV-1 recognition by the broadly neutralizing monoclonal antibody 2F5: epitope conformation, antigen-recognition loop mobility, and anion-binding site. J Mol Biol 384:377–392
Julien J-P, Khayat R, Sok D et al (2013a) Broadly neutralizing antibody PGT121 allosterically modulates CD4 binding via recognition of the HIV-1 gp120 V3 base and multiple surrounding glycans. PLoS Pathog 9:e1003342
Julien JP, Lee JH, Cupo A et al (2013b) Asymmetric recognition of the HIV-1 trimer by broadly neutralizing antibody PG9. Proc Natl Acad Sci U S A 110:4351–4356
Kabanova A, Adamo R, Proietti D et al (2010) Preparation, characterization and immunogenicity of HIV-1 related high-mannose oligosaccharides-CRM197 glycoconjugates. Glycoconj J 27: 501–513
Knipe DM, Howley PM (2007) Fields virology. Lippincott Williams & Wilkins, Philadelphia, pp 2107–2186
Kong L, Sheppard NC, Stewart-Jones GB et al (2010) Expression-system-dependent modulation of HIV-1 envelope glycoprotein antigenicity and immunogenicity. J Mol Biol 403:131–147
Kong L, Julien J-P, Calarese D et al (2012) Toward a carbohydrate-based HIV-1 vaccine. In: Klyosov AA (ed) Glycobiology and drug design, vol 1102. ACS Publications, Washington, DC, pp 187–215
Kong L, Lee JH, Doores KJ et al (2013) Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat Struct Mol Biol 20:796–803
Kunert R, Ruker F, Katinger H (1998) Molecular characterization of five neutralizing anti-HIV type 1 antibodies: identification of nonconventional D segments in the human monoclonal antibodies 2G12 and 2F5. AIDS Res Hum Retroviruses 14:1115–1128
Kwong PD, Wyatt R, Robinson J et al (1998) Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393:648–659
Kwong PD, Wyatt R, Majeed S et al (2000a) Structures of HIV-1 gp120 envelope glycoproteins from laboratory-adapted and primary isolates. Structure 8:1329–1339
Kwong PD, Wyatt R, Sattentau QJ et al (2000b) Oligomeric modeling and electrostatic analysis of the gp120 envelope glycoprotein of human immunodeficiency virus. J Virol 74:1961–1972
Leonard CK, Spellman MW, Riddle L et al (1990) Assignment of intrachain disulfide bonds and characterization of potential glycosylation sites of the type 1 recombinant human immunodeficiency virus envelope glycoprotein (gp120) expressed in Chinese hamster ovary cells. J Biol Chem 265:10373–10382
Liang PH, Wang SK, Wong CH (2007) Quantitative analysis of carbohydrate-protein interactions using glycan microarrays: determination of surface and solution dissociation constants. J Am Chem Soc 129:11177–11184
Liao HX, Chen X, Munshaw S et al (2011) Initial antibodies binding to HIV-1 gp41 in acutely infected subjects are polyreactive and highly mutated. J Exp Med 208:2237–2249
Liao HX, Lynch R, Zhou T et al (2013) Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496:469–476
Liu J, Bartesaghi A, Borgnia MJ et al (2008) Molecular architecture of native HIV-1 gp120 trimers. Nature 455:109–113
Luallen RJ, Lin J, Fu H et al (2008) An engineered Saccharomyces cerevisiae strain binds the broadly neutralizing human immunodeficiency virus type 1 antibody 2G12 and elicits mannose-specific gp120-binding antibodies. J Virol 82:6447–6457
Luallen RJ, Fu H, Agrawal-Gamse C et al (2009) A yeast glycoprotein shows high-affinity binding to the broadly neutralizing human immunodeficiency virus antibody 2G12 and inhibits gp120 interactions with 2G12 and DC-SIGN. J Virol 83:4861–4870
Lutteke T (2009) Analysis and validation of carbohydrate three-dimensional structures. Acta Crystallogr D65:156–168
Lutteke T, Bohne-Lang A, Loss A et al (2006) GLYCOSCIENCES.de: an Internet portal to support glycomics and glycobiology research. Glycobiology 16:71R–81R
Manrique A, Rusert P, Joos B et al (2007) In vivo and in vitro escape from neutralizing antibodies 2G12, 2F5, and 4E10. J Virol 81:8793–8808
Marradi M, Di Gianvincenzo P, Enriquez-Navas PM et al (2011) Gold nanoparticles coated with oligomannosides of HIV-1 glycoprotein gp120 mimic the carbohydrate epitope of antibody 2G12. J Mol Biol 410:798–810
Mascola JR, Lewis MG, Stiegler G et al (1999) Protection of macaques against pathogenic simian/human immunodeficiency virus 89.6PD by passive transfer of neutralizing antibodies. J Virol 73:4009–4018
Mascola JR, Stiegler G, VanCott TC et al (2000) Protection of macaques against vaginal transmission of a pathogenic HIV-1/SIV chimeric virus by passive infusion of neutralizing antibodies. Nat Med 6:207–210
McLellan JS, Pancera M, Carrico C et al (2011) Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480:336–343
Mehandru S, Vcelar B, Wrin T et al (2007) Adjunctive passive immunotherapy in human immunodeficiency virus type 1-infected individuals treated with antiviral therapy during acute and early infection. J Virol 81:11016–11031
Montero M, van Houten NE, Wang X et al (2008) The membrane-proximal external region of the human immunodeficiency virus type 1 envelope: dominant site of antibody neutralization and target for vaccine design. Microbiol Mol Biol Rev 72:54–84
Moore PL, Crooks ET, Porter L et al (2006) Nature of nonfunctional envelope proteins on the surface of human immunodeficiency virus type 1. J Virol 80:2515–2528
Moore PL, Gray ES, Wibmer CK et al (2012) Evolution of an HIV glycan-dependent broadly neutralizing antibody epitope through immune escape. Nat Med 18:1688–1692
Moore PL, Sheward D, Nonyane M et al (2013) Multiple pathways of escape from HIV broadly cross-neutralizing V2-dependent antibodies. J Virol 87:4882–4894
Mouquet H, Klein F, Scheid JF et al (2011) Memory B cell antibodies to HIV-1 gp140 cloned from individuals infected with clade A and B viruses. PLoS One 6:e24078
Ofek G, Tang M, Sambor A et al (2004) Structure and mechanistic analysis of the anti-human immunodeficiency virus type 1 antibody 2F5 in complex with its gp41 epitope. J Virol 78: 10724–10737
Ohgimoto S, Shioda T, Mori K et al (1998) Location-specific, unequal contribution of the N glycans in simian immunodeficiency virus gp120 to viral infectivity and removal of multiple glycans without disturbing infectivity. J Virol 72:8365–8370
Pancera M, McLellan JS, Wu X et al (2010) Crystal structure of PG16 and chimeric dissection with somatically related PG9: structure-function analysis of two quaternary-specific antibodies that effectively neutralize HIV-1. J Virol 84:8098–8110
Pejchal R, Walker LM, Stanfield RL et al (2010) Structure and function of broadly reactive antibody PG16 reveal an H3 subdomain that mediates potent neutralization of HIV-1. Proc Natl Acad Sci U S A 107:11483–11488
Pejchal R, Doores KJ, Walker LM et al (2011) A potent and broad neutralizing antibody recognizes and penetrates the HIV glycan shield. Science 334:1097–1103
Petrescu AJ, Petrescu SM, Dwek RA et al (1999) A statistical analysis of N- and O-glycan linkage conformations from crystallographic data. Glycobiology 9:343–352
Petrescu AJ, Milac AL, Petrescu SM et al (2004) Statistical analysis of the protein environment of N-glycosylation sites: implications for occupancy, structure, and folding. Glycobiology 14: 103–114
Plotkin SA (2010) Correlates of protection induced by vaccination. Clin Vaccine Immunol 17: 1055–1065
Price JL, Culyba EK, Chen W et al (2012) N-glycosylation of enhanced aromatic sequons to increase glycoprotein stability. Biopolymers 98:195–211
Roth J, Ziak M, Zuber C (2003) The role of glucosidase II and endomannosidase in glucose trimming of asparagine-linked oligosaccharides. Biochimie 85:287–294
Sanders RW, Venturi M, Schiffner L et al (2002) The mannose-dependent epitope for neutralizing antibody 2G12 on human immunodeficiency virus type 1 glycoprotein gp120. J Virol 76: 7293–7305
Scanlan CN, Pantophlet R, Wormald MR et al (2002) The broadly neutralizing anti-human immunodeficiency virus type 1 antibody 2G12 recognizes a cluster of alpha1→2 mannose residues on the outer face of gp120. J Virol 76:7306–7321
Scanlan CN, Ritchie GE, Baruah K et al (2007) Inhibition of mammalian glycan biosynthesis produces non-self antigens for a broadly neutralising, HIV-1 specific antibody. J Mol Biol 372:16–22
Schneider J, Kaaden O, Copeland TD et al (1986) Shedding and interspecies type sero-reactivity of the envelope glycopolypeptide gp120 of the human immunodeficiency virus. J Gen Virol 67:2533–2538
Schwarz F, Aebi M (2011) Mechanisms and principles of N-linked protein glycosylation. Curr Opin Struct Biol 21:576–582
Stanfield RL, Gorny MK, Williams C et al (2004) Structural rationale for the broad neutralization of HIV-1 by human monoclonal antibody 447-52D. Structure 12:193–204
Stanfield RL, Gorny MK, Zolla-Pazner S et al (2006) Crystal structures of human immunodeficiency virus type 1 (HIV-1) neutralizing antibody 2219 in complex with three different V3 peptides reveal a new binding mode for HIV-1 cross-reactivity. J Virol 80:6093–6105
Tran EE, Borgnia MJ, Kuybeda O et al (2012) Structural mechanism of trimeric HIV-1 envelope glycoprotein activation. PLoS Pathog 8:e1002797
Trkola A, Purtscher M, Muster T et al (1996) Human monoclonal antibody 2G12 defines a distinctive neutralization epitope on the gp120 glycoprotein of human immunodeficiency virus type 1. J Virol 70:1100–1108
Vcelar B, Stiegler G, Wolf HM et al (2007) Reassessment of autoreactivity of the broadly neutralizing HIV antibodies 4E10 and 2F5 and retrospective analysis of clinical safety data. AIDS 21: 2161–2170
Walker LM, Phogat SK, Chan-Hui PY et al (2009) Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326:285–289
Walker LM, Huber M, Doores KJ et al (2011a) Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477:466–470
Walker LM, Sok D, Nishimura Y et al (2011b) Rapid development of glycan-specific, broad, and potent anti-HIV-1 gp120 neutralizing antibodies in an R5 SIV/HIV chimeric virus infected macaque. Proc Natl Acad Sci U S A 108:20125–20129
Wang SK, Liang PH, Astronomo RD et al (2008) Targeting the carbohydrates on HIV-1: interaction of oligomannose dendrons with human monoclonal antibody 2G12 and DC-SIGN. Proc Natl Acad Sci U S A 105:3690–3695
Wei X, Decker JM, Wang S et al (2003) Antibody neutralization and escape by HIV-1. Nature 422:307–312
WHO (2011) Global HIV/AIDS response: epidemic update and health sector progress towards universal access. Progress report 2011, WHO Press.
Woods RJ, Tessier MB (2010) Computational glycoscience: characterizing the spatial and temporal properties of glycans and glycan-protein complexes. Curr Opin Struct Biol 20:575–583
Wu X, Yang ZY, Li Y et al (2010) Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329:856–861
Wyatt R, Kwong PD, Desjardins E et al (1998) The antigenic structure of the HIV gp120 envelope glycoprotein. Nature 393:705–711
Zhang M, Gaschen B, Blay W et al (2004) Tracking global patterns of N-linked glycosylation site variation in highly variable viral glycoproteins: HIV, SIV, and HCV envelopes and influenza hemagglutinin. Glycobiology 14:1229–1246
Zhou T, Xu L, Dey B et al (2007) Structural definition of a conserved neutralization epitope on HIV-1 gp120. Nature 445:732–737
Zhou T, Georgiev I, Wu X et al (2010) Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science 329:811–817
Zhu P, Liu J, Bess J Jr et al (2006) Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature 441:847–852
Zhu P, Winkler H, Chertova E et al (2008) Cryoelectron tomography of HIV-1 envelope spikes: further evidence for tripod-like legs. PLoS Pathog 4:e1000203
Acknowledgments
Support is acknowledged from NIH Scripps CHAVI-ID AI100663, HIVRAD P01 AI082362, R01 AI084817 (IAW), and the International AIDS Vaccine Initiative (IAVI). LK is supported by an American Foundation for AIDS Research Mathilde Krim Fellowship in Basic Biomedical Research. This is publication #23081 from The Scripps Research Institute.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Kong, L., Stanfield, R.L., Wilson, I.A. (2014). Molecular Recognition of HIV Glycans by Antibodies. In: Pantophlet, R. (eds) HIV glycans in infection and immunity. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-8872-9_5
Download citation
DOI: https://doi.org/10.1007/978-1-4614-8872-9_5
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-8871-2
Online ISBN: 978-1-4614-8872-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)