Abstract
There is a now a large body of evidence supporting the notion that cancer cells have vastly altered cell cycle networks that serve to maintain their high rate of proliferation. Consequently, targeting these pathways pharmacologically has been long studied, but only recently have some promising compounds progressed into the clinic. In this chapter, we review cell cycle function in both normal cells and describe how cancer cells deregulate this fundamental process. Next we describe in detail the development of different classes of CDK inhibitors and review the failures and successes so far, and provide insight into some future directions for research and clinical trials in order to exploit the ever-expanding molecular characterization of tumors with the drugs available and in the pipelines. In addition, we present a short overview of using differential cell cycle characteristics of normal and tumor cells as a way of protecting normal cells from cytotoxic chemotherapies. Finally we describe other potential targets such as regulating p27, inhibiting PIM and MELK kinases as well as some of the mitotic kinases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Hanahan and Weinberg recognized the importance of cell cycle and checkpoints in their original and updated “hallmarks of cancer” papers, which describe the key features that normal cells must acquire during transformation into a tumor [1, 2]. Cell cycle deregulation has long been appreciated as a fundamental early event during tumorigenesis, which contributes to several of these hallmarks, namely self-sufficiency in growth signals” and “insensitivity to anti-growth signals”, and results in genomic instability, one of the newly added hallmarks. Since these alterations are almost universal among different tumor types, cancer biologists have expended considerable effort in interrogating these pathways as therapeutic targets for 20 years. In spite of the substantial body of literature focused on identifying the biological roles of many cell cycle pathway proteins in both normal development and in describing tumor-associated defects, the progress in the clinic has not been as rapid as desired. With this in mind, we felt that this chapter would be an ideal opportunity for us to review what is known about cell cycle deregulation in cancer, with a focus on personalized treatment strategies. Ultimately, we hope to suggest future directions for research and clinical trials to utilize the wealth of genomic knowledge we now have about cancer, and design more rational strategies likely to be effective in defined genetic contexts, as well as using the cell cycle as a means of protecting normal tissues from the chemotherapeutic insults. We will utilize a particularly promising strategy from our work as an example in a subsequent section.
2 Core Cell Cycle Proteins as Targets for Therapy
2.1 Cell Cycle Regulation
Some of the best characterized cell cycle targets are key proteins that have been highly conserved throughout evolution in all eucaryotes. These include cyclins, cyclin-dependent kinases (CDKs) and cyclin-dependent kinase inhibitors (CDKi). Cyclins are master regulators of the cell cycle, via activating CDKs which in turn stimulate downstream signaling (Fig. 14.1).
Cell cycle progression is regulated by 4 major families of cyclins, cyclin D, E, A and B, and 4 respective CDKs (CDK4, 6, 2, or 1). Cyclin D is the first cyclin that is involved in the entry of cells from G0 into G1, in response to ample growth factors and other mitogens. Cyclin D exerts its activity via its catalytic partners CDK4 and CDK6, (and can also bind to CDK2 and CDK3) which phosphorylate many substrates. One of the most studied substrates of CDK4/6 is the retinoblastoma protein (Rb), which is a negative regulator of E2F transcriptional activity. Prior to phosphorylation by CDK4/6, Rb is in a hypophosphorylated state, and is bound to E2F and DP proteins, keeping E2F inactive. However CDK4/6 phosphorylation induces conformational change in Rb, releasing E2F to bind DNA and facilitate transcription. The next cyclins that are transcriptionally regulated are cyclin E and A. Cyclin E bound to CDK2 helps drive cells through G1/S transition by further phosphorylating Rb and other substrates involved in DNA replication such as cdc6. Later during S phase, CDK2 is also regulated by cyclin A levels. After DNA replication is complete, cells enter G2 phase where they prepare to enter mitosis by upregulating microtubule formation and other biosynthetic pathways necessary for chromosome segregation. Towards the end of G2 phase, CDK1 takes over as the predominant kinase, since cyclin B levels begin to rise and translocate to the nucleus to bind CDK1 to initiate the G2/M transition. This complex was first identified as the M-phase promoting factor since its main function is to break down the nuclear envelope and initiate prophase. Once mitosis is almost complete, CDK1 is deactivated via dephosphorylation, and a negative feedback loop is engaged via the anaphase-promoting complex which degrades cyclin B, allowing cells to exit mitosis.
The requirement for all of the cyclins and CDKs to control the cell cycle in normal cells as described in the previous paragraph has been recently challenged based on the findings from genetic studies in knockout mice. Tables 14.1 and 14.2 summarize the phenotypes observed in the knockout models. Each interphase CDK has been knocked out individually, and except for CDK1, all of the mice are viable. However, each mouse model has cell-type specific defects, which reveal tissue-specific roles for individual CDKs. For example, the CDK2-deficient model is viable (although born at slightly lower than predicted Mendelian ratio), but sterile due to an absolute requirement for CDK2 during meiosis in both male and female germ cells [3, 4]. Cell cycle analysis and proliferation rate of mouse embryonic fibroblasts from both CDK2 wild-type and knockout embryos showed no significant difference in cell cycle distribution, and a similar rate of proliferation for the first 4 days in culture, after which the knockout cells reached a plateau phase. Similarly CDK4 and CDK6 are not necessary for cell cycle progression in most cells, although CDK4 is required for proliferation of pancreatic β-cells, leading to a diabetic phenotype in the knockout mouse [5]. Additionally, CDK6 is important in the hematopoietic system, both in the lymphocytes and erythrocytes. CDK6-deficient mice have small, less cellular thymi, since CDK6 is downstream of Notch and AKT signaling which is critical in early thymic T cell commitment to the T-cell lineage, and also have smaller spleens with less erythroid cells [6]. Double knockout of CDK4 and CDK6 induces late embryonic lethality, primarily due to the hematopoietic defects in erythroid cell production, however MEF cells from these embryos do proliferate and can become immortalized by continuous passage. CDK2 can partially compensate to phosphorylate Rb in these double-knockout cells by binding cyclin D, and therefore promote cell cycle progression, however this is not thought to fully explain the lack of cell cycle defects systemically. In stark contrast to the phenotypes seen in CDK2/4/6 knockout models, CDK1 deficiency causes cell cycle arrest and prevents embryos from developing beyond the 2-cell stage, demonstrating a lack of compensation between the mitotic CDK and the interphase CDKs [7].
Unlike normal cells that do not depend on any single cyclin or CDK for growth, there is beginning to be some evidence that in tumor cells, altered cellular wiring can lead to oncogenic addiction to CDK signaling. For example, in a mouse model of triple negative breast cancer driven by low-molecular weight cyclin E, tumors are highly dependent upon CDK2 signaling, even though CDK2 is dispensable in normal cells [8]. This type of variation on synthetic lethality involving CDK signaling is not limited to breast cancer but can be observed in a K-Ras mutant lung cancer model. In a K-Ras-driven mouse model of lung cancer, CDK2 and CDK6 knockout only partially inhibited tumor initiation, whereas CDK4 knockout significantly decreased tumorigenesis due to an immediate induction of senescence [9]. Even though the K-Ras-mutant transgene is expressed in several other epithelial tissues in this mouse model, none of these undergo hyperplasia or tumorigenesis, and senescence is not observed in these normal tissues. CDK4 was also shown to be essential for progression of established K-Ras driven NSCLC lesions, and pharmacological CDK4 inhibition significantly inhibited tumorigenesis. The reasons for lack of immediate compensation mechanisms involving other CDKs in tumors are not clear, but this phenomenon may allow us to turn this frequent observation into an Achilles heel in cancer cells if we carefully dissect true dependencies in well-planned genetic experiments.
2.2 CDK Inhibitors
2.2.1 Pan-CDK Inhibitors
Since CDKs are the catalytically active drivers of cell cycle progression, targeting them pharmacologically has been a major effort. The early generation inhibitors, developed more than 15 years ago were pan-inhibitors targeting a large spectrum of CDKs. These drugs were somewhat promising based upon cell line and xenograft work, but when moved into early stage clinical studies failed to show considerable net benefit. The reasons for failure are likely multi-factorial, and include both biological issues as well clinical trial design flaws. In the forthcoming section we will describe the development of several generations of inhibitors and their related trials, and provide insights into future development of these classes of compounds. Figure 14.2 shows the structures of all the CDK inhibitors discussed in this section.
The most extensively tested pan-CDK inhibitor is flavopiridol, which inhibits all four of the interphase CDKs as well as CDK7 and is also the most potent known CDK9 inhibitor [10, 11]. CDK7 is both a cell cycle and a transcriptional CDK, since it is a part of the transcription factor IIH (TFIIH) complex with cyclin H and MAT [12, 13]. CDK7 promotes transcription elongation by phosphorylating the C-terminus of RNA polymerase II. CDK9 is also thought to be involved in transcriptional regulation, in complex with cyclin T, via phosphorylating different sites in the C-terminus of RNA polymerase II [14]. There is a profound response to flavopiridol in cells, that encompasses both cell cycle arrest in G1 and G2, but also transcriptional changes especially in mRNAs with short half lives such as early response transcription factors, apoptosis regulating genes (like Mcl1) and NFκB responsive genes [11, 15, 16]. Whether these responses truly translated when this agent was tested in the clinic was not well studied.
Preclinical data had suggested that prolonged exposure to flavopiridol was necessary for maximal anti-tumor effect, so the two phase 1 trials that opened in 1994 used long infusions (72 h). The dose-limiting toxicities seen were primarily diarrhea, and at higher doses hypotension, anorexia and muscle weakness, and 21 % of patients had venous thromboses. Pharmaokinetic analysis of steady state plasma concentration revealed that 200–400nM was the range reached at the maximum tolerated dose [17]. The prior preclinical studies had found that for maximum activity, a higher concentration in the micromolar range would be desirable, so future studies attempted to reach these levels via bolus dosing on a 1 h per day for five consecutive days schedule. In these later trials, low micromolar peak concentrations were observed, and similar toxicities were observed [18]. However, when several phase 2 studies in solid tumors were analyzed, the enthusiasm for this agent waned, since no objective responses were seen in tumors ranging from melanoma to endometrial carcinoma [19–21]. In contrast, the results seen in hematopoietic malignancies appeared more promising [17]. For example, in chronic lymphocytic leukemia, 40 % of patients had partial responses, and the dose-limiting toxicity observed was tumor-lysis syndrome, indicative of strong anti-tumor activity of this agent [22].
In addition to studies using flavopiridol as a single agent, combination studies were pursued based on the hypothesis that flavopiridol may have benefit as a chemosensitizer. This hypothesis was generated based on the pre-clinical observation that synchronizing the cells into S phase sensitized them to flavopiridol-induced cytotoxicity, resulting in E2F dependent cell death that is selective to transformed cells [23]. These studies used a variety of classes of cytotoxic drugs including platinum agents, anthracyclines, taxanes and 5-fluorouracil. These studies had more promising results, including a 30–40 % rate of partial responses in some studies [24–31].
In spite of some of these promising activities in both solid tumors and leukemias, recently, there has not been significant progress with this agent. The chemistry of the agent does have some challenges, since it binds to plasma proteins and also is poorly water soluble [32, 33]. There has been a novel liposomal formulation reported a few years ago, which aimed to improve the therapeutic index by slowly releasing the drug to effectively synchronize a large portion of the tumor cell population, while not being bound up in the circulation by plasma proteins [34]. As of writing, there have not been any clinical studies presented or registered using this formulation.
2.2.2 Selective CDK4/6 Inhibitors
The G1-S checkpoint (see Fig. 14.3) is altered in close to 90 % of human tumors, by various mechanisms, indicating that this phenotype provides a selective advantage for proliferation and/or survival. With this in mind, selectively targeting CDK4 and CDK6 has been considered as an alternative strategy in several diseases including breast cancer due to the prevalence of amplification/overexpression of cyclin D1 observed (15–20 % amplification/50–70 % overexpression overall) [35, 36]. One drug candidate, PD-0332991 (see Fig. 14.2 for structure), has quickly moved to the top of its class and has rapidly moved into clinical studies [37]. This compound was selected from a high-throughput screen of pyridopyrimidines, in which both potent anti-proliferative and selective inhibition of CDK4 were used as criteria [38]. When tested against a large panel of other kinases, PD-0332291 (Pablociclib) had a highly selectivity index towards CDK4 and CDK6 (IC50’s 11/16nM, versus >8–10 μM for 36 other kinases tested). When tested in MDA-MB435 breast carcinoma cells, PD-0332991 induced a robust G1 arrest, and concomitant reduction of phosphorylation of Rb at the CDK4/6 phosphorylation sites (Ser 780–795). As expected, in Rb-negative cell lines, this compound has no activity, further demonstrating that its mechanism of action includes inhibition of CDK4/6 phosphorylation of Rb [39]. Another marker of resistance that has been identified is elevated p16 expression, since CDK4/6 is physically bound and unable to be inhibited [40].
Breast cancer is the model system which has been best studied so far in terms of understanding mechanism of action, potential synergistic combinations and predictors of resistance. At the molecular level, breast cancers can be divided into luminal or basal based on gene expression signatures. Most of the sensitive cell lines are luminal in nature, and all have intact Rb signaling, whereas the resistant cells tend to be basal-like and lack Rb activity [41, 42]. Intriguingly, basal cell lines, which retain Rb activity are still unresponsive to PD-0332991, and have hyperphosphorylated Rb. It is unclear at present what the precise mechanism is that drives hyperphosphorylation of Rb in these cell lines. It is possible that there is a greater dependence upon CDK2/CDK1, in which case these cells might respond to a combination of CDK2/1 and CDK4/6 inhibitors. Luminal tumors encompass both estrogen-receptor (ER) positive and many HER2-amplified tumors, so naturally combinations of ER antagonists or HER2 inhibiting drugs with CDK4/6 inhibitors were tested. In ER-positive cell lines, treatment with tamoxifen and PD-0332991 resulted in synergism and G1 arrest, and similarly trastuzumab and PD-0332991 are synergistic in HER2-amplified cell lines [41].
Apart from breast cancer cell lines, PD-0332991 has now been evaluated in a variety of other solid tumor types, including pancreatic neuroendocrine tumors, glioblastoma multiforme, rhabdomyosarcoma and mantle cell lymphoma with similar results [43–47]. In xenograft experiments, this drug is mostly cytostatic, with a few examples of cytotoxicity. In addition, PD-0332991 has been explored as a radiosensitizer in glioblastoma, due to its high penetrance of the blood-brain barrier, and preclinically appears to be useful in this scenario [47].
However not all tumor contexts are ideal candidates for such a strategy even if the underlying genetic changes would predict sensitivity. A recent paper described an unanticipated effect observed in pancreatic adenocarcinoma (PDAC). In PDAC cell lines examined, PD-0332991 had anti-proliferative activity, induced robust G1 arrest and hypophosphorylation of Rb [48]. However, gene expression analysis revealed that PD-0332991 upregulated genes involved in pro-angiogenic signaling, cell adhesion, cell migration/ECM remodeling, and inflammatory pathways. In addition, EMT was induced correlating with increased invasion via TGFβ-SMAD4 signaling, suggesting perhaps combinations of TGFβ inhibitors with CDK4/6 inhibitors might be a way forward in PDAC tumors expressing wild-type SMAD4. Genetic manipulation of CDK4/6 recapitulated this phenotype, ruling out a drug-mediated off-target kinase inhibition.
The first phase 1 study performed using PD-0332991 was recently published, and examined patients with Rb-positive advanced cancers [49]. This study showed that the drug was generally well tolerated, with the main toxicity being myelosuppression, consistent with other cell cycle targeted therapies. Pharmacokinetic analysis suggested favorable properties including slow absorption and elimination. The response rate was moderate (~27 %), however given all the usual caveats of generalizing based on phase 1 studies, the patients who derived some benefit (i.e. stable disease) could tolerate the drug well enough to remain on study for 10+ cycles. In breast cancer patients, a randomized phase I-II study utilizing PD-0332991 in combination with letrozole in ER-positive, HER2-normal post-menopausal patients has been completed (personal communication). In the phase 1 portion, there has been no biomarker selection, but in the phase II portion, the trial is specifically focused on patients with cyclin D1 amplification and/or loss of p16, since these are the patients predicted to respond best. So far, the clinical benefit rate in this combination trial was 70%, which resulted in statistically significant increase in progression-free survival and the adverse event profiles are very similar to what was reported in the single-agent phase 1 studies trials previously discussed i.e. this combination is generally well tolerated. The few patients so far in the study have been safely treated with some patients having partial responses. These well-designed trials with integrated biomarkers built in, are likely to provide more useful information not only about safety and pharmacokinetics but also pharmacogenomics information about responders and the biology behind responses. Caution must be taken though in considering combinations with chemotherapies that depend on actively cycling cells, based on a recent study that showed that PD-0332991 actually protected RB-positive MDA-MB-231 cells from paclitaxel-induced mitotic catastrophe [50]. Pre-treatment with PD-0322991 also resulted in a switch from homologous recombination (HR) DNA repair mechanism to non-homologous end-joining (NHEJ), which is an error prone pathway that could potentially induce further genomic instability in cancer cells. While this finding is intriguing and worthy of further mechanistic study in a broader panel of cell lines and normal mammary epithelial cells, it still remains to be determined how this result will be translated into ER+/HER2+ cell lines, which have been the focus of the majority of the breast cancer studies using this agent.
Going forward in hormone-receptor positive breast cancer, targeting CDK4/6 should eventually become a first line therapy in combination with endocrine therapy for a number of reasons. Certainly, as described above, the preclinical data regarding this combination is compelling, and this includes cell lines that acquired tamoxifen resistance being re-sensitized by PD-0332991. Secondly, genomic data have confirmed the relevance of this pathway in resistance to endocrine therapy alone, such as the fact that cyclin D1 is overexpressed/amplified in endocrine-therapy resistant tumors [51, 52]. In addition to deregulating the cell cycle, amplified cyclin D can directly activate ER in a hormonally-independent manner that does not require CDK/Rb activity [53]. Thirdly, the fact that cyclin D1-CDK4 is downstream of multiple pathways that mediate resistance to anti-estrogens (e.g. EGFR/HER2, ERK, AKT, NFκB) may make this strategy useful regardless of which pathways are upregulated in any particular patient [54]. Despite being a good target however, resistance to CDK4/6 inhibition is likely to occur, since resistance arises to every targeted therapy tested so far. Indeed, there is evidence currently for activation of CDK2 due to p27 down regulation as a mechanism of resistance to these agents, which could potentially be targeted via CDK2 inhibitors as will be discussed in the next section [55].
Further clinical studies in other diseases are also underway, for example in mantle cell lymphoma. The single agent study showed some evidence of benefit, and now a subsequent study has been designed using PD-0332991 in combination with bortezomib in this patient subset.
2.2.3 Selective CDK1/2 Inhibitors
The other subclass of CDK inhibitors that have been developed are more specific for CDK1 and CDK2 (versus CDK4/6), such as R-roscovitine (also known as CYC202 or seliciclib), SNS-032 and the newer agent SCH727965 (see Fig. 14.2 for the structures). Roscovitine, a 2,6,9-trisubstituted purine was generated in a screen of olomoucine-related analogues for CDK1/cyclin B inhibition, and found to potently inhibit CDK1 kinase activity (IC50 of 0.45 μM) [56]. Once olomoucine was shown to co-crystalize with CDK2, roscovitine was also confirmed to bind directly to CDK2 [57]. Several years later, roscovitine became the first orally bioavailable drug from this class to go into clinical trials based on the preclinical data showing that CDK1/2 inhibition causes S and G2 arrest followed by apoptosis in tumor cells [58–60]. Apart from the effects on these cell cycle CDKs, roscovitine also inhibits CDK7 and CDK9, thereby inhibiting transcription as well, via reducing key anti-apoptotic proteins such as Mcl1 [61–64].
Several studies have shown that apoptosis is further induced when CDK1/2 inhibitors are combined with most cytotoxic therapies including taxanes, anthracyclines as well as radiation. For example, a combination of purvalanol A and taxol caused profound apoptosis in Hela cells, when taxol was used first to stabilize microtubules then purvalanol A was added [60]. Intriguingly when the drugs were used in the reverse order, the response was decreased, demonstrating that synchronization of cells in mitosis (i.e. the end results of taxol) is important for the mechanism of CDK inhibitor-induced cell death. A similar synergistic combination strategy was demonstrated in MCF7 breast cancer xenografts using roscovitine and doxorubicin, however in this context, cell cycle synchronization in G2/M phase with roscovitine was used to prime the cells to respond to doxorubicin (versus the taxol→CDKi strategy in Hela cells) [65, 66]. Similar to the Hela cell study described above, the taxol-purvalanol A combination was found to be similarly effective in MCF7 xenografts [67]. These dichotomous results in two different systems just illustrate one of the challenges we have moving forward with sequential combination therapies that exploit mechanism of action of drugs. Likely a number of factors could contribute to which direction of treatment is likely to be best, including genomic factors (such as Rb status, p53 pathway status), timing of exposure to agents as well as which specific drugs under investigation. Clearly, further mechanistic work is still needed to dissect out these details in order to rationally match treatments to individual patients.
In spite of this incomplete understanding of mechanism of action of these agents in both solid tumors and hematopoietic malignancies, roscovitine was moved into clinical trials in the early 2000s [68]. The phase 1 studies demonstrated that this agent could be administered both in an intravenous formulation as well as orally [69, 70]. It has reasonable pharmacokinetic properties including high bioavailability, slow GI absorption, however it is rapidly metabolized to an inactive metabolite, making its half-life fairly short (~1 h). However, there were a number of dose-limiting side effects observed including liver and kidney toxicity, electrolyte disturbances, rashes and fatigue that accumulated over time, making repeated administration daily for more than 5 days too challenging for patients. Responses were unimpressive over a few phase 1 studies, with primarily stable disease and very few partial responses seen as monotherapy. Two phase 2 studies were undertaken in non-small cell lung cancer and nasopharngeal carcinoma which essentially replicated the results in the phase 1 studies [71]. Looking at the pharmacological and response data together, the researchers concluded that one of the major challenges was maintaining a plasma dose that is high enough for anti-tumor activity based on the preclinical studies, and even when white blood cells were used as a surrogate for tumor cells, Rb phosphorylation was not decreased, further supporting the claim of insufficient dose reaching tumor cells. One way of potentially overcoming this problem might be to use a more frequent dosing schedule such as 2–3 times a day, since the preclinical studies showed that 8–16 h of continued exposure is needed to effectively inhibit tumor growth. Whether this would actually work might not be known since the excitement regarding this agent has waned, in light of newer compounds that have been developed. One such potential compound is CR8, which is a N6-biaryl-substituted derivative of roscovitine that is 2–4 fold more potent at inhibiting CDK1, CDK2 and CDK7, which translated to 40–70 fold higher potency in cellular activity measures such as PARP cleavage and caspase activation [72–74]. Since discovery of this compound a few years ago, animal studies have not been published in cancer models, although a very recent paper utilizing CR8 in a mouse model of traumatic brain injury demonstrated that this drug could be delivered safely in vivo.
The other CDK inhibitor that targets CDK1 and CDK2 (as well as CDK5 and CDK9) that appears promising is SCH727965 (Dinaciclib) (see Fig. 14.2 for structure), a compound developed to address some of the issues with previous generation inhibitors with respect to therapeutic index [75]. Indeed, in direct comparison to flavopiridol this agent had > tenfold greater therapeutic index in the A2780 ovarian cancer xenograft model (defined as the ratio between MTD defined as 20 % body weight loss and minimal effective dose to cause 50 % inhibition of tumor growth when given i.p. once daily for 7 days). In addition, cell line studies showed that even a brief 2 h exposure to SCH727965 was sufficient to inhibit progression of cells into S phase. In vivo, this drug was at least as effective as paclitaxel in A2780 xenografts, and well tolerated with the main toxicity being reversible myelosuppression.
In addition to ovarian cancer, SCH727965 has been tested and found to be potentially effective in pancreatic cancer, melanoma and various forms of sarcoma including osteosarcoma [76–79]. The pancreatic study was particularly exciting, as it was performed using low-passage patient-derived xenografts (PDX) as opposed to cell line xenografts. These PDX models are thought to more faithfully recapitulate human tumorigenesis for multiple reasons including the fact that they maintain human stroma for multiple passages [80]. This intense desmoplastic stroma and hypovascular microenvironment which characterizes pancreatic cancer, is known to be a major barrier to chemotherapy drug access. Therefore the data showing efficacy in multiple mouse models with these characteristics bodes well for subsequent trials in humans. In addition to using these better disease models, the authors performed gene set enrichment analysis on the tumors to interrogate potential mechanisms of resistance, an area of research that is very undeveloped in the cell cycle field. They compared sensitive and resistant tumors, and found that in the most resistant tumors, the Notch and TGFβ pathways were upregulated, suggesting that perhaps combinations of these inhibitors may be future directions for research.
Preclinical studies in adult and pediatric leukemia are also underway, and a report of efficacy in CLL cells showed promise independent of high-risk genomic features (del 17p13.1 and gVHI unmutated). Short-term exposure of CLL cells directly taken from patients was sufficient to induce apoptosis [81]. Moreover, SCH727965 was shown to abrogate microenvironment-derived cytokine-induced survival signaling, in a PI3K-dependent mechanism, suggesting that a logical combination to explore might be SCH727965 in combination with PI3K inhibitors.
These pre-clinical studies provided strong rationale for moving this agent into clinical studies. A phase 1 study using SCH727965 dosed once every 3 weeks as a single agent in unselected adult patients has been performed which showed moderate responses (mainly stable disease) and similar to the preclinical studies, myelosuppression was the DLT [82]. Notably, unlike flavopiridol, there was no diarrhea, and much less fatigue, making this agent better tolerated. Since nausea and vomiting were common side effects as well, a subset of patients was given the anti-emetic drug aprepitant [83]. Aprepitant is known to weakly inhibit CYP3A4 which is one of the enzymes involved in metabolizing SCH727965, so the pharmacokinetic parameters were compared in patients treated with aprepitant with those not given it. These results showed no interaction between aprepitant and SCH727965, suggesting that use of this agent prophylactically in the clinic is a feasible and safe strategy moving forward to maximize use of SCH727965 in different patient populations.
Several phase 2 studies with SCH727965 as a single agent have also begun in both solid tumors and hematopoietic diseases, and some have been presented as abstracts at meetings. The first and most promising was performed in adult acute myeloid or lymphocytic leukemia patients and reported at ASH in 2010 [84]. The response rate was 60 %, and many patients had rapid decrease in their blast counts. Correlative studies including pharmacodynamics analyses showed that CDK activity (i.e. Rb phosphorylation and Mcl1 decreased expression) was effectively inhibited in the samples taken at 4 h post infusion, however these biomarkers returned almost to baseline by 24 h, suggesting a need for frequent dosing. A similar study is also underway in multiple myeloma, but no results are currently available [85].
With respect to solid tumors being examined in single-agent phase 2 trials, the progress has been slower. One single arm study has been reported in melanoma, which had ~72 patients enrolled [86]. The response profiles were very modest, with 22 % of patients with stable disease and no partial or complete responses, and toxicities were common. Another phase 1–2 study in unresectable melanoma is now approved and about to open. The only other trial that has been started is a multi-arm randomized phase 2 in breast cancer and NSCLC, in which SCH727965 is being compared to active treatments for each respective disease (oral capecitabine for breast cancer, and erlotinib for NSCLC). Importantly crossover from the control arm was allowed after disease progression, which is likely to make detection of a significant difference in overall survival extremely challenging. The study has been completed but no data has been presented as of writing (Nov 2012) [87].
Similar to the other CDK inhibitors, combinations with chemotherapy/other targeted agents are ultimately going to be necessary for optimal activity of SCH727965, and already a number of combination studies have been started. These trials include combinations with the PARP inhibitor Veliparib with or without carboplatin, rituximab in CLL, bortezomib and dexamethasone in myeloma, and our own trial with epirubicin in triple negative breast cancer [88–91]. Our trial differs from the others in that by limiting our patients to a specific subtype of breast cancer in which we have preclinical data showing dependence on CDK2 signaling because of LMW-E expression [92]. In addition, another group has shown elevated c-Myc expression is synthetically lethal with CDK inhibition in triple negative breast cancer [93]. By pre-selecting patients with a high likelihood of being oncogenically addicted to CDKs either via the LMW-E pathway or via amplification of c-Myc, we believe we will enrich for potential responders. Further biomarkers of response or markers predicting resistance will be needed in order to make this drug clinically useful. For example, basal like breast cancers or serous ovarian cancers with defects in DNA repair such as BRCA1/2 or ATM mutations may be more sensitive to CDK2 inhibition (in combination with chemotherapy), analogous to their propensity for sensitivity to PARP inhibition. BRCA1 or ATM knockdown sensitized various cell lines to CDK2 inhibitors, since CDK2 can regulate DNA repair independently of its effect on the cell cycle [94].
2.3 Targeting p27
Moving away from directly targeting components of the cell cycle that promote growth, the other strategy that has been proposed is targeting negative regulators such as p27. p27 is a small nuclear protein involved in negatively regulating G1 to S phase transition via inhibiting cyclin E-CDK2, cyclin A-CDK2 and cyclin D-CDK4/6 activity [95]. However, in addition, it has novel functions when mislocalized in the cytoplasm that contribute to tumor cell survival and cancer progression [95]. For example, in response to stress, p27 is driven into the cytoplasm via AMPK-mediated phosphorylation at Thr198 which blocks apoptosis and induces a cytoprotective autophagy response [96, 97]. Other cytoplasmic functions also include increased invasion and metastatic potential, perhaps via binding to RhoA and stimulating changes in actin cytoskeleton formation [98]. A p27 knock-in mouse model where p27 has been mutated to be unable to bind cyclins and CDK2, was generated to determine whether there are cell-cycle independent roles for p27 [99]. One of the major findings from this model includes a role for cytoplasmic p27 in stem/progenitor cell enrichment, leading to lung tumorigenesis [100]. Taken together these results clearly show that p27 has pleiotropic functions and therefore merely upregulating it without understanding cellular context may not be very effective.
In many cancers p27 levels are decreased as a result of deregulation of transcriptional, translational or post-translational pathways. Much effort has been expended into understanding these mechanisms with the end goal being to determine ways of upregulating nuclear p27 to prevent cell cycle progression. To briefly summarize these studies, which have been reviewed extensively elsewhere, p27 transcriptional regulation is complex, and involves both repression via oncogenic transcription factors such as c-Myc and Id3, as well as activation via FOXO transcription factors and E2F1 [101]. In addition to transcriptional regulation, p27 has been shown to be regulated via miRNAs such as miR-221/222, which are overexpressed in some tumors [102]. p27 is also extensively regulated at a post-translational level, including multiple phosphorylation sites that dictate localization, as well as that phosphorylation sites that stabilize the protein via inhibiting ubiquitination. Some of these most important sites include Ser10, Tyr74/88/89, Thr157, Thr187, and Thr198.
Beyond phosphorylation, p27 localization can also be regulated via protein-protein interactions, such as Jab1, which binds to p27 and induces its nuclear export and degradation in the cytoplasm [103]. Jab1 is an interesting potential target for a number of reasons. Jab1 expression in tumors is inversely correlated with p27 levels and overexpression is correlated with poor prognosis [104–107]. This proto-oncogene has also been linked to radiation resistance and cisplatin-resistance due to inhibiting several apoptotic and DNA repair pathways [108]. Apart from p27, Jab1 can induce degradation of other tumor suppressors such as Smad4 and p53 as well as inhibit DNA repair via HR pathways that involve Rad51 [109–111].
p27 proteolysis via the proteasome is regulated by Skp2 [112]. Skp2 is part of a larger SCF complex, comprised of cullin1, Skp1, ROC1/Rbx1 and requires an adaptor protein called Cks1. Together, this complex functions as an E3-ligase that regulates a number of substrates including p27, p21, p57, FOXO1 and c-Myc [113–116]. Towards the end of G1 phase, Skp2 recognizes p27 phosphorylated at Thr187, which serves as a degradation signal, allowing cells to enter S phase. However, in cancer cells, Skp2 can be amplified and/or overexpressed, leading to decreased p27 levels constitutively [117–119]. As a result of this reciprocal relationship, the concept of targeting Skp2 has been proposed, however these studies have not progressed well. Genetic studies using a Skp2 deficient mouse model have demonstrated that acute inactivation of Skp2 in the context of Pten or Arf heterozygosity induced senescence but not in the Skp2-deficient mice without other oncogenic signals [120]. Senescence induction correlated with decreased tumorigenesis and p27 induction in the preneoplastic lesions that could be detected in some mice, providing additional rationale for the development of Skp2 inhibitors.
A high-throughput screen for inhibitors of Skp2 was performed using purified components of the complex and p27 as a substrate [121]. A compound designated CpdA was discovered to induce cell cycle arrest at low micromolar doses in multiple myeloma cells, and this led to caspase-independent cell death via autophagy. In addition, because low levels of p27 is associated with resistance to therapy in myeloma cells, CpdA was examined as a chemosensitizer, and was found to synergize with bortezomib, and overcome resistance to doxorubicin and melphalan. Despite this useful spectrum of activity however, further progress has not been made using this agent since it is difficult to make, and was not potent enough to use in vivo (required 5–10 μM dose in cell lines). More recently, another screen was performed using a chemical-genetics approach using automated microscopy to identify compounds that upregulate p27 in prostate cancer cell lines [122]. After several rounds of stringent validation, two compounds were identified – SMIP001 and SMIP004. These compounds are not broad proteasome inhibitors, but specifically target Skp2, resulting in elevated p27 and decreased CDK2 activity at low micromolar doses. At this point is it not known whether either of these compounds will be effective in vivo.
Targeting p27 presents a considerable challenge, despite the large body of knowledge regarding the mechanisms of its regulation and their redundancy. Some open questions include which of the upstream regulatory enzymes would be the best target to induce a sustained increase in p27 cells in tumor cells specifically. In some ways, the lack of complete specificity of substrates presents the largest conundrum. Assuming it would be possible to move one or more of the identified compounds (or a derivative) into the clinic, it is possible that some of the off-target effects may be undesirable e.g. in some contexts the upregulation of cyclin E or c-Myc may drive additional genomic instability. In fact there is data suggesting this might be the case using siRNA in A549 lung cancer cells targeting p27, Skp2 or the combination of both mRNAs [123]. In the dual-siRNA treated cells, there was an increase in centrosome number, abnormal mitoses/nuclear atypia, which could be attributed to an increase in both full length and low-molecular weight forms of cyclin E. In tumors, such as triple negative breast cancers, which already have LMW-E expression we propose that a Skp2-targeting strategy could be detrimental because of this off-target effect, unless combined with CDK inhibitors.
The other concern with Skp2 targeting relates to the specific genetic background of p27. As mentioned previously, p27 transcriptional silencing such as by miRNAs or promoter methylation are not uncommon events, and could co-exist with Skp2 overexpression. In this scenario even a potent Skp2 inhibitor would be ineffective at restoring p27 expression, with similar potential consequences as described above. In addition, because cyclin D-CDK complexes also bind p27, the level of p27 induction that may be necessary to slow down the cell cycle might be fairly high. Clearly, much work lies ahead in developing more potent inhibitors and understanding in greater depth the cellular contexts in which this strategy could be beneficial and which contexts to avoid.
3 Exploiting Normal and Cancer Cell Differences for Protection of Normal Tissue
One of the most challenging problems in cancer therapy involving cytotoxic chemotherapy is how to selectively kill tumor cells while sparing normal dividing cells. Previous work from our group has demonstrated two potential strategies that utilize cell cycle synchronization as a mechanism of selectively arresting normal cells. We proposed that using UCN-01, a staurosporine analog which was developed as a PKC inhibitor, but was later shown to inhibit other kinases including CDK1, CDK2 and CDK4 at low nanomolar concentrations could be used in this manner. UCN-01 has been found to induce a reversible G1 arrest in normal cells, while Rb-deficient tumor cells arrest in S phase instead [124]. Cytostatic doses of staurosporine (0.5–10nM) can also be used to arrest normal cells in G1 without any detectable effect on tumor cells [125]. Importantly, staurosporine priming does not compromise the ability for tumor cells to respond to cytotoxic therapies, while normal cells are arrested in G1 and therefore not responsive to chemotherapy that targets cycling cells.
More recently with the availability of specific CDK4/6 inhibitors that are more clinically relevant than staurosporine, this hypothesis has been revived and tested in mouse models using PD-0332991 [126]. Myelosuppression induced by platinum drugs and anthracyclines are one of the most life-threatening toxicities seen in cancer patients, due to both heightened risk of infection while immunosuppressed and also due to the subsequent chemotherapy delays or dose reduction, which can compromise treatment efficacy. A study was performed comparing two different mouse models of breast cancer, one with intact Rb signaling (the MMTV-neu model) and the other with inactive Rb (C3-Tag), to examine whether CDK4/6 inhibition could protect the hematopoietic progenitor cells from carboplatin-induced quiescence. The C3-Tag model which best resembles basal-like breast cancers, is Rb-deficient and, unsurprisingly, does not respond to CDK4/6 inhibition as a single agent. Carboplatin is highly active in basal-like breast cancer, though, and co-treatment with PD-0332991 did not protect tumor cells from death. However, these mice had reduced thrombocytopenia. In contrast, MMTV-neu mice, which have previously been shown to be dependent upon CDK4 and cyclin D, were sensitive to PD-0332991 as a single agent. In addition, when carboplatin was combined with PD-0332291, tumors grew back faster, indicating the CDK4/6 inhibition in this context protects tumor cells as well normal cells from toxicity, therefore not gaining any significant therapeutic index. These results suggest that CDK4/6 inhibitors may have a new utility – in tumors that are CDK4/6 sensitive, these drugs can be used for anti-tumor effect (and should be used separately from other cytotoxic therapies), and in other tumors that are insensitive (e.g. Rb deficient, p16 overexpressed), these drugs can be normal tissue protectors from other cytotoxic therapies. As a practical consideration as a result of these discoveries, we advocate for Rb mutation status to become one of the biomarkers tested routinely in the clinic.
4 Newer Cell Cycle-Related Targets for Therapy
In this last section, we briefly outline rationale for targeting other cell cycle-related proteins and review the state of drug development for each class of agent. Some of these proteins are intimately involved in mitosis regulation, and will be discussed separately.
4.1 PIM Family Kinases
One of the most interesting emerging targets in the PIM family of kinases, which regulate multiple pathways including the cell cycle. The PIM family of serine/threonine kinases consists of three isoforms that have a significant degree of sequence homology, but differ in their tissue distribution [127]. Overall these proteins are expressed throughout in hematopoietic progenitors as well as liver, spleen and other epithelial and mesenchymal tissues, and have considerable functional redundancy. PIM1 is the best studied member of this family, and is thought to be the most widely relevant gene (of the 3 PIM isoforms) in cancer.
The PIM1 gene was identified in the 1980s as a frequent proviral integration site for Moloney murine-leukemia virus (MuLV) which induced T-cell lymphomas in transgenic mice [128]. Subsequently PIM1 was shown to cooperate with c-Myc in inducing lymphomas in utero or around birth, whereas Eμ-Myc transgenic mouse crossed onto a PIM1 and PIM2-deficient background had delayed lymphomagenesis [129–131]. More recently, PIM family kinases have been discovered to be overexpressed or mutated in other solid tumors such as prostate, pancreatic, ER-negative breast cancer and head and neck squamous carcinomas as well as many leukemias and lymphomas (AML and CLL), leading to the question of whether they could be targeted [132–137].
PIM kinases are unusual in that they are constitutively active, but are regulated largely at the transcriptional and translational level [127–138]. A wide range of cytokines and growth factors can activate PIM kinases, mainly via the JAK-STAT pathway and NFκB pathways [139, 140]. Since the mRNA has a short half-life, inhibitors of JAK-STAT could potentially be used to inhibit PIM signaling as well. An emerging paradigm places PIM1 at the center of a cellular stress response, since PIM1 can be induced by hypoxia and DNA damage via various mechanisms. For example hypoxia can induce PIM1 expression rapidly in a HIF1α independent mechanism, as well as induce nuclear translocation [141, 142]. PIM1 induction in response to hypoxia promotes cell survival via inhibition of apoptosis and is linked to chemoresistance under these conditions. PIM1 can also be induced in response to DNA damage by Kruppel-like factor 5 [143]. In a study of head and neck squamous carcinoma patients, upregulation of PIM1 in response to irradiation was shown to be associated with a poor response [144]. Since EGFR expression is also correlated with radiation resistance and EGFR is autophosphorylated and nuclear localized after IR, the authors asked with EGFR can regulate PIM1 levels/activity. In cell lines, EGF-ligands induced PIM1 nuclear-translocation, and this effect could be blocked by the EGFR antibody cetuximab or tyrosine kinase inhibitor, gefitinib. Similarly, HNSCC cells that were irradiated had more nuclear PIM1, and PIM1 knockdown demonstrated the pro-survival role that PIM1 plays in this context. Taken together, these studies show that PIM1 may be a good target in a number of different cancer systems.
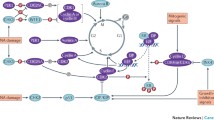
PIM1 plays a number of cellular functions that all contribute to tumorigenesis, as depicted in Fig. 14.4. One of these is regulation of the cell cycle via phosphorylating several substrates such as p21, p27, cdc25A, cdc25C and HP1 [145–151]. One of the most robust readouts of PIM1 activity is phosphorylation of the pro-apoptotic protein BAD at Ser 112, which inactivates it, therefore enhancing anti-apoptotic activity of Bcl2 [152]. As mentioned previously, PIM1/PIM2 cooperates with c-Myc to regulate lymphomagenesis, and one of the mechanisms it does so is via phosphorylation of c-Myc which stabilizes the protein [153]. Overexpression of PIM1 also induces genomic instability via deregulating the mitotic spindle checkpoint, which causes abnormal mitoses, centrosome amplification and aneuploidy [154]. In hematopoietic malignancies in which PIM2 is highly expressed, 4EBP1 is also a target that is involved in promoting cap-dependent translation initiation of proteins that have growth promoting roles such as c-Myc and cyclin D1 [136]. In prostate cancer specifically, the androgen receptor is also a substrate of PIM1, and this phosphorylated form is transcriptionally inactivated and degraded [155, 156].
Structurally PIM kinases are distinct from other kinases in terms of how ATP binds to them, which has allowed chemists to design highly selective inhibitors. One of the most attractive features of PIM1 as a drug target is the lack of obvious phenotype in the knockout mouse, which is viable and fertile [157]. Compensation by other PIM family members is unlikely since compound knockout mice are also viable and fertile. The only phenotype that was observed in the Pim1-/- mouse is a subtle hematopoietic effect, such that the red blood cells are abnormally small but this did not lead to any physiological effects. When other potential hematopoietic functions were examined closely, it was found that bone-marrow-derived cells in culture had a significant impairment in IL-3 and IL-7 growth factor response [131].
The first compound that has been developed that has moved into cellular and in vivo studies is SGI-1776, which is an imidazo [1, 2-b] pyridazine compound that inhibits all three PIM kinases with IC50s of 7, 363 and 69nM (PIM1, PIM2 and PIM3 respectively), and has some activity against FLT3, another target in AML (see Fig. 14.5 for structure of this and other PIM inhibitors) [158]. In xenograft models of AML cells, this drug was highly active as an oral agent, inducing complete regression of blasts [159]. In addition, SGI-1776 can re-sensitize chemoresistant prostate cells to taxanes due to inhibiting multidrug resistance proteins including MDR1 [160]. Unfortunately when moved into phase 1 trials in humans (one trial was focused on prostate cancer and the other was non-Hodgkin’s lymphoma) this drug was found to cause dose-limiting cardiac toxicity for reasons that are not clear, and the studies were stopped [161]. Another two structurally-related PIM1 inhibitors were identified in a chemical library screen called Smi-4a and Smi-16a which are benzylidene-thiazolidine-2, 4-diones [162]. When tested in vitro, these agents both had growth inhibitory activity in leukemia and prostate cancer cell lines, induced G1 arrest and induced p27 nuclear translocation. In addition, PIM1 inhibitors including Smi-4a synergize with both rapamycin in prostate cancer cells, and more recently with the Bcl2 inhibitor ABT-737 [163]. In our opinion despite these compounds having promising pre-clinical activity, the drug discovery market relating to PIM1 kinase is wide open right now. Special attention should be focused on the potential cardiac toxicity profile of future inhibitors (compared to SGI-1776), to try to understand what the off-target mechanisms are that underlie the Qt prolongation seen with SGI-1776 in patients.
4.2 Mitotic Kinases: Aurora Kinase Family
In much of this chapter so far we have discussed targets that function early in the cell cycle in regulating G1 and S phases. However, G2 and M phases are also very kinase-rich and have tremendous potential as drug targets. Several classes of chemotherapies already target these processes, such as taxanes, which bind to tubulin and disrupts the assembly of the spindle. In order to design better therapies against proteins that act in G2 and M phase, we must understand their functions at a mechanistic level and how they contribute to the events that are necessary for progression through these stages. During G2 phase when cells are preparing for mitosis, the cell is very active in ensuring the DNA was replicated correctly, and dividing the other organelles. In addition, microtubule proteins are being synthesized in order to form the mitotic spindle along which the chromosomes will segregate during mitosis. G2-M phase targets include proteins that are involved in entering mitosis (such as Aurora kinase A), the spindle assembly checkpoint (such as BUB1), and mitotic exit (such as APC). Each of these proteins and processes could be the focus of entire chapters, so we will provide a high-level overview of each here, and point the readers to recent reviews on these proteins.
Aurora kinases, of which there are three highly related isoforms (A, B and C) in mammalian cells, are key regulators of mitosis that have been well conserved throughout eukaryotic organisms. All three isoforms have been the focus of drug development over the past several years, and most of the inhibitors target two or three of them due to the highly conserved catalytic domain [164]. Aurora kinase A (AURKA) and Aurora kinase B (AURKB) have the strongest evidence for a role in tumor cell growth, whereas Aurora kinase C has scant evidence. This may be because for many years Aurora kinase C was thought to be primarily expressed in the testes where it plays a role in spermatogenesis, by playing similar roles to AURKB [165]. However more recently it has also been found to be expressed at high levels in some cancer lines, and several point mutations have been found in lung tumors but relatively little is known about its function [166, 167].
AURKA is ubiquitously expressed and is the first of the family members to be activated starting in late S phase and working through to completion of mitosis. Many related processes are regulated by AURKA, including maturation and separation of the centrosomes, assembling the mitotic spindle, chromosome alignment and cytokinesis [168–170]. Regulation of AURKA levels is also important for mitotic exit, as either too much or too little activity leads to failure of cytokinesis and multinucleation [171]. AURKA regulation occurs at both transcriptional and posttranslational levels, including activation by autophosphorylation at Thr288 on the activation loop, and deactivation via protein phosphatase 1 [172]. In cancer, AURKA is frequently amplified/overexpressed due to various mechanisms, especially in higher grade tumors and has been demonstrated to be a poor prognostic factor [173–176]. It was established as a bona fide oncogene when it was shown to be capable of inducing rodent fibroblast cell transformation due to formation of multipolar mitotic spindles that induce genomic instability [177]. Interestingly, these chromosomal abnormalities that occur in AURKA overexpressing cells does not lead to cell death, because AURKA also promotes cell survival pathways including AKT-mTOR and nuclear accumulation of cyclin D1 [178]. NFκB is another anti-apoptotic pathway that is regulated by AURKA phosphorylation of its inhibitor, IκB [179]. AURKA has also been shown to interact with the p53 network, specifically via phosphorylating p53 inducing its degradation via MDM2, as well as phosphorylating p53 on Ser 215 which inhibits its DNA binding ability [180, 181]. These findings demonstrate the wide spectrum of roles that AURKA plays in cellular transformation and provide significant rationale for targeting this kinase.
Similar to Aurora kinase A, Aurora kinase B (AURKB) is also expressed in all proliferating cells, however it plays more limited roles as a chromosome passenger protein. AURKB is primarily expressed starting during prophase, where it is localized at the kinetochore to ensure correct chromosome alignment to the spindle and also helps ensure chromosomes segregate correctly [182]. In addition, AURKB phosphorylates Histone H3 at Ser10 and Ser28, which facilitates chromosome condensation [183, 184]. In cancer AURKB is not amplified, however it is still highly expressed in several tumor types [185–187]. Apart from regulating kinetochore-spindle interactions, in cancer cells, AURKB has been linked to degradation of p53 via phosphorylation at multiple sites [188], providing further rationale for targeting this kinase.
Inhibition of AURKA leads to G2 arrest, and has been shown to increase chemo- and radiosensitivity in cancer cells [189]. Quite a number of inhibitors have been generated by most of the major pharmaceutical companies and are currently being tested in early stage clinical trials. Many of these target both AURKA and AURKB, although Millennium has two AURKA specific compounds, MLN8054 and MLN8237. For a recent review with information about the clinical development of these agents, see [169]. Inhibition of AURKB are known as mitotic drivers, since they cause overriding of the mitotic checkpoints and results in aberrant mitosis and aneuploidy. This contrasts with AURKA inhibitors which block passage through mitosis. The question of whether inhibiting both AURKA and AURKB is better than either kinase alone has still not been answered. In preclinical genetic studies, the results have been equivocal. In one study in pancreatic cancer, antisense oligonucleotides to AURKA, AURKB or the combination were added to cells and responses compared [190]. The combination of both oligonucleotides was not better at inducing caspase activation, accumulating tetraploid cells or reducing formation in soft agar than either one alone. Targeting AURKA alone had a slightly better response overall versus AURKB alone, and this correlated with cells rounding up and detaching from the plate versus becoming large and multinucleated with the AURKB oligonucleotide. In contrast to this pancreatic study however, in colon cancer cells, AURKB inhibition was better than AURKA [191]. In order to move these targets forward, greater emphasis will have to be placed on understanding what contexts predict response to inhibition of each protein, and multiple readouts of each kinase inhibition should be analyzed since it is possible that each drug will have a slightly different profile. One clue that has already emerged is that p53-deficient cells more readily undergo apoptosis in response to the VX-680 inhibitor, however since this is a pan-aurora inhibitor, it is difficult to dissect out which target is most relevant in p53-deficient tumors [192]. Further molecular and pharmacodynamic characterization of sensitive and resistant patients in the many clinical trials in progress should elucidate more such factors, as well as more detailed preclinical work with patient derived xenograft models should be the in vivo assay of choice in these studies.
4.3 Other Mitotic Targets of Interest
Moving forward as genomic studies are completed and more functional screens are performed it is likely more novel cell cycle targets will be found. Some examples of this nature that have been identified so far include MELK, Bub1, and Mps1. We will briefly summarize some of these proteins and how targeting them might be useful in cancer.
Maternal embryonic leucine zipper (MELK) is an atypical member of Snf1/AMPK family of kinases that has received only a little research attention so far. MELK is upregulated in several solid tumors including high-grade prostate cancer, astrocytoma, medulloblastoma and in breast cancer [193–195]. In addition, MELK is highly expressed in neural and breast cancer stem cells, making it a potentially attractive target to eradicate this population of cells thought to be the main drivers of drug resistance and eventual disease progression [196–198]. Expression of MELK is known to be increased in mitotically-arrested cells, and in prostate cancer cells is highly correlated with several other cell cycle/proliferation related genes including AURKB, cyclin B2 and DNA topoisomerase 2 alpha [193, 199, 200]. A few recent studies have suggested a role for MELK in radioresistance and chemoresistance, and have provided some in vitro evidence that knockdown can sensitize cancer cells to additional therapies [201, 202]. The only known pharmacological agent that targets MELK so far is the antibiotic siomycin A which reduces MELK expression and has been shown to decrease glioblastoma growth in vivo via targeting the neural stem cells [203].
The spindle assembly checkpoint (SAC) is a mechanism of delaying anaphase if the kinetochores are unattached to microtubules. There are at least 14 proteins involved in this process, four of which are kinases that are potentially targetable. These kinases are Bub1, BubR1, Mps1 and aurora B, although aurora B is dispensable for the checkpoint. If the SAC checkpoint is active, some of the components, such as Bub1, sequester Cdc20 which is the active part of the APC/C complex which degrades cyclin B. Bub1 may be a master regulator of the spindle assembly checkpoint by recruiting other important proteins involved such as BubR1, Mad1 and Mad2 [204]. The underlying concept behind targeting this checkpoint is that by preventing SAC activation, severe chromosome segregation occurs occur which causes cell death. Even partial inhibition of any of these essential mitotic checkpoint components can sensitize tumor cells to mitotic-targeting chemotherapies such as taxanes, whereas normal cells are not sensitized since normal cells can maintain a diploid population of cells [205]. Proof of this principle in vitro has been obtained for inhibitors of Mps1 [206, 207].
Cdc20 has also been proposed as a target in cancer due to the strong phenotypes seen in genetic studies from blocking mitotic exit. The cdc20 homozygous knockout mouse is embryonic lethal at the two-cell stage due to a metaphase arrest [208]. When an inducible knockout model was generated, a similar phenotype could be observed upon induction, and very high levels of cyclin B was observed in the cell, consistent with a defect in APC/C function [209]. When tumors of either epithelial or mesenchymal origin were induced in this model, and then cdc20 knockout was induced, the tumors rapidly regressed due to mitotic arrest and apoptosis.
In summary, there are various strategies that are being investigated to interfere with mitosis including delaying mitotic entry and spindle formation, preventing activation of the spindle assembly checkpoint or targeting mitotic exit via the APC/C-cdc20 complex. Such strategies may synergize with current chemotherapies that act in mitosis such as taxanes and vinca alkaloids, allowing lower doses of these agents to be administered. The question remains however whether a sufficient therapeutic index can be reached since normal cells also require these processes to be intact to undergo normal mitosis.
References
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70, PubMed PMID: 10647931. eng
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674, PubMed PMID: 21376230. eng
Berthet C, Aleem E, Coppola V, Tessarollo L, Kaldis P (2003) Cdk2 knockout mice are viable. Curr Biol 13(20):1775–1785, PubMed PMID: 14561402. eng
Ortega S, Prieto I, Odajima J, Martín A, Dubus P, Sotillo R et al (2003) Cyclin-dependent kinase 2 is essential for meiosis but not for mitotic cell division in mice. Nat Genet 35(1):25–31, PubMed PMID: 12923533. eng
Rane SG, Dubus P, Mettus RV, Galbreath EJ, Boden G, Reddy EP et al (1999) Loss of Cdk4 expression causes insulin-deficient diabetes and Cdk4 activation results in beta-islet cell hyperplasia. Nat Genet 22(1):44–52, PubMed PMID: 10319860. eng
Malumbres M, Sotillo R, Santamaría D, Galán J, Cerezo A, Ortega S et al (2004) Mammalian cells cycle without the D-type cyclin-dependent kinases Cdk4 and Cdk6. Cell 118(4):493–504, PubMed PMID: 15315761. eng
Malumbres M, Barbacid M (2005) Mammalian cyclin-dependent kinases. Trends Biochem Sci 30(11):630–641, PubMed PMID: 16236519. eng
Akli S, Van Pelt CS, Bui T, Meijer L, Keyomarsi K (2011) Cdk2 is required for breast cancer mediated by the low-molecular-weight isoform of cyclin E. Cancer Res 71(9):3377–3386, PubMed PMID: 21385896. Pubmed Central PMCID: PMC3085722. eng
Puyol M, Martín A, Dubus P, Mulero F, Pizcueta P, Khan G et al (2010) A synthetic lethal interaction between K-Ras oncogenes and Cdk4 unveils a therapeutic strategy for non-small cell lung carcinoma. Cancer Cell 18(1):63–73, PubMed PMID: 20609353. eng
Chao SH, Price DH (2001) Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J Biol Chem 276(34):31793–31799, PubMed PMID: 11431468. eng
Sedlacek H, Czech J, Naik R, Kaur G, Worland P, Losiewicz M et al (1996) Flavopiridol (L86 8275; NSC 649890), a new kinase inhibitor for tumor therapy. Int J Oncol 9(6):1143–1168, PubMed PMID: 21541623. eng
Fisher RP, Morgan DO (1994) A novel cyclin associates with MO15/CDK7 to form the CDK-activating kinase. Cell 78(4):713–724, PubMed PMID: 8069918. eng
Roy R, Adamczewski JP, Seroz T, Vermeulen W, Tassan JP, Schaeffer L et al (1994) The MO15 cell cycle kinase is associated with the TFIIH transcription-DNA repair factor. Cell 79(6):1093–1101, PubMed PMID: 8001135. eng
Kim YK, Bourgeois CF, Isel C, Churcher MJ, Karn J (2002) Phosphorylation of the RNA polymerase II carboxyl-terminal domain by CDK9 is directly responsible for human immunodeficiency virus type 1 Tat-activated transcriptional elongation. Mol Cell Biol 22(13):4622–4637, PubMed PMID: 12052871. Pubmed Central PMCID: PMC133925. eng
Lam LT, Pickeral OK, Peng AC, Rosenwald A, Hurt EM, Giltnane JM et al (2001) Genomic-scale measurement of mRNA turnover and the mechanisms of action of the anti-cancer drug flavopiridol. Genome Biol 2(10): RESEARCH0041 PubMed PMID: 11597333. Pubmed Central PMCID: PMC57796. eng http://genomebiology.com/content/pdf/gb-2001-2-10-research0041.pdf
Garriga J, Graña X (2004) Cellular control of gene expression by T-type cyclin/CDK9 complexes. Gene 337:15–23, PubMed PMID: 15276198. eng
Byrd JC, Lin TS, Dalton JT, Wu D, Phelps MA, Fischer B et al (2007) Flavopiridol administered using a pharmacologically derived schedule is associated with marked clinical efficacy in refractory, genetically high-risk chronic lymphocytic leukemia. Blood 109(2):399–404, PubMed PMID: 17003373. Pubmed Central PMCID: PMC1785084. eng
Tan AR, Headlee D, Messmann R, Sausville EA, Arbuck SG, Murgo AJ et al (2002) Phase I clinical and pharmacokinetic study of flavopiridol administered as a daily 1-hour infusion in patients with advanced neoplasms. J Clin Oncol 20(19):4074–4082, PubMed PMID: 12351605. eng
Burdette-Radoux S, Tozer RG, Lohmann RC, Quirt I, Ernst DS, Walsh W et al (2004) Phase II trial of flavopiridol, a cyclin dependent kinase inhibitor, in untreated metastatic malignant melanoma. Invest New Drugs 22(3):315–322, PubMed PMID: 15122079. eng
Grendys EC, Blessing JA, Burger R, Hoffman J (2005) A phase II evaluation of flavopiridol as second-line chemotherapy of endometrial carcinoma: a gynecologic oncology group study. Gynecol Oncol 98(2):249–253, PubMed PMID: 15978659. eng
Dispenzieri A, Gertz MA, Lacy MQ, Geyer SM, Fitch TR, Fenton RG et al (2006) Flavopiridol in patients with relapsed or refractory multiple myeloma: a phase 2 trial with clinical and pharmacodynamic end-points. Haematologica 91(3):390–393, PubMed PMID: 16503551. eng
Phelps MA, Lin TS, Johnson AJ, Hurh E, Rozewski DM, Farley KL et al (2009) Clinical response and pharmacokinetics from a phase 1 study of an active dosing schedule of flavopiridol in relapsed chronic lymphocytic leukemia. Blood 113(12):2637–2645, PubMed PMID: 18981292. Pubmed Central PMCID: PMC2661854. eng
Matranga CB, Shapiro GI (2002) Selective sensitization of transformed cells to flavopiridol-induced apoptosis following recruitment to S-phase. Cancer Res 62(6):1707–1717, PubMed PMID: 11912144. eng
Shah MA, Kortmansky J, Motwani M, Drobnjak M, Gonen M, Yi S et al (2005) A phase I clinical trial of the sequential combination of irinotecan followed by flavopiridol. Clin Cancer Res 11(10):3836–3845, PubMed PMID: 15897584. eng
Rathkopf D, Dickson MA, Feldman DR, Carvajal RD, Shah MA, Wu N et al (2009) Phase I study of flavopiridol with oxaliplatin and fluorouracil/leucovorin in advanced solid tumors. Clin Cancer Res 15(23):7405–7411, PubMed PMID: 19934304. Pubmed Central PMCID: PMC2787644. eng
George S, Kasimis BS, Cogswell J, Schwarzenberger P, Shapiro GI, Fidias P et al (2008) Phase I study of flavopiridol in combination with paclitaxel and carboplatin in patients with non-small-cell lung cancer. Clin Lung Cancer 9(3):160–165, PubMed PMID: 18621626. eng
El-Rayes BF, Gadgeel S, Parchment R, Lorusso P, Philip PA (2006) A phase I study of flavopiridol and docetaxel. Invest New Drugs 24(4):305–310, PubMed PMID: 16683073. eng
Fornier MN, Rathkopf D, Shah M, Patil S, O’Reilly E, Tse AN et al (2007) Phase I dose-finding study of weekly docetaxel followed by flavopiridol for patients with advanced solid tumors. Clin Cancer Res 13(19):5841–5846, PubMed PMID: 17908977. eng
Bible KC, Lensing JL, Nelson SA, Lee YK, Reid JM, Ames MM et al (2005) Phase 1 trial of flavopiridol combined with cisplatin or carboplatin in patients with advanced malignancies with the assessment of pharmacokinetic and pharmacodynamic end points. Clin Cancer Res 11(16):5935–5941, PubMed PMID: 16115936. eng
Karp JE, Passaniti A, Gojo I, Kaufmann S, Bible K, Garimella TS et al (2005) Phase I and pharmacokinetic study of flavopiridol followed by 1-beta-D-arabinofuranosylcytosine and mitoxantrone in relapsed and refractory adult acute leukemias. Clin Cancer Res 11(23): 8403–8412, PubMed PMID: 16322302. eng
Bible KC, Peethambaram PP, Oberg AL, Maples W, Groteluschen DL, Boente M et al (2012) A phase 2 trial of flavopiridol (alvocidib) and cisplatin in platin-resistant ovarian and primary peritoneal carcinoma: MC0261. Gynecol Oncol 127(1):55–62, PubMed PMID: 22664059. eng
Rudek MA, Bauer KS, Lush RM, Stinson SF, Senderowicz AM, Headlee DJ et al (2003) Clinical pharmacology of flavopiridol following a 72-hour continuous infusion. Ann Pharmacother 37(10):1369–1374, PubMed PMID: 14519054. eng
Li P, Tabibi SE, Yalkowsky SH (1999) Solubilization of flavopiridol by pH control combined with cosolvents, surfactants, or complexants. J Pharm Sci 88(9):945–947, PubMed PMID: 10479359. eng
Yang X, Zhao X, Phelps MA, Piao L, Rozewski DM, Liu Q et al (2009) A novel liposomal formulation of flavopiridol. Int J Pharm 365(1-2):170–174, PubMed PMID: 18778761. Pubmed Central PMCID: PMC3035394. eng
Santarius T, Shipley J, Brewer D, Stratton MR, Cooper CS (2010) A census of amplified and overexpressed human cancer genes. Nat Rev Cancer 10(1):59–64, PubMed PMID: 20029424. eng
Arnold A, Papanikolaou A (2005) Cyclin D1 in breast cancer pathogenesis. J Clin Oncol 23(18):4215–4224, PubMed PMID: 15961768. eng
Fry DW, Harvey PJ, Keller PR, Elliott WL, Meade M, Trachet E et al (2004) Specific inhibition of cyclin-dependent kinase 4/6 by PD 0332991 and associated antitumor activity in human tumor xenografts. Mol Cancer Ther 3(11):1427–1438, PubMed PMID: 15542782. eng
Toogood PL, Harvey PJ, Repine JT, Sheehan DJ, VanderWel SN, Zhou H et al (2005) Discovery of a potent and selective inhibitor of cyclin-dependent kinase 4/6. J Med Chem 48(7):2388–2406, PubMed PMID: 15801831. eng
Dean JL, Thangavel C, McClendon AK, Reed CA, Knudsen ES (2010) Therapeutic CDK4/6 inhibition in breast cancer: key mechanisms of response and failure. Oncogene 29(28):4018–4032, PubMed PMID: 20473330. eng
Ertel A, Dean JL, Rui H, Liu C, Witkiewicz AK, Knudsen KE et al (2010) RB-pathway disruption in breast cancer: differential association with disease subtypes, disease-specific prognosis and therapeutic response. Cell Cycle 9(20):4153–4163, PubMed PMID: 20948315. Pubmed Central PMCID: PMC3055199. eng
Finn RS, Dering J, Conklin D, Kalous O, Cohen DJ, Desai AJ et al (2009) PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res 11(5):R77 PubMed PMID: 19874578. Pubmed Central PMCID: PMC2790859. Eng
Dean JL, McClendon AK, Hickey TE, Butler LM, Tilley WD, Witkiewicz AK et al (2012) Therapeutic response to CDK4/6 inhibition in breast cancer defined by ex vivo analyses of human tumors. Cell Cycle 11(14):2756–2761, PubMed PMID: 22767154. Pubmed Central PMCID: PMC3409015. eng
Tang LH, Contractor T, Clausen R, Klimstra DS, Du YC, Allen PJ et al (2012) Attenuation of the retinoblastoma pathway in pancreatic neuroendocrine tumors due to increased cdk4/cdk6. Clin Cancer Res 18(17):4612–4620, PubMed PMID: 22761470. eng
Cen L, Carlson BL, Schroeder MA, Ostrem JL, Kitange GJ, Mladek AC et al (2012) p16-Cdk4-Rb axis controls sensitivity to a cyclin-dependent kinase inhibitor PD0332991 in glioblastoma xenograft cells. Neuro Oncol 14(7):870–881, PubMed PMID: 22711607. Pubmed Central PMCID: PMC3379801. eng
Saab R, Bills JL, Miceli AP, Anderson CM, Khoury JD, Fry DW et al (2006) Pharmacologic inhibition of cyclin-dependent kinase 4/6 activity arrests proliferation in myoblasts and rhabdomyosarcoma-derived cells. Mol Cancer Ther 5(5):1299–1308, PubMed PMID: 16731763. eng
Leonard JP, LaCasce AS, Smith MR, Noy A, Chirieac LR, Rodig SJ et al (2012) Selective CDK4/6 inhibition with tumor responses by PD0332991 in patients with mantle cell lymphoma. Blood 119(20):4597–4607, PubMed PMID: 22383795. eng
Michaud K, Solomon DA, Oermann E, Kim JS, Zhong WZ, Prados MD et al (2010) Pharmacologic inhibition of cyclin-dependent kinases 4 and 6 arrests the growth of glioblastoma multiforme intracranial xenografts. Cancer Res 70(8):3228–3238, PubMed PMID: 20354191. Pubmed Central PMCID: PMC2855904. eng
Liu F, Korc M (2012) Cdk4/6 inhibition induces epithelial-mesenchymal transition and enhances invasiveness in pancreatic cancer cells. Mol Cancer Ther 11(10):2138–2148, PubMed PMID: 22869556. eng
Flaherty KT, Lorusso PM, Demichele A, Abramson VG, Courtney R, Randolph SS et al (2012) Phase I, dose-escalation trial of the oral cyclin-dependent kinase 4/6 inhibitor PD 0332991, administered using a 21-day schedule in patients with advanced cancer. Clin Cancer Res 18(2):568–576, PubMed PMID: 22090362. eng
Dean JL, McClendon AK, Knudsen ES (2012) Modification of the DNA damage response by therapeutic CDK4/6 inhibition. J Biol Chem 287(34):29075–29087, PubMed PMID: 22733811. Pubmed Central PMCID: PMC3436568. eng
Zwart W, Rondaij M, Jalink K, Sharp ZD, Mancini MA, Neefjes J et al (2009) Resistance to antiestrogen arzoxifene is mediated by overexpression of cyclin D1. Mol Endocrinol 23(9):1335–1345, PubMed PMID: 19477949. Pubmed Central PMCID: PMC2737554. eng
Kilker RL, Planas-Silva MD (2006) Cyclin D1 is necessary for tamoxifen-induced cell cycle progression in human breast cancer cells. Cancer Res 66(23):11478–11484, PubMed PMID: 17145896. eng
Zwijsen RM, Wientjens E, Klompmaker R, van der Sman J, Bernards R, Michalides RJ (1997) CDK-independent activation of estrogen receptor by cyclin D1. Cell 88(3):405–415, PubMed PMID: 9039267. eng
Musgrove EA, Sutherland RL (2009) Biological determinants of endocrine resistance in breast cancer. Nat Rev Cancer 9(9):631–643, PubMed PMID: 19701242. eng
Wang L, Wang J, Blaser BW, Duchemin AM, Kusewitt DF, Liu T et al (2007) Pharmacologic inhibition of CDK4/6: mechanistic evidence for selective activity or acquired resistance in acute myeloid leukemia. Blood 110(6):2075–2083, PubMed PMID: 17537993. eng
Meijer L, Raymond E (2003) Roscovitine and other purines as kinase inhibitors. From starfish oocytes to clinical trials. Acc Chem Res 36(6):417–425, PubMed PMID: 12809528. eng
De Azevedo WF, Leclerc S, Meijer L, Havlicek L, Strnad M, Kim SH (1997) Inhibition of cyclin-dependent kinases by purine analogues: crystal structure of human cdk2 complexed with roscovitine. Eur J Biochem 243(1-2):518–526, PubMed PMID: 9030780. eng
Raynaud FI, Whittaker SR, Fischer PM, McClue S, Walton MI, Barrie SE et al (2005) In vitro and in vivo pharmacokinetic-pharmacodynamic relationships for the trisubstituted aminopurine cyclin-dependent kinase inhibitors olomoucine, bohemine and CYC202. Clin Cancer Res 11(13):4875–4887, PubMed PMID: 16000586. eng
Wesierska-Gadek J, Gueorguieva M, Horky M (2003) Dual action of cyclin-dependent kinase inhibitors: induction of cell cycle arrest and apoptosis. A comparison of the effects exerted by roscovitine and cisplatin. Pol J Pharmacol 55(5):895–902, PubMed PMID: 14704484. eng
Whittaker SR, Walton MI, Garrett MD, Workman P (2004) The cyclin-dependent kinase inhibitor CYC202 (R-roscovitine) inhibits retinoblastoma protein phosphorylation, causes loss of cyclin D1, and activates the mitogen-activated protein kinase pathway. Cancer Res 64(1):262–272, PubMed PMID: 14729633. eng
McClue SJ, Blake D, Clarke R, Cowan A, Cummings L, Fischer PM et al (2002) In vitro and in vivo antitumor properties of the cyclin dependent kinase inhibitor CYC202 (R-roscovitine). Int J Cancer 102(5):463–468, PubMed PMID: 12432547. eng
Wang D, de la Fuente C, Deng L, Wang L, Zilberman I, Eadie C et al (2001) Inhibition of human immunodeficiency virus type 1 transcription by chemical cyclin-dependent kinase inhibitors. J Virol 75(16):7266–7279, PubMed PMID: 11461999. Pubmed Central PMCID: PMC114962. eng
MacCallum DE, Melville J, Frame S, Watt K, Anderson S, Gianella-Borradori A et al (2005) Seliciclib (CYC202, R-roscovitine) induces cell death in multiple myeloma cells by inhibition of RNA polymerase II-dependent transcription and down-regulation of Mcl-1. Cancer Res 65(12):5399–5407, PubMed PMID: 15958589. eng
Fischer PM, Gianella-Borradori A (2003) CDK inhibitors in clinical development for the treatment of cancer. Expert Opin Investig Drugs 12(6):955–970, PubMed PMID: 12783600. eng
Appleyard MV, O’Neill MA, Murray KE, Paulin FE, Bray SE, Kernohan NM et al (2009) Seliciclib (CYC202, R-roscovitine) enhances the antitumor effect of doxorubicin in vivo in a breast cancer xenograft model. Int J Cancer 124(2):465–472, PubMed PMID: 19003963. eng
Nanos-Webb A, Jabbour NA, Multani AS, Wingate H, Oumata N, Galons H et al (2012) Targeting low molecular weight cyclin E (LMW-E) in breast cancer. Breast Cancer Res Treat 132(2):575–588, PubMed PMID: 21695458. eng
O’Connor DS, Wall NR, Porter AC, Altieri DC (2002) A p34(cdc2) survival checkpoint in cancer. Cancer Cell 2(1):43–54, PubMed PMID: 12150824. eng
Guzi T (2004) CYC-202 cyclacel. Curr Opin Investig Drugs 5(12):1311–1318, PubMed PMID: 15648953. eng
Benson C, White J, De Bono J, O’Donnell A, Raynaud F, Cruickshank C et al (2007) A phase I trial of the selective oral cyclin-dependent kinase inhibitor seliciclib (CYC202; R-roscovitine), administered twice daily for 7 days every 21 days. Br J Cancer 96(1):29–37, PubMed PMID: 17179992. Pubmed Central PMCID: PMC2360206. eng
Le Tourneau C, Faivre S, Laurence V, Delbaldo C, Vera K, Girre V et al (2010) Phase I evaluation of seliciclib (R-roscovitine), a novel oral cyclin-dependent kinase inhibitor, in patients with advanced malignancies. Eur J Cancer 46(18):3243–3250, PubMed PMID: 20822897. eng
Hsieh WS, Soo R, Peh BK, Loh T, Dong D, Soh D et al (2009) Pharmacodynamic effects of seliciclib, an orally administered cell cycle modulator, in undifferentiated nasopharyngeal cancer. Clin Cancer Res 15(4):1435–1442, PubMed PMID: 19228744. eng
Bettayeb K, Oumata N, Echalier A, Ferandin Y, Endicott JA, Galons H et al (2008) CR8, a potent and selective, roscovitine-derived inhibitor of cyclin-dependent kinases. Oncogene 27(44):5797–5807, PubMed PMID: 18574471. eng
Bettayeb K, Baunbæk D, Delehouze C, Loaëc N, Hole AJ, Baumli S et al (2010) CDK inhibitors roscovitine and CR8 trigger Mcl-1 down-regulation and apoptotic cell death in neuroblastoma cells. Genes Cancer 1(4):369–380, PubMed PMID: 21779453. Pubmed Central PMCID: PMC3092200. eng
Kabadi SV, Stoica BA, Hanscom M, Loane DJ, Kharebava G, Murray Ii MG et al (2012) CR8, a selective and potent CDK inhibitor, provides neuroprotection in experimental traumatic brain injury. Neurotherapeutics 9(2):405–421, PubMed PMID: 22167461. Pubmed Central PMCID: PMC3324621. eng
Parry D, Guzi T, Shanahan F, Davis N, Prabhavalkar D, Wiswell D et al (2010) Dinaciclib (SCH 727965), a novel and potent cyclin-dependent kinase inhibitor. Mol Cancer Ther 9(8):2344–2353, PubMed PMID: 20663931. eng
Feldmann G, Mishra A, Bisht S, Karikari C, Garrido-Laguna I, Rasheed Z et al (2011) Cyclin-dependent kinase inhibitor dinaciclib (SCH727965) inhibits pancreatic cancer growth and progression in murine xenograft models. Cancer Biol Ther 12(7):598–609, PubMed PMID: 21768779. Pubmed Central PMCID: PMC3218385. eng
Abdullah C, Wang X, Becker D (2011) Expression analysis and molecular targeting of cyclin-dependent kinases in advanced melanoma. Cell Cycle 10(6):977–988, PubMed PMID: 21358262. Pubmed Central PMCID: PMC3100877. eng
Fu W, Ma L, Chu B, Wang X, Bui MM, Gemmer J et al (2011) The cyclin-dependent kinase inhibitor SCH 727965 (dinacliclib) induces the apoptosis of osteosarcoma cells. Mol Cancer Ther 10(6):1018–1027, PubMed PMID: 21490307. eng
Gorlick R, Kolb EA, Houghton PJ, Morton CL, Neale G, Keir ST et al (2012) Initial testing (stage 1) of the cyclin dependent kinase inhibitor SCH 727965 (dinaciclib) by the pediatric preclinical testing program. Pediatr Blood Cancer 59(7):1266–1274, PubMed PMID: 22315240. Pubmed Central PMCID: PMC3349821. eng
Tentler JJ, Tan AC, Weekes CD, Jimeno A, Leong S, Pitts TM et al (2012) Patient-derived tumour xenografts as models for oncology drug development. Nat Rev Clin Oncol 9(6):338–350, PubMed PMID: 22508028. eng
Johnson AJ, Yeh YY, Smith LL, Wagner AJ, Hessler J, Gupta S et al (2012) The novel cyclin-dependent kinase inhibitor dinaciclib (SCH727965) promotes apoptosis and abrogates microenvironmental cytokine protection in chronic lymphocytic leukemia cells. 12:2554–7, Leukemia.PubMed PMID: 22791353. ENG
Shapiro G, Bannerji R, Small K, Black S, Statkevich P, Abutarif M (eds) (2008) A phase I dose-escalation study of the safety, pharmacokinetics (PK) and pharmacodynamics (PD) of the novel cyclin-dependent kinase inhibitor SCH 727965 administered every 3 weeks in subjects with advanced malignancies. ASCO, Chicago, ASCO Annual Meeting
Zhang D, Mita M, Shapiro GI, Poon J, Small K, Tzontcheva A et al (2012) Effect of aprepitant on the pharmacokinetics of the cyclin-dependent kinase inhibitor dinaciclib in patients with advanced malignancies. Cancer Chemother Pharmacol. ASH stands for the American Society of Hematology. Headquarters are in Washington DC 6:891–8, PubMed PMID: 23053255. ENG
Gojo I, Walker AR, Cooper M, Feldman EJ, Padmanabhan S, Baer MR et al (eds) (2010) Phase II Study of the Cyclin-Dependent Kinase (CDK) Inhibitor Dinaciclib (SCH 727965) In Patients with Advanced Acute Leukemias ASH
Available from: http://clinicaltrials.gov/ct2/show/NCT01096342
Lao CD, Moon J, Fruehauf JP, Flaherty LE, Bury MJ, Ribas A et al (eds) (2012) SWOG S0826: A phase II trial of SCH 727965 (NSC 747135) in patients with stage IV melanoma. 2012 ASCO annual meeting, Chicago
SCH 727965 in Patients with advanced breast and lung cancers (Study P04716AM3) [Internet]. Available from: http://clinicaltrials.gov/ct2/show/NCT00732810
Veliparib and dinaciclib with or without carboplatin in treating patients with advanced solid tumors: http://clinicaltrials.gov/show/NCT01434316
A study of dinaciclib in combination with rituximab in participants with chronic lymphocytic leukemia and small lymphocytic lymphoma (P07974 AM1) http://clinicaltrials.gov/ct2/show/NCT01650727
Dinaciclib, bortezomib, and dexamethasone in treating patients with relapsed multiple myeloma [Internet]. Available from: http://www.clinicaltrials.gov/ct2/show/NCT01711528
Dinaciclib and epirubicin hydrochloride in treating patients with metastatic triple-negative breast cancer [Internet]. Available from: http://clinicaltrials.gov/show/NCT01624441
Akli S, Van Pelt CS, Bui T, Meijer L, Keyomarsi K (2011) Cdk2 Is required for breast cancer mediated by the Low-molecular-weight isoform of cyclin E. Cancer Res 71(9):3377–3386, PubMed PMID: 21385896. Pubmed Central PMCID: PMC3085722. eng
Horiuchi D, Kusdra L, Huskey NE, Chandriani S, Lenburg ME, Gonzalez-Angulo AM et al (2012) MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J Exp Med 209(4):679–696, PubMed PMID: 22430491. Pubmed Central PMCID: PMC3328367. eng
Deans AJ, Khanna KK, McNees CJ, Mercurio C, Heierhorst J, McArthur GA (2006) Cyclin-dependent kinase 2 functions in normal DNA repair and is a therapeutic target in BRCA1-deficient cancers. Cancer Res 66(16):8219–8226, PubMed PMID: 16912201. eng
Lee J, Kim SS (2009) The function of p27 KIP1 during tumor development. Exp Mol Med 41(11):765–771, PubMed PMID: 19887899. Pubmed Central PMCID: PMC2788730 eng
Liang J, Shao SH, Xu ZX, Hennessy B, Ding Z, Larrea M et al (2007) The energy sensing LKB1-AMPK pathway regulates p27(kip1) phosphorylation mediating the decision to enter autophagy or apoptosis. Nat Cell Biol 9(2):218–224, PubMed PMID: 17237771. eng
Short JD, Houston KD, Dere R, Cai SL, Kim J, Johnson CL et al (2008) AMP-activated protein kinase signaling results in cytoplasmic sequestration of p27. Cancer Res 68(16):6496–6506, PubMed PMID: 18701472. Pubmed Central PMCID: PMC3011867. eng
Besson A, Gurian-West M, Schmidt A, Hall A, Roberts JM (2004) p27Kip1 modulates cell migration through the regulation of RhoA activation. Genes Dev 18(8):862–876, PubMed PMID: 15078817. Pubmed Central PMCID: PMC395846. eng
Besson A, Gurian-West M, Chen X, Kelly-Spratt KS, Kemp CJ, Roberts JM (2006) A pathway in quiescent cells that controls p27Kip1 stability, subcellular localization, and tumor suppression. Genes Dev 20(1):47–64, PubMed PMID: 16391232. Pubmed Central PMCID: PMC1356100. eng
Besson A, Hwang HC, Cicero S, Donovan SL, Gurian-West M, Johnson D et al (2007) Discovery of an oncogenic activity in p27Kip1 that causes stem cell expansion and a multiple tumor phenotype. Genes Dev 21(14):1731–1746, PubMed PMID: 17626791. Pubmed Central PMCID: PMC1920168. eng
Borriello A, Bencivenga D, Criscuolo M, Caldarelli I, Cucciolla V, Tramontano A et al (2011) Targeting p27Kip1 protein: its relevance in the therapy of human cancer. Expert Opin Ther Targets 15(6):677–693, PubMed PMID: 21355788. eng
Martínez-Sánchez A, Gebauer F (2010) Regulation of p27(kip1) mRNA expression by microRNAs. Prog Mol Subcell Biol 50:59–70, PubMed PMID: 19841881. eng
Tomoda K, Kubota Y, Kato J (1999) Degradation of the cyclin-dependent-kinase inhibitor p27Kip1 is instigated by Jab1. Nature 398(6723):160–165, PubMed PMID: 10086358. eng
Sui L, Dong Y, Ohno M, Watanabe Y, Sugimoto K, Tai Y et al (2001) Jab1 expression is associated with inverse expression of p27(kip1) and poor prognosis in epithelial ovarian tumors. Clin Cancer Res 7(12):4130–4135, PubMed PMID: 11751512. eng
Rassidakis GZ, Claret FX, Lai R, Zhang Q, Sarris AH, McDonnell TJ et al (2003) Expression of p27(Kip1) and c-Jun activation binding protein 1 are inversely correlated in systemic anaplastic large cell lymphoma. Clin Cancer Res 9(3):1121–1128, PubMed PMID: 12631617. eng
Kouvaraki MA, Rassidakis GZ, Tian L, Kumar R, Kittas C, Claret FX (2003) Jun activation domain-binding protein 1 expression in breast cancer inversely correlates with the cell cycle inhibitor p27(Kip1). Cancer Res 63(11):2977–2981, PubMed PMID: 12782606. eng
Fukumoto A, Ikeda N, Sho M, Tomoda K, Kanehiro H, Hisanaga M et al (2004) Prognostic significance of localized p27Kip1 and potential role of Jab1/CSN5 in pancreatic cancer. Oncol Rep 11(2):277–284, PubMed PMID: 14719054. eng
Pan Y, Zhang Q, Atsaves V, Yang H, Claret FX (2012) Suppression of Jab1/CSN5 induces radio- and chemo-sensitivity in nasopharyngeal carcinoma through changes to the DNA damage and repair pathways, Oncogene. PubMed PMID: 22797071. ENG
Wan M, Cao X, Wu Y, Bai S, Wu L, Shi X et al (2002) Jab1 Antagonizes TGF-beta signaling by inducing Smad4 degradation. EMBO Rep 3(2):171–176, PubMed PMID: 11818334. Pubmed Central PMCID: PMC1083965. eng
Lee EW, Oh W, Song J (2006) Jab1 As a mediator of nuclear export and cytoplasmic degradation of p53. Mol Cells 22(2):133–140, PubMed PMID: 17085963. eng
Tian L, Peng G, Parant JM, Leventaki V, Drakos E, Zhang Q et al (2010) Essential roles of Jab1 in cell survival, spontaneous DNA damage and DNA repair. Oncogene 29(46):6125–6137, PubMed PMID: 20802511. Pubmed Central PMCID: PMC3495558. eng
Carrano AC, Eytan E, Hershko A, Pagano M (1999) SKP2 Is required for ubiquitin-mediated degradation of the CDK inhibitor p27. Nat Cell Biol 1(4):193–199, PubMed PMID: 10559916. eng
Kamura T, Hara T, Kotoshiba S, Yada M, Ishida N, Imaki H et al (2003) Degradation of p57Kip2 mediated by SCFSkp2-dependent ubiquitylation. Proc Natl Acad Sci U S A 100(18):10231–10236, PubMed PMID: 12925736. Pubmed Central PMCID: PMC193544. eng
Huang H, Regan KM, Wang F, Wang D, Smith DI, van Deursen JM et al (2005) Skp2 inhibits FOXO1 in tumor suppression through ubiquitin-mediated degradation. Proc Natl Acad Sci U S A 102(5):1649–1654, PubMed PMID: 15668399. Pubmed Central PMCID: PMC545492. eng
Kim SY, Herbst A, Tworkowski KA, Salghetti SE, Tansey WP (2003) Skp2 regulates Myc protein stability and activity. Mol Cell 11(88):1177–1188, PubMed PMID: 12769843. eng
von der Lehr N, Johansson S, Wu S, Bahram F, Castell A, Cetinkaya C et al (2003) The F-box protein Skp2 participates in c-Myc proteosomal degradation and acts as a cofactor for c-Myc-regulated transcription. Mol Cell 11(5):1189–1200, PubMed PMID: 12769844. eng
Latres E, Chiarle R, Schulman BA, Pavletich NP, Pellicer A, Inghirami G et al (2001) Role of the F-box protein Skp2 in lymphomagenesis. Proc Natl Acad Sci U S A 98(5):2515–2520, PubMed PMID: 11226270. Pubmed Central PMCID: PMC30169. eng
Yokoi S, Yasui K, Saito-Ohara F, Koshikawa K, Iizasa T, Fujisawa T et al (2002) A novel target gene, SKP2, within the 5p13 amplicon that is frequently detected in small cell lung cancers. Am J Pathol 161(1):207–216, PubMed PMID: 12107105. Pubmed Central PMCID: PMC1850681. eng
Gstaiger M, Jordan R, Lim M, Catzavelos C, Mestan J, Slingerland J et al (2001) Skp2 is oncogenic and overexpressed in human cancers. Proc Natl Acad Sci U S A 98(9):5043–5048, PubMed PMID: 11309491. Pubmed Central PMCID: PMC33160. eng
Lin HK, Chen Z, Wang G, Nardella C, Lee SW, Chan CH et al (2010) Skp2 Targeting suppresses tumorigenesis by Arf-p53-independent cellular senescence. Nature 464(9):374–379, PubMed PMID: 20237562. Pubmed Central PMCID: PMC2928066. eng
Chen Q, Xie W, Kuhn DJ, Voorhees PM, Lopez-Girona A, Mendy D et al (2008) Targeting the p27 E3 ligase SCF(Skp2) results in p27- and Skp2-mediated cell-cycle arrest and activation of autophagy. Blood 111(9):4690–4699, PubMed PMID: 18305219. Pubmed Central PMCID: PMC2343599. eng
Rico-Bautista E, Yang CC, Lu L, Roth GP, Wolf DA (2010) Chemical genetics approach to restoring p27Kip1 reveals novel compounds with antiproliferative activity in prostate cancer cells. BMC Biol 8:153, PubMed PMID: 21182779. Pubmed Central PMCID: PMC3025922. eng
Koutsami M, Velimezi G, Kotsinas A, Evangelou K, Papavassiliou AG, Kittas C et al (2008) Is exclusive Skp2 targeting always beneficial in cancer therapy? Blood 112(12):4777–4779, PubMed PMID: 19029458. eng
Chen X, Lowe M, Keyomarsi K (1999) UCN-01-mediated G1 arrest in normal but not tumor breast cells is pRb-dependent and p53-independent. Oncogene 18(41):5691, PubMed PMID: 10577141. eng
Chen X, Lowe M, Herliczek T, Hall MJ, Danes C, Lawrence DA et al (2000) Protection of normal proliferating cells against chemotherapy by staurosporine-mediated, selective, and reversible G(1) arrest. J Natl Cancer Inst 92(24):1999–2008, PubMed PMID: 11121462. eng
Roberts PJ, Bisi JE, Strum JC, Combest AJ, Darr DB, Usary JE et al (2012) Multiple roles of cyclin-dependent kinase 4/6 inhibitors in cancer therapy. J Natl Cancer Inst 104(6):476–487, PubMed PMID: 22302033. Pubmed Central PMCID: PMC3309128. eng
Nawijn MC, Alendar A, Berns A (2011) For better or for worse: the role of Pim oncogenes in tumorigenesis. Nat Rev Cancer 11(1):23–34, PubMed PMID: 21150935. eng
Cuypers HT, Selten G, Quint W, Zijlstra M, Maandag ER, Boelens W et al (1984) Murine leukemia virus-induced T-cell lymphomagenesis: integration of proviruses in a distinct chromosomal region. Cell 37(1):141–150, PubMed PMID: 6327049. eng
van Lohuizen M, Verbeek S, Krimpenfort P, Domen J, Saris C, Radaszkiewicz T et al (1989) Predisposition to lymphomagenesis in pim-1 transgenic mice: cooperation with c-myc and N-myc in murine leukemia virus-induced tumors. Cell 56(4):673–682, PubMed PMID: 2537153. eng
Verbeek S, van Lohuizen M, van der Valk M, Domen J, Kraal G, Berns A (1991) Mice bearing the E mu-myc and E mu-pim-1 transgenes develop pre-B-cell leukemia prenatally. Mol Cell Biol 11(2):1176–1179, PubMed PMID: 1990273. Pubmed Central PMCID: PMC359805. eng
Mikkers H, Nawijn M, Allen J, Brouwers C, Verhoeven E, Jonkers J et al (2004) Mice deficient for all PIM kinases display reduced body size and impaired responses to hematopoietic growth factors. Mol Cell Biol 24(13):6104–6115, PubMed PMID: 15199164. Pubmed Central PMCID: PMC480904. eng
Valdman A, Fang X, Pang ST, Ekman P, Egevad L (2004) Pim-1 expression in prostatic intraepithelial neoplasia and human prostate cancer. Prostate 60(4):367–371, PubMed PMID: 15264249. eng
Reiser-Erkan C, Erkan M, Pan Z, Bekasi S, Giese NA, Streit S et al (2008) Hypoxia-inducible proto-oncogene Pim-1 is a prognostic marker in pancreatic ductal adenocarcinoma. Cancer Biol Ther 7(9):1352–1359, PubMed PMID: 18708761. eng
Speers C, Tsimelzon A, Sexton K, Herrick AM, Gutierrez C, Culhane A et al (2009) Identification of novel kinase targets for the treatment of estrogen receptor-negative breast cancer. Clin Cancer Res 15(20):6327–6340, PubMed PMID: 19808870. Pubmed Central PMCID: PMC2763053. eng
Beier UH, Weise JB, Laudien M, Sauerwein H, Görögh T (2007) Overexpression of Pim-1 in head and neck squamous cell carcinomas. Int J Oncol 30(6):1381–1387, PubMed PMID: 17487358. eng
Tamburini J, Green AS, Bardet V, Chapuis N, Park S, Willems L et al (2009) Protein synthesis is resistant to rapamycin and constitutes a promising therapeutic target in acute myeloid leukemia. Blood 114(8):1618–1627, PubMed PMID: 19458359. eng
Cohen AM, Grinblat B, Bessler H, Kristt D, Kremer A, Schwartz A et al (2004) Increased expression of the hPim-2 gene in human chronic lymphocytic leukemia and non-Hodgkin lymphoma. Leuk Lymphoma 45(5):951–955, PubMed PMID: 15291354. eng
Qian KC, Wang L, Hickey ER, Studts J, Barringer K, Peng C et al (2005) Structural basis of constitutive activity and a unique nucleotide binding mode of human Pim-1 kinase. J Biol Chem 280(7):6130–6137, PubMed PMID: 15525646. eng
Amaravadi R, Thompson CB (2005) The survival kinases Akt and Pim as potential pharmacological targets. J Clin Invest 115(10):2618–2624, PubMed PMID: 16200194. Pubmed Central PMCID: PMC1236693. eng
Zhu N, Ramirez LM, Lee RL, Magnuson NS, Bishop GA, Gold MR (2002) CD40 Signaling in B cells regulates the expression of the Pim-1 kinase via the NF-kappa B pathway. J Immunol 168(2):744–754, PubMed PMID: 11777968. eng
Chen J, Kobayashi M, Darmanin S, Qiao Y, Gully C, Zhao R et al (2009) Hypoxia-mediated up-regulation of Pim-1 contributes to solid tumor formation. Am J Pathol 400(1):400–411, PubMed PMID: 19528349. Pubmed Central PMCID: PMC2708825. eng
Chen J, Kobayashi M, Darmanin S, Qiao Y, Gully C, Zhao R et al (2009) Pim-1 plays a pivotal role in hypoxia-induced chemoresistance. Oncogene 28(28):2581–2592, PubMed PMID: 19483729. Pubmed Central PMCID: PMC3358117. eng
Zhao Y, Hamza MS, Leong HS, Lim CB, Pan YF, Cheung E et al (2008) Kruppel-like factor 5 modulates p53-independent apoptosis through Pim1 survival kinase in cancer cells. Oncogene 27(1):1–8, PubMed PMID: 17603560. eng
Peltola K, Hollmen M, Maula SM, Rainio E, Ristamäki R, Luukkaa M et al (2009) Pim-1 kinase expression predicts radiation response in squamocellular carcinoma of head and neck and is under the control of epidermal growth factor receptor. Neoplasia 11(7):629–636, PubMed PMID: 19568408. Pubmed Central PMCID: PMC2697349. eng
Zhang Y, Wang Z, Magnuson NS (2007) Pim-1 kinase-dependent phosphorylation of p21Cip1/WAF1 regulates its stability and cellular localization in H1299 cells. Mol Cancer Res 5(9):909–922, PubMed PMID: 17855660. eng
Morishita D, Katayama R, Sekimizu K, Tsuruo T, Fujita N (2008) Pim kinases promote cell cycle progression by phosphorylating and down-regulating p27Kip1 at the transcriptional and posttranscriptional levels. Cancer Res 68(13):5076–5085, PubMed PMID: 18593906. eng
Mochizuki T, Kitanaka C, Noguchi K, Muramatsu T, Asai A, Kuchino Y (1999) Physical and functional interactions between Pim-1 kinase and Cdc25A phosphatase. Implications for the Pim-1-mediated activation of the c-Myc signaling pathway. J Biol Chem 274(26):18659–18666, PubMed PMID: 10373478. eng
Bachmann M, Hennemann H, Xing PX, Hoffmann I, Möröy T (2004) The oncogenic serine/threonine kinase Pim-1 phosphorylates and inhibits the activity of Cdc25C-associated kinase 1 (C-TAK1): a novel role for Pim-1 at the G2/M cell cycle checkpoint. J Biol Chem 279((46):48319–48328, PubMed PMID: 15319445. eng
Bachmann M, Kosan C, Xing PX, Montenarh M, Hoffmann I, Möröy T (2006) The oncogenic serine/threonine kinase Pim-1 directly phosphorylates and activates the G2/M specific phosphatase Cdc25C. Int J Biochem Cell Biol 38(3):430–443, PubMed PMID: 16356754. eng
Koike N, Maita H, Taira T, Ariga H, Iguchi-Ariga SM (2000) Identification of heterochromatin protein 1 (HP1) as a phosphorylation target by Pim-1 kinase and the effect of phosphorylation on the transcriptional repression function of HP1(1). FEBS Lett 467(1):17–21, PubMed PMID: 10664448. eng
Zhao T, Heyduk T, Eissenberg JC (2001) Phosphorylation site mutations in heterochromatin protein 1 (HP1) reduce or eliminate silencing activity. J Biol Chem 276(12):9512–9518, PubMed PMID: 11121421. eng
Aho TL, Sandholm J, Peltola KJ, Mankonen HP, Lilly M, Koskinen PJ (2004) Pim-1 kinase promotes inactivation of the pro-apoptotic Bad protein by phosphorylating it on the Ser112 gatekeeper site. FEBS Lett 571(1-3):43–49, PubMed PMID: 15280015. eng
Zhang Y, Wang Z, Li X, Magnuson NS (2008) Pim kinase-dependent inhibition of c-Myc degradation. Oncogene 27(35):4809–4819, PubMed PMID: 18438430. eng
Roh M, Gary B, Song C, Said-Al-Naief N, Tousson A, Kraft A et al (2003) Overexpression of the oncogenic kinase Pim-1 leads to genomic instability. Cancer Res 63(23):8079–8084, PubMed PMID: 14678956. Eng
Ha S, Iqbal NJ, Mita P, Ruoff R, Gerald WL, Lepor H et al (2012) Phosphorylation of the androgen receptor by PIM1 in hormone refractory prostate cancer. Oncogene. PubMed PMID: 22986532. ENG
Linn DE, Yang X, Xie Y, Alfano A, Deshmukh D, Wang X et al (2012) Differential regulation of androgen receptor by PIM-1 kinases via phosphorylation-dependent recruitment of distinct ubiquitin E3 ligases. J Biol Chem 287(27):22959–22968, PubMed PMID: 22584579. Pubmed Central PMCID: PMC3391098. Eng
Laird PW, van der Lugt NM, Clarke A, Domen J, Linders K, McWhir J et al (1993) Nucleic Acids Res 21(20):4750–4755, PubMed PMID: 8233823. Pubmed Central PMCID: PMC331501. eng
Chen LS, Redkar S, Bearss D, Wierda WG, Gandhi V (2009) Pim kinase inhibitor, SGI-1776, induces apoptosis in chronic lymphocytic leukemia cells. Blood 114(19):4150–4157, PubMed PMID: 19734450. Pubmed Central PMCID: PMC2774551. eng
Chen LS, Redkar S, Taverna P, Cortes JE, Gandhi V (2011) Mechanisms of cytotoxicity to Pim kinase inhibitor, SGI-1776, in acute myeloid leukemia. Blood 118(3):693–702, PubMed PMID: 21628411. Pubmed Central PMCID: PMC3142906. eng
Mumenthaler SM, Ng PY, Hodge A, Bearss D, Berk G, Kanekal S et al (2009) Pharmacologic inhibition of Pim kinases alters prostate cancer cell growth and resensitizes chemoresistant cells to taxanes. Mol Cancer Ther 8(10):2882–2893, PubMed PMID: 19825806. Pubmed Central PMCID: PMC2808126. eng
SuperGen discontinues clinical development of SGI-1776. Press release 18 Nov 2012. Available from: http://www.businesswire.com/news/home/20101110007218/en/SuperGen-Discontinues-Clinical-Development-SGI-1776
Beharry Z, Zemskova M, Mahajan S, Zhang F, Ma J, Xia Z et al (2009) Novel benzylidene-thiazolidine-2,4-diones inhibit Pim protein kinase activity and induce cell cycle arrest in leukemia and prostate cancer cells. Mol Cancer Ther 8(6):1473–1483, PubMed PMID: 19509254. Pubmed Central PMCID: PMC3415237. eng
Song JH, Kraft AS (2012) Pim kinase inhibitors sensitize prostate cancer cells to apoptosis triggered by Bcl-2 family inhibitor ABT-737. Cancer Res 72(1):294–303, PubMed PMID: 22080570. Pubmed Central PMCID: PMC3251634. eng
Dar AA, Goff LW, Majid S, Berlin J, El-Rifai W (2010) Aurora kinase inhibitors – rising stars in cancer therapeutics? Mol Cancer Ther 9(2):268–278, PubMed PMID: 20124450. Pubmed Central PMCID: PMC2820587. eng
Kimmins S, Crosio C, Kotaja N, Hirayama J, Monaco L, Höög C et al (2007) Differential functions of the Aurora-B and Aurora-C kinases in mammalian spermatogenesis. Mol Endocrinol 21(3):726–739, PubMed PMID: 17192404. eng
Khan J, Ezan F, Crémet JY, Fautrel A, Gilot D, Lambert M et al (2011) Overexpression of active Aurora-C kinase results in cell transformation and tumour formation. PLoS One 6(10):e26512, PubMed PMID: 22046298. Pubmed Central PMCID: PMC3203144. eng
Kimura M, Matsuda Y, Yoshioka T, Okano Y (1999) Cell cycle-dependent expression and centrosome localization of a third human aurora/Ipl1-related protein kinase, AIK3. J Biol Chem 274(11):7334–7340, PubMed PMID: 10066797. eng
Glover DM, Leibowitz MH, McLean DA, Parry H (1995) Mutations in aurora prevent centrosome separation leading to the formation of monopolar spindles. Cell 81(1):95–105, PubMed PMID: 7720077. eng
Kelly KR, Ecsedy J, Mahalingam D, Nawrocki ST, Padmanabhan S, Giles FJ et al (2011) Targeting aurora kinases in cancer treatment. Curr Drug Targets 12(14):2067–2078, PubMed PMID: 21777198. eng
Sasai K, Parant JM, Brandt ME, Carter J, Adams HP, Stass SA et al (2008) Targeted disruption of Aurora A causes abnormal mitotic spindle assembly, chromosome misalignment and embryonic lethality. Oncogene 27(29):4122–4177, PubMed PMID: 18345035. eng
Marumoto T, Honda S, Hara T, Nitta M, Hirota T, Kohmura E et al (2003) Aurora-A kinase maintains the fidelity of early and late mitotic events in HeLa cells. J Biol Chem 278(51):51786–51795, PubMed PMID: 14523000. eng
Littlepage LE, Wu H, Andresson T, Deanehan JK, Amundadottir LT, Ruderman JV (2002) Identification of phosphorylated residues that affect the activity of the mitotic kinase Aurora-A. Proc Natl Acad Sci U S A 99(24):15440–15445, PubMed PMID: 12422018. Pubmed Central PMCID: PMC137735. eng
Tanaka T, Kimura M, Matsunaga K, Fukada D, Mori H, Okano Y (1999) Centrosomal kinase AIK1 is overexpressed in invasive ductal carcinoma of the breast. Cancer Res 59(9):2041–2044, PubMed PMID: 10232583. eng
Lehman NL, O’Donnell JP, Whiteley LJ, Stapp RT, Lehman TD, Roszka KM et al (2012) Aurora A is differentially expressed in gliomas, is associated with patient survival in glioblastoma and is a potential chemotherapeutic target in gliomas. Cell Cycle 11(3):489–502, PubMed PMID: 22274399. Pubmed Central PMCID: PMC3315093. eng
Lo Iacono M, Monica V, Saviozzi S, Ceppi P, Bracco E, Papotti M et al (2011) Aurora kinase a expression is associated with lung cancer histological-subtypes and with tumor de-differentiation. J Transl Med 9:100, PubMed PMID: 21718475. Pubmed Central PMCID: PMC3148570. eng
Lassus H, Staff S, Leminen A, Isola J, Butzow R (2011) Aurora-A overexpression and aneuploidy predict poor outcome in serous ovarian carcinoma. Gynecol Oncol 120(1):11–17, PubMed PMID: 20937525. eng
Zhou H, Kuang J, Zhong L, Kuo WL, Gray JW, Sahin A et al (1998) Tumour amplified kinase STK15/BTAK induces centrosome amplification, aneuploidy and transformation. Nat Genet 20(2):189–193, PubMed PMID: 9771714. eng
Dar AA, Zaika A, Piazuelo MB, Correa P, Koyama T, Belkhiri A et al (2008) Frequent overexpression of Aurora kinase A in upper gastrointestinal adenocarcinomas correlates with potent antiapoptotic functions. Cancer 112(8):1688–1698, PubMed PMID: 18311783. eng
Chefetz I, Holmberg JC, Alvero AB, Visintin I, Mor G (2011) Inhibition of Aurora-A kinase induces cell cycle arrest in epithelial ovarian cancer stem cells by affecting NFĸB pathway. Cell Cycle 10(13):2206–2214, PubMed PMID: 21623171. Pubmed Central PMCID: PMC3154367. eng
Katayama H, Sasai K, Kawai H, Yuan ZM, Bondaruk J, Suzuki F et al (2004) Phosphorylation by aurora kinase A induces Mdm2-mediated destabilization and inhibition of p53. Nat Genet 36(1):55–62, PubMed PMID: 14702041. eng
Liu Q, Kaneko S, Yang L, Feldman RI, Nicosia SV, Chen J et al (2004) Aurora-A abrogation of p53 DNA binding and transactivation activity by phosphorylation of serine 215. J Biol Chem 279(50):52175–52182, PubMed PMID: 15469940. eng
Hauf S, Cole RW, LaTerra S, Zimmer C, Schnapp G, Walter R et al (2003) The small molecule hesperadin reveals a role for Aurora B in correcting kinetochore-microtubule attachment and in maintaining the spindle assembly checkpoin. J Cell Biol 161(2):281–294, PubMed PMID: 12707311. Pubmed Central PMCID: PMC2172906. eng
Crosio C, Fimia GM, Loury R, Kimura M, Okano Y, Zhou H et al (2002) Mitotic phosphorylation of histone H3: spatio-temporal regulation by mammalian Aurora kinases. Mol Cell Biol 22(3):874–885, PubMed PMID: 11784863. Pubmed Central PMCID: PMC133550. eng
Goto H, Yasui Y, Nigg EA, Inagaki M (2002) Aurora-B phosphorylates histone H3 at serine28 with regard to the mitotic chromosome condensation. Genes Cells 7(1):11–17, PubMed PMID: 11856369. eng
Hetland TE, Nymoen DA, Holth A, Brusegard K, Flørenes VA, Kærn J et al (2012) Aurora B expression in metastatic effusions from advanced-stage ovarian serous carcinoma is predictive of intrinsic chemotherapy resistance. Hum Pathol. 5:777–85, PubMed PMID: 23114921. ENG
Hartsink-Segers SA, Zwaan CM, Exalto C, Luijendijk MW, Calvert VS, Petricoin EF et al (2012) Aurora kinases in childhood acute leukemia: the promise of aurora B as therapeutic target. Leukemia. 3:560–8, PubMed PMID: 22940834. ENG
Buczkowicz P, Zarghooni M, Bartels U, Morrison A, Misuraca KL, Chan T, et al. (2012) Aurora kinase B is a potential therapeutic target in pediatric diffuse intrinsic pontine glioma. Brain Pathol. 3:244–53, PubMed PMID: 22971244. ENG
Gully CP, Velazquez-Torres G, Shin JH, Fuentes-Mattei E, Wang E, Carlock C et al (2012) Aurora B kinase phosphorylates and instigates degradation of p53. Proc Natl Acad Sci U S A 109(24):E1513–E1522, PubMed PMID: 22611192. Pubmed Central PMCID: PMC3386093. eng
Hata T, Furukawa T, Sunamura M, Egawa S, Motoi F, Ohmura N et al (2005) RNA interference targeting aurora kinase a suppresses tumor growth and enhances the taxane chemosensitivity in human pancreatic cancer cells. Cancer Res 65(7):2899–2905, PubMed PMID: 15805292. eng
Warner SL, Munoz RM, Stafford P, Koller E, Hurley LH, Von Hoff DD et al (2006) Comparing Aurora A and Aurora B as molecular targets for growth inhibition of pancreatic cancer cells. Mol Cancer Ther 5(10):2450–2458, PubMed PMID: 17041088. eng
Girdler F, Gascoigne KE, Eyers PA, Hartmuth S, Crafter C, Foote KM et al (2006) Validating aurora B as an anti-cancer drug target. J Cell Sci 119(Pt 17):3664–3675, PubMed PMID: 16912073. eng
Gizatullin F, Yao Y, Kung V, Harding MW, Loda M, Shapiro GI (2006) The Aurora kinase inhibitor VX-680 induces endoreduplication and apoptosis preferentially in cells with compromised p53-dependent postmitotic checkpoint function. Cancer Res 66(15):7668–7677, PubMed PMID: 16885368. eng
Kuner R, Fälth M, Pressinotti NC, Brase JC, Puig SB, Metzger J et al (2012) The maternal embryonic leucine zipper kinase (MELK) is upregulated in high-grade prostate cancer. J Mol Med (Berl). 2:237–48, PubMed PMID: 22945237. ENG
Marie SK, Okamoto OK, Uno M, Hasegawa AP, Oba-Shinjo SM, Cohen T et al (2008) Maternal embryonic leucine zipper kinase transcript abundance correlates with malignancy grade in human astrocytomas. Int J Cancer 122(4):807–815, PubMed PMID: 17960622. eng
Pickard MR, Green AR, Ellis IO, Caldas C, Hedge VL, Mourtada-Maarabouni M et al (2009) Dysregulated expression of Fau and MELK is associated with poor prognosis in breast cancer. Breast Cancer Res 11(4):60, PubMed PMID: 19671159. Pubmed Central PMCID: PMC2750122. eng
Nakano I, Paucar AA, Bajpai R, Dougherty JD, Zewail A, Kelly TK et al (2005) Maternal embryonic leucine zipper kinase (MELK) regulates multipotent neural progenitor proliferation. J Cell Biol 170(3):413–427, PubMed PMID: 16061694. Pubmed Central PMCID: PMC2171475. eng
Nakano I, Masterman-Smith M, Saigusa K, Paucar AA, Horvath S, Shoemaker L et al (2008) J Neurosci Res 86(1):48–60, PubMed PMID: 17722061. eng
Hebbard LW, Maurer J, Miller A, Lesperance J, Hassell J, Oshima RG et al (2010) Maternal embryonic leucine zipper kinase is upregulated and required in mammary tumor-initiating cells in vivo. Cancer Res 70(21):8863–8873, PubMed PMID: 20861186. eng
Badouel C, Körner R, Frank-Vaillant M, Couturier A, Nigg EA, Tassan JP (2006) M-phase MELK activity is regulated by MPF and MAPK. Cell Cycle 5(8):883–889, PubMed PMID: 16628004. eng
Badouel C, Chartrain I, Blot J, Tassan JP (2010) Maternal embryonic leucine zipper kinase is stabilized in mitosis by phosphorylation and is partially degraded upon mitotic exit. Exp Cell Res 316(13):2166–2173, PubMed PMID: 20420823. eng
Choi S, Ku JL (2011) Resistance of colorectal cancer cells to radiation and 5-FU is associated with MELK expression. Biochem Biophys Res Commun 412(2):207–213, PubMed PMID: 21806965. eng
Liu G, Yuan X, Zeng Z, Tunici P, Ng H, Abdulkadir IR et al (2006) Analysis of gene expression and chemoresistance of CD133+ cancer stem cells in glioblastoma. Mol Cancer 5:67, PubMed PMID: 17140455. Pubmed Central PMCID: PMC1697823. eng
Nakano I, Joshi K, Visnyei K, Hu B, Watanabe M, Lam D et al (2011) Siomycin A targets brain tumor stem cells partially through a MELK-mediated pathway. Neuro Oncol 13(6):622–634, PubMed PMID: 21558073. Pubmed Central PMCID: PMC3107094. eng
Klebig C, Korinth D, Meraldi P (2009) Bub1 regulates chromosome segregation in a kinetochore-independent manner. J Cell Biol 185(5):841–858, PubMed PMID: 19487456. Pubmed Central PMCID: PMC2711590. eng
Janssen A, Kops GJ, Medema RH (2009) Elevating the frequency of chromosome mis-segregation as a strategy to kill tumor cells. Proc Natl Acad Sci U S A 106(45):19108–19113, PubMed PMID: 19855003. Pubmed Central PMCID: PMC2776415. eng
Kwiatkowski N, Jelluma N, Filippakopoulos P, Soundararajan M, Manak MS, Kwon M et al (2010) Small-molecule kinase inhibitors provide insight into Mps1 cell cycle function. Nat Chem Biol 6(5):359–368, PubMed PMID: 20383151. Pubmed Central PMCID: PMC2857554. eng
Hewitt L, Tighe A, Santaguida S, White AM, Jones CD, Musacchio A et al (2010) Sustained Mps1 activity is required in mitosis to recruit O-Mad2 to the Mad1-C-Mad2 core complex. J Cell Biol 190(1):25–34, PubMed PMID: 20624899. Pubmed Central PMCID: PMC2911659. eng
Li M, York JP, Zhang P (2007) Loss of Cdc20 causes a securin-dependent metaphase arrest in two-cell mouse embryos. Mol Cell Biol 27(9):3481–3488, PubMed PMID: 17325031. Pubmed Central PMCID: PMC1899968. eng
Manchado E, Guillamot M, de Cárcer G, Eguren M, Trickey M, García-Higuera I et al (2010) Argeting mitotic exit leads to tumor regression in vivo: modulation by Cdk1, Mastl, and the PP2A/B55α,δ phosphatase. Cancer Cell 18(6):641–654, PubMed PMID: 21156286. eng
Ye X, Zhu C, Harper JW (2001) A premature-termination mutation in the Mus musculus cyclin-dependent kinase 3 gene. Proc Natl Acad Sci U S A 98(4):1682–1686, PubMed PMID: 11172011. Pubmed Central PMCID: PMC29317. eng
Tsutsui T, Hesabi B, Moons DS, Pandolfi PP, Hansel KS, Koff A et al (1999) Targeted disruption of CDK4 delays cell cycle entry with enhanced p27(Kip1) activity. Mol Cell Biol 19(10):7011–7019, PubMed PMID: 10490638. Pubmed Central PMCID: PMC84696. eng
Atanasoski S, Shumas S, Dickson C, Scherer SS, Suter U (2001) Differential cyclin D1 requirements of proliferating Schwann cells during development and after injury. Mol Cell Neurosci 18(6):581–592, PubMed PMID: 11749035. eng
Ciemerych MA, Sicinski P (2005) Cell cycle in mouse development. Oncogene 24(17):2877–2898, PubMed PMID: 15838522. eng
Fantl V, Stamp G, Andrews A, Rosewell I, Dickson C (1995) Mice lacking cyclin D1 are small and show defects in eye and mammary gland development. Genes Dev 9(19):2364–2372, PubMed PMID: 7557388. eng
Sicinski P, Donaher JL, Parker SB, Li T, Fazeli A, Gardner H et al (1995) Cyclin D1 provides a link between development and oncogenesis in the retina and breast. Cell 82(4):621–630, PubMed PMID: 7664341. eng
Georgia S, Bhushan A (2004) Beta cell replication is the primary mechanism for maintaining postnatal beta cell mass. J Clin Invest 114(7):963–968, PubMed PMID: 15467835. Pubmed Central PMCID: PMC518666. eng
Sicinski P, Donaher JL, Geng Y, Parker SB, Gardner H, Park MY et al (1996) Cyclin D2 is an FSH-responsive gene involved in gonadal cell proliferation and oncogenesis. Nature 384(6608):470–474, PubMed PMID: 8945475. Eng
Sicinska E, Aifantis I, Le Cam L, Swat W, Borowski C, Yu Q et al (2003) Requirement for cyclin D3 in lymphocyte development and T cell leukemias. Cancer Cell 4(6):451–461, PubMed PMID: 14706337. eng
Ciemerych MA, Kenney AM, Sicinska E, Kalaszczynska I, Bronson RT, Rowitch DH et al (2002) Development of mice expressing a single D-type cyclin. Genes Dev 16(24):3277–3289, PubMed PMID: 12502747. Pubmed Central PMCID: PMC187507. Eng
Kozar K, Ciemerych MA, Rebel VI, Shigematsu H, Zagozdzon A, Sicinska E et al (2004) Mouse development and cell proliferation in the absence of D-cyclins. Cell 118(4):477–491, PubMed PMID: 15315760. eng
Geng Y, Yu Q, Sicinska E, Das M, Schneider JE, Bhattacharya S et al (2003) Cyclin E ablation in the mouse. Cell 114(4):431–443, PubMed PMID: 12941272. eng
Parisi T, Beck AR, Rougier N, McNeil T, Lucian L, Werb Z et al (2003) Cyclins E1 and E2 are required for endoreplication in placental trophoblast giant cells. EMBO J 22(18):4794–4803, PubMed PMID: 12970191. Pubmed Central PMCID: PMC212738. eng
Liu D, Matzuk MM, Sung WK, Guo Q, Wang P, Wolgemuth DJ (1998) Cyclin A1 is required for meiosis in the male mouse. Nat Genet 20(4):377–380, PubMed PMID: 9843212. eng
Liu D, Liao C, Wolgemuth DJ (2000) A role for cyclin A1 in the activation of MPF and G2-M transition during meiosis of male germ cells in mice. Dev Biol 224(2):388–400, PubMed PMID: 10926775. eng
Brandeis M, Rosewell I, Carrington M, Crompton T, Jacobs MA, Kirk J et al (1998) Cyclin B2-null mice develop normally and are fertile whereas cyclin B1-null mice die in utero. Proc Natl Acad Sci U S A 95(8):4344–4349, PubMed PMID: 9539739. Pubmed Central PMCID: PMC22491. eng
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Alexander, A., Keyomarsi, K. (2014). Exploiting Cell Cycle Pathways in Cancer Therapy: New (and Old) Targets and Potential Strategies. In: Kumar, R. (eds) Nuclear Signaling Pathways and Targeting Transcription in Cancer. Cancer Drug Discovery and Development. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4614-8039-6_14
Download citation
DOI: https://doi.org/10.1007/978-1-4614-8039-6_14
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4614-8038-9
Online ISBN: 978-1-4614-8039-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)