Abstract
One of the main causes of the relatively low reproductive success rates seen in humans is numerical chromosome abnormalities (aneuploidy). The advent of in vitro fertilisation (IVF) and associated diagnostic methods such as preimplantation genetic diagnosis and screening enabled a detailed investigation of the chromosome content of human gametes and embryos. Such investigations clearly showed that the incidence of numerical chromosome abnormalities was high and as would be expected had an adverse impact on the progress of embryo development and viability.
Embryonic aneuploidy can have a meiotic origin with chromosome malsegregation taking place during gametogenesis, and a post-zygotic origin with abnormalities occurring after fertilisation. Direct analysis of the human female gamete using a variety of cytogenetic methods has shown that the majority of meiotically derived aneuploidies arise during oogenesis. The close relationship between advancing female age and increasing aneuploidy rates was also confirmed, as more than 50 % of oocytes generated by women over the age of 40 years were demonstrated to be chromosomally abnormal. Graphs illustrating the changing oocyte aneuploidy rate with advancing maternal age are a mirror image of those showing the declining implantation rate of IVF embryos with age, suggesting that the increase in meiotic errors might explain the reduced success rate of IVF treatments for women in their late 30s and 40s.
The contribution of aneuploidy of male meiotic origin to the embryo is not as clear. Several studies have been carried out, and it was concluded that the incidence of chromosome anomalies ranged between 3 and 5 % in the sperm of fertile men, but significantly (approximately threefold) increased in the sperm obtained by infertile men.
Studies investigating two different preimplantation development stages, namely the cleavage stage and blastocyst stage, have suggested that post-zygotic chromosome anomalies take place even more frequently compared to the meiotic ones, especially at the cleavage stage. A result of post-zygotic anomalies is chromosome mosaicism, the presence of two or more karyotypically distinct cell lines within the same embryo.
During this chapter, the data obtained from cytogenetic investigations of human oocytes, sperm and also embryos at the cleavage and blastocyst stage of preimplantation development will be summarised. In addition, the different mechanisms leading to aneuploidy of meiotic and post-zygotic origin will be described and their frequencies and impact on embryonic survival will be discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Comparative Genomic Hybridisation
- Preimplantation Genetic Diagnosis
- Inner Cell Mass
- Whole Genome Amplification
- Aneuploidy Rate
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Humans as a species are not as efficient in reproducing compared to other mammals. Specifically, a couple of proven fertility will achieve a viable pregnancy in only 20–25 % of all natural cycles [1, 2]. This rate decreases during in vitro fertilisation (IVF) cycles as only approximately 13 % of all morphologically normal embryos transferred will go on to produce a live birth [3]. One of the main causes of these relatively low reproductive success rates seen in humans is numerical chromosome abnormalities (aneuploidy). The advent of IVF and associated diagnostic methods such as preimplantation genetic diagnosis (PGD) and screening (PGS) enabled a detailed investigation of the chromosome content of human gametes and embryos. Such investigations clearly showed that the incidence of numerical chromosome abnormalities was high and as would be expected had an adverse impact on the progress of embryo development and viability [4, 5].
Embryonic aneuploidy can have a meiotic origin with chromosome malsegregation taking place during gametogenesis, and a post-zygotic origin with abnormalities occurring after fertilisation. Direct analysis of the human female gamete using a variety of cytogenetic methods has shown that the majority of meiotically derived aneuploidies arise during oogenesis. The close relationship between advancing female age and increasing aneuploidy rates was also confirmed, as more than 50 % of oocytes generated by women over the age of 40 years were demonstrated to be chromosomally abnormal [6–12]. Graphs illustrating the changing oocyte aneuploidy rate with advancing maternal age are a mirror image of those showing the declining implantation rate of IVF embryos with age, suggesting that the increase in meiotic errors might explain the reduced success rate of IVF treatments for women in their late 30s and 40s.
The contribution of aneuploidy of male meiotic origin to the embryo is not as clear. Several studies have been carried out and examined the chromosome complement of sperm generated by fertile as well as infertile men. What was concluded was that the incidence of chromosome anomalies ranged between 3 and 5 % in the sperm of fertile men, but significantly (approximately threefold) increased in the sperm obtained by infertile men [13].
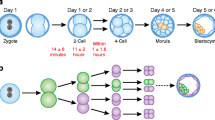
Two different preimplantation development stages have been examined in order to establish the frequency of aneuploidy arising after fertilisation. These are the cleavage stage, which is reached 3 days after fertilisation when the embryo consists of 6–8 totipotent cells called blastomeres, and the blastocyst stage. Embryos become blastocysts 5–6 days after fertilisation and consist of two different cell types, the outer trophectoderm (TE) and the inner cell mass (ICM). Studies investigating these two different preimplantation development stages have suggested that post-zygotic chromosome anomalies take place even more frequently compared to the meiotic ones, especially at the cleavage stage [14–17]. A result of post-zygotic anomalies is chromosome mosaicism, the presence of two or more karyotypically distinct cell lines within the same embryo.
During this chapter, the data obtained from cytogenetic investigations of human oocytes, sperm and also embryos at the cleavage and blastocyst stage of preimplantation development will be summarised. In addition, the different mechanisms leading to aneuploidy of meiotic and post-zygotic origin will be described and their frequencies and impact on embryonic survival will be discussed.
Aneuploidy of Female Meiotic Origin
It has been demonstrated that the vast majority of meiotic chromosome abnormalities in the embryo originate from the oocyte. As they mature, both the male and female gametes begin a specialised cellular division called meiosis. As a result, the diploid chromosome number is reduced by half, leading to the formation of haploid gametes.
Meiosis is separated into two different divisions, meiosis I (MI) and II (MII). The whole process starts with the replication of DNA during S phase, leading to the generation of two sister chromatids. During MI the homologous chromosomes align, and exchange of material between the chromatids of different homologues may occur. At the end of MI, bivalents separate, with homologues travelling to opposite poles of the meiotic spindle, one entering the first polar body (PB) while the other stays in the oocyte. During MI, sister chromatids are held together, and it is not until MII that they are finally separated, one of the chromatids passing into the second PB, the other remaining in the oocyte. The end result is the generation of haploid gametes, sperm cells in the male and an oocyte and two PBs in the female. Figure 1 describes the normal segregation of chromosomes during meiosis.
Normal chromosome segregation during meiosis. Meiosis is made up of two divisions, MI and MII. During prophase I, the 46 chromosomes condense and form 23 homologous pairs of bivalents. Formation of chiasmata and exchange of genetic material between homologues follows. During metaphase I, these bivalents align on the metaphase plate orientated by attachment to the spindle. They then disjoin and segregate to the resulting daughter cells at anaphase I and telophase I. MI is a reduction division, whereas MII is a mitotic type of division during which chromosomes align once again on the metaphase II spindle. Separation and segregation of sister chromatids to opposite poles follows at anaphase II and telophase II. Cytokinesis produces four haploid products. (From Fragouli E, Wells D, Delhanty JD. Chromosome abnormalities in the human oocyte. Cytogenet Genome Res. 2011;133 (2–4):107–18, with permission.)
There are three different stages during which a chromosome could malsegregate and cause aneuploidy in the embryo. These are the initial set of premeiotic mitotic divisions, MI and MII. Two different mechanisms of oocyte chromosome malsegregation have been described, whole chromosome nondisjunction and unbalanced chromatid predivision.
Whole chromosome nondisjunction occurs in MI as well as MII and is the result of homologous chromosomes not segregating to opposite poles of the meiotic spindle, but instead moving together towards the same pole. As a result, an oocyte with an extra chromosome is generated whereas the corresponding PB (either the first or second) is missing this chromosome, or vice versa. Risk factors associated with this aneuploidy-causing mechanism depend on the chromosome and the meiotic division, but generally include the number and positioning of the chiasmata in relation to the centromere (too proximal or too distal). It is also believed that female age has differing effects on the various recombination patterns predisposing to nondisjunction [11, 18].
Unbalanced chromatid predivision was described when a group of researchers in their analysis of metaphase II oocytes noticed that some cells consisted of an extra one or two single chromatids [19, 20]. This observation led them to suggest that imbalance can result from premature division (predivision) of the chromosome centromere after which chromatids segregate at random, potentially moving to the same pole at anaphase of MII [19, 20]. Both whole chromosome nondisjunction and unbalanced chromatid predivision are illustrated in Figs. 2 and 3 respectively.
Whole chromosome non-disjunction. The first aneuploidy-causing mechanism could take place during MI (a) and also MII (b). It involves the segregation of entire chromosomes towards the same pole of the meiotic spindle which leads to the formation of disomic and nullisomic gametes. (From Fragouli E, Wells D, Delhanty JD. Chromosome abnormalities in the human oocyte. Cytogenet Genome Res. 2011;133 (2–4):107–18, with permission.)
Unbalanced chromatid predivision. This second aneuploidy-causing mechanism could take place only during MI and involves the premature separation of a chromosome into its sister chromatids. These are subsequently distributed at random during anaphase I. Unbalanced chromatid predivision could have as an effect the formation of gametes which have either an extra or a missing chromatid. (From Fragouli E, Wells D, Delhanty JD. Chromosome abnormalities in the human oocyte. Cytogenet Genome Res. 2011;133 (2–4):107–18, with permission.)
Both whole chromosome and single chromatid malsegregation have been shown to be active and responsible for anomalies of female meiotic origin. The difference between them is that whole chromosome nondisjunction will produce an aneuploid oocyte and subsequent embryo in all cases, whereas a single chromatid abnormality will result to embryonic aneuploidy in 50 % of cases, depending initially on whether the unattached chromatids segregate to opposite spindle poles or move together towards the same pole, and finally on events at anaphase II after fertilisation. As with whole chromosome nondisjunction, absent or altered recombination, combined with advancing maternal age, predispose oocytes to become aneuploid via random segregation of single chromatids.
To elucidate the causal mechanisms of maternally derived aneuploidy, researchers employed several different cytogenetic methods to examine human oocytes and/or their corresponding PBs. The following sections will summarise findings for MI and MII obtained with the use of classical and molecular cytogenetic methods.
Meiosis I
Classical cytogenetic techniques such as G and R-banding, along with various molecular methods such as fluorescence in situ hybridisation (FISH), spectral karyotyping (SKY) and comparative genomic hybridisation [metaphase, (CGH) or array (aCGH)] have been employed for the study of MI. For this purpose, oocytes arrested at the metaphase II stage of development and, whenever available, their corresponding first PBs have been examined.
The largest data set of karyotyped metaphase II oocytes was reported by Pellestor et al. [8]. Specifically, a total of 1,397 oocytes were analysed with the use of R-banding. These were donated by 792 women undergoing IVF with an average age of ∼34 years (age range 19–46 years). Of all the examined oocytes, chromosome errors were scored in 10.8 %. This investigation confirmed the expected relationship between advancing female age and global aneuploidy rate. It was also concluded that advancing age had a more pronounced effect on the predivision and unbalanced segregation of single chromatids [8]. This study along with data obtained from similar investigations clearly showed that smaller chromosomes, belonging to groups D–G, were more frequently participating in aneuploidy events [8, 21–24]. The poor morphology of oocyte metaphase preparations, however, made the exact identification of individual chromosomes difficult. The use of a different molecular cytogenetic method, FISH, solved this problem.
FISH is advantageous over classical karyotyping methods because it can yield results regardless of the quality of the cell under analysis. Using FISH, it was not only possible to examine the chromosomes of the metaphase II oocyte, but also those of the even more challenging first PB. The chromosomes targeted were selected on the basis of data obtained from the analysis of miscarriages and abnormal live births, and generally included smaller size ones (starting from 13 onwards).
A combination of the larger chromosomes 1, 9, 12 and X and the smaller 13, 16, 18 and 21 were investigated during the course of two different studies carried out by our group [25, 26]. A three-round FISH protocol was employed and a total of 236 spare, mostly unfertilised oocytes and their corresponding first PBs were analysed. These were donated by 124 patients, whose average maternal age was 32.5 years (age range 22–44 years). Only gains of chromosomes were scored, as it was considered that chromosome loss could be due to the spreading of the oocyte and/or PB on the microscope slide. The hyperploidy rate was calculated to be approximately 4 %, the smaller chromosomes 13, 16, 18 and 21 were mostly affected, and both whole chromosome nondisjunction and unbalanced chromatid predivision were identified to be active. Moreover, a third aneuploidy-causing mechanism was observed, involving the presence of a trisomic cell line in the gonads of some patients (germinal/gonadal mosaicism) [25, 26].
The use of alternative cytogenetic methods such as SKY enabled the analysis of all 23 chromosomes of human oocytes. Sandalinas and colleagues employed SKY to examine 47 fresh non-inseminated metaphase II oocytes, donated by 16 patients with a female age range of 24–48 years [6]. A total of 9 whole chromosome nondisjunction events were observed, along with 11 errors involving single chromatids. Chromo-somes 9, 11, 12, 14, 18, 20, 21 and 22 were affected. More importantly, 12 different oocytes with balanced chromatid predivision were observed. It was therefore confirmed that this type of abnormality is real and not an artefact of oocyte ageing in culture, and it is closely associated with advancing female age and decreasing chromosome size [6].
Even though classical karyotyping, FISH and SKY are all capable of accurately examining the chromosomes of human oocytes, their main disadvantage is that they all require the spreading of a single cell arrested in metaphase on a microscope slide, risking in this way the artefactual loss of chromosomes. Clearly an alternative cytogenetic method, which could not only avoid spreading but could examine all 23 chromosomes as well, was needed. Ultimately, this was achieved by a combination of whole genome amplification (WGA) and CGH [27, 28]. CGH is related to FISH and employs simultaneous hybridisation of differentially labelled DNA samples (sample DNA: green; chromosomally normal reference DNA: red) to normal metaphase chromosomes. The ratio of green:red fluorescence along the length of each chromosome indicates whether there has been any gain or loss of chromosomal material in the test sample.
The first application of this approach took place in a clinical context, and involved the chromosome complement analysis of ten first PBs biopsied from oocytes generated by a 40-year old IVF patient with a history of repeated implantation failure (RIF) [29]. CGH was further validated and used to examine larger sets of metaphase II oocytes and their corresponding first PBs, by our group and others [9, 10]. During our investigation, 107 oocyte-PB complexes were analysed via CGH. These were donated by 46 women who were undergoing routine IVF procedures, and whose average age was 32.5 years (range 18–42 years). It was evident from the obtained results that aneuploidy frequently affected even the larger chromosomes (1–12), as well as the smaller ones. Both aneuploidy-causing mechanisms, i.e. whole chromosome nondisjunction and unbalanced predivision of single chromatids were observed, and the resulting aneuploidy rate was 22 %. Interestingly, the larger chromosomes (1–12) participated solely in whole chromosome nondisjunction events, whereas a combination of whole chromosome and single chromatid anomalies were seen for smaller chromosomes (13 onwards) and chromosome X. CGH was also capable of accurately detecting structural abnormalities resulting from chromosome breakage. Additionally, evidence of age-independent factors influencing aneuploidy of MI origin was provided, with certain patients exhibiting unexpectedly high frequencies of abnormal oocytes [10].
In summary, studies using a variety of cytogenetic methods to examine human oocytes have uniformly shown that even though all chromosomes can be affected by MI malsegregation errors, there seems to be a preferential involvement of the smaller ones. Combining data obtained in a research and clinical setting suggests that for an average female age of 32 years, the expected MI aneuploidy rate is in the range of 22 % and can increase to over 45 % for women of an average age of 40 years or more [9, 10, 30, 31]. Table 1 shows a summary of results obtained during the analysis of metaphase II oocytes using methods such as karyotyping, FISH and CGH.
Meiosis II
Mature oocytes are arrested at the metaphase stage of the second meiotic division (MII). At this stage oocytes consist of 23 chromosomes with sister chromatids held together at the centromeres. Fertilisation leads to resumption of MII, followed by centromere separation. As a result, 23 chromatids remain in the fertilised oocyte, while the other 23 move to the second PB, which is extruded soon after oocyte activation. As it would be difficult to directly examine the fertilised oocyte, data on the incidence of MII aneuploidy has been obtained almost exclusively from the investigation of the second PB. FISH, CGH and recently aCGH, have all been used to examine MII [7, 12, 32, 33].
Verlinsky and colleagues have reported FISH results from large numbers of first as well as second PBs [7, 32]. Five different chromosomes were targeted, namely 13, 16, 18, 21 and 22 in a single hybridisation round. It was shown for the first time that women of advanced reproductive age generate oocytes carrying an almost equal amount of MI and MII chromosomes errors (41.8 % and 37.3 % respectively). They also reported findings on the meiotic origin of abnormalities affecting chromosomes 16 and 18 that contradict data obtained from analysis of spontaneous abortion material. Specifically, such studies demonstrated that most abnormalities affecting chromosome 16 have an MI origin, and those affecting 18 originate during MII [34–36], whereas the complete opposite was observed from the FISH analysis of first and second PBs. Another important finding was that almost one third (32.5 %) of oocytes classified as abnormal due to an MI chromatid error led to the formation of apparently normal zygotes, following segregation of the chromatid to the second PB in MII [7, 32].
FISH analysis of second PBs provided a valuable insight into the frequency and type of MII chromosome malsegregation errors. However, as with MI, PBs were spread on microscope slides, risking the artefactual loss of chromosome material, and only 5 of the 23 chromosomes were examined. In a more recent investigation, CGH was applied to analyse first and second PBs biopsied from a total of 117 zygotes generated by couples with a poor reproductive history [12]. The total meiotic aneuploidy rate was 65.5 % and this was seen for an average maternal age of 39.8 years. It was evident from the obtained results that anomalies of MII origin occurred slightly more frequently, compared to those taking place during MI (45.8 % and 36.5 % respectively), and as with the FISH data, the mechanism of MII ‘correction’ of MI chromatid errors was also observed. Combining the MI and MII cytogenetic data predicted that 60 % of abnormalities would lead to a trisomy in the resulting embryo. The fact that chromosome losses in the PBs were more frequent than gains suggests that anaphase lag or chromosome failure to congress on the spindle is a significant aneuploidy-causing mechanism. Additionally, it was demonstrated that MII malsegregation errors affected chromosomes from all groups, but the smaller 16, 21, 22, 15 and 19 were found to more frequently participate in aneuploidy events [12]. A summary of the data obtained from the cytogenetic analysis of first and second PBs is shown on Table 2.
It can therefore be concluded that both MI and MII are prone to mistakes in chromosome segregation, at least in older women. Additionally, even though aneuploidy events can affect the larger chromosomes, it is the smaller ones that seem to be preferentially involved during both meiotic divisions. It is also very clear that advancing female age has a significant adverse effect on both meiotic divisions, but MII may be more susceptible to age-related errors than MI.
Aneuploidy of Male Meiotic Origin
Estimates of sperm aneuploidy have been arrived at by two methods. First, by fusing sperm with hamster eggs when all the chromosomes may be visualised; secondly by FISH analysis of the nucleus when a few chromosomes at a time may be analysed. The first method is very labour intensive, the second less so, but many thousand sperm need to be analysed to obtain a reliable result. Templado and colleagues reviewed aneuploidy levels in healthy men from 30 FISH studies that used a minimum of five donors and employed strict scoring criteria and obtained a lower estimate of total aneuploidy of 4.5 % (2× disomy of 2.26 %) [35]. This figure was more than double that found by sperm karyotyping. Disomy for the autosomes is about 0.1 % on average but ranges between 0.03 (chromosome 8) and 0.47 (chromosome 22). Most authors investigating a limited range of chromosomes found that chromosome 21 (0.17 %) has an increased disomy frequency and that the sex chromosomes at 0.27 % have the highest, reflecting the significant male component in causing Down and Klinefelter syndromes and the sole male origin of the XYY cases. However, the studies revealed a considerable level of interindividual variability making it clear that certain men are far more prone to the production of sperm with unbalanced chromosomes than others. A doubling of sperm disomy for the chromosome pair involved was seen in fathers of offspring with Down, Turner and Klinefelter syndromes which had been shown to be of paternal origin [35]. Variability has also been observed in male meiotic cells; the frequency of unpaired sex chromosomes at MI in spermatocytes ranged between 3.2 and 43.7 % in fertile men [37]. The rate of sex chromosome abnormalities seen at MI of meiosis (mean 27.4 %) is far higher than the 0.5 % seen in spermatozoa. This difference can be explained by the arrest of chromosomally abnormal (asynapsed) cells by the activation of meiotic checkpoints that operate with much greater efficiency in males than in females [38].
How does the mechanism of origin of sperm aneuploidy compare with that in female gametes? Clearly errors in chromosome segregation, as in females, may result from failure to pair properly initially, altered or reduced recombination, or failure to maintain chromatid cohesion beyond anaphase I of meiosis. As with autosomal trisomies, sex chromosome trisomy is clearly linked with reduced recombination of the XY bivalent [11]. Recently, for the first time, the two types of nondisjunction seen in oocytes, that involving whole chromosomes and premature separation of chromatids, have been seen in metaphase I and II of male meiosis [39]. Premature separation of chromatids was the more frequent type in that study, seen in MII spermatocytes as extra or missing chromatids of chromosomes 21, X or Y. Very recently, evidence for the existence of a postmeiotic checkpoint that monitors numerical abnormalities was found by the application of Multiplex fluorescence in situ hybridisation (M-FISH) to metaphase I and II spermatocytes from three fertile donors [40]. This technique allows identification of each and every chromosome by a combination of different fluorochromes. As expected, the frequency of numerical abnormalities (disomy) at MII (14.5 %) was found to be far higher than that seen at MI (trisomy) (1.3 %). A significant proportion of spermatocytes at MI (27.7 %) were seen to display a low chiasma count, with small chromosomes present as two univalents. The chromosomes most frequently involved were X, Y and 21 and these were also those most frequently exhibiting disomy at MII.
The conclusion was that achiasmate nondisjunction of whole chromosomes and premature separation of chromatids appear as the main mechanisms generating aneuploidy in human male meiosis and that both contribute equally. Since the level of disomy observed in spermatocytes II is about threefold higher than that seen in spermatozoa the existence of a postmeiotic checkpoint is postulated [40]. The detection of spermatocytes II with separated sister chromatids is the first evidence that balanced predivision, seen in fresh oocytes, also occurs in male meiosis and may lead to aneuploid sperm by random segregation [6]. In conclusion, it appears that there are many similarities between the mechanisms leading to aneuploidy in the two sexes; however in contrast to the situation in females, it has not been possible to obtain definitive evidence for a paternal age effect [41].
Sperm Aneuploidy and Male Infertility
About 5 % of infertile men have a chromosomal abnormality; these subjects will clearly be at increased risk of producing aneuploid sperm. However, the efficiency of the meiotic cell cycle checkpoints that detect unpaired segments of DNA ensures that most prospective aneuploid gametes undergo apoptosis; a lowered sperm count is the result. In men with a normal somatic karyotype, an inverse correlation between sperm aneuploidy and sperm concentration has been well documented [42]. Analysis of sperm chromosomal content in 46 male carriers of reciprocal translocations shows that a mean of 40 % of sperm have a normal or balanced complement (by FISH) with a range between 19 and 77 % [43]. The risk is highly dependent upon the type of chromosomes involved (acrocentric vs. metacentric) and the position of the breakpoints. In contrast the sperm of Robertsonian translocation carriers have a majority of normal or balanced types, mean 85 %, range 60–96 % [44]. Couples where the male is the carrier of a chromosomal rearrangement may opt for PGD if they are unsuccessful in achieving a normal pregnancy, but follow-up studies on embryos not transferred clearly show that post-zygotic mitotic anomalies are at least as important as errors in meiotic segregation in causing embryonic aneuploidy [45, 46]. The high proportion of errors developing during cleavage in embryos from such couples is probably related to their reproductive failure.
Germline Mosaicism: Inherited Aneuploidy
Germinal or germline mosaicism may be defined as genetic abnormality of premeiotic origin in an otherwise normal organism. It can be the result of mosaicism that affects a proportion of the premeiotic germ cells and will thus lead to recurrent genetic abnormality of the same type in the offspring. This type could also be described as gonadal mosaicism. Alternatively, if the error occurred during one of the mitotic divisions prior to the onset of meiosis it could be confined to a single gamete. The term germline mosaicism covers both types. It has generally been considered to be a rare phenomenon.
Evidence for the existence of chromosomal germline mosaicism is obtained from both genetic and cytological studies. Recurrent aneuploid conceptions of the same type may be due to chance, especially if the mother is of advanced age [47]. However, careful analysis using both karyotyping and molecular studies in 151 families with Down syndrome offspring identified 8 families with germline mosaicism. In all cases the mother was younger than 35 years. Thus the prevalence of germinal mosaicism in young couples with a Down syndrome child was estimated to be 5.3 % [48].
Mosaicism for Chromosomal Rearrangements
Most cases of this type of mosaicism are revealed because of a history of repeated conceptions with similar cytogenetic abnormalities but in other cases a single conception with an unbalanced karyotype can result in the identification of the presence of a low level of balanced cells in a parent. In one interesting case, a couple were referred for PGD with a history of recurrent miscarriage. Two cycles of IVF for PGD were carried out; in all, 4 of 13 embryos were found to have a partial duplication of chromosome 21q. The same duplication was then detected by FISH analysis in 6 % of sperm nuclei, thereby proving paternal gonadal mosaicism for the duplication [49].
Cytological Proof of Germinal Mosaicism
The application of PGD led to the first cytological proof of gonadal mosaicism for trisomy 21 in the case of a couple with normal lymphocyte chromosomes that had a history of 3 conceptions with Down syndrome and one normal child [50]. The couple was referred for PGD; of 7 preimplantation embryos tested 4 had trisomy 21 and 3 of 4 unfertilised oocytes were abnormal. In 3 of the oocytes there was either an extra chromosome or chromatid 21; a crucial observation was that in one oocyte an extra chromatid was found in both the MII and in the first PB. This provided the first direct evidence of a maternal trisomic germ cell line and showed that the extra chromosome 21 had precociously divided into two chromatids before completion of MI.
Studies on Oocytes
The great majority of studies on metaphase II oocytes have been carried out without the analysis of the corresponding first PB. In this situation it is not possible to determine whether any extra chromosomal material present has arisen due to a meiotic error or is caused by a premeiotic trisomic cell. The other option is to analyse oocytes at the MI stage; no studies of this kind have so far been published. Thus it is very difficult to obtain a reliable estimate of the frequency of germinal mosaicism in the human female. Four studies that do provide some information, since oocyte/first PB doublets were analysed, are listed in Table 3. With the exception of Mahmood et al. all studies used either oocytes collected at GV or MI (and allowed to mature to the MII/first PB stage—IVM) or a mixture of these and oocytes that had failed to show evidence of fertilisation after IVF or ICSI [25]. By far the highest frequency of germinal mosaicism (20 %) was seen by Pujol et al.; virtually all of their samples were IVM [51]. For the other three studies, the average frequency was about 6 %. It is to be expected that a higher frequency of this type of aneuploidy will be found in IVM oocytes, since preexisting chromosomal anomalies will lead to delay in completing the cell cycle.
Studies on Spermatocytes
Metaphase I of meiosis is more easily studied in the male. In their study of material from three fertile donors, Uroz and Templado found 1.3 % of MI spermatocytes to be trisomic [40]. Four of the 317 spermatocytes I contained an extra chromosome (18, 19, 22 or Y); this was considered to be caused by preexisting aneuploidy in the germ cells.
Direct Studies on Cells from Foetal Ovaries and Testes
An alternative cytological approach to search for evidence of germinal mosaicism was used by Hultén et al. [52]. Ovarian cells from eight phenotypically normal female foetuses (14–22 weeks gestation) were analysed by FISH using two chromosome 21 specific probes that mapped to different locations. The number of cells studied per case varied between 967 and 2,200 and consisted of premeiotic and stromal cells as well as those of meiotic origin. Mosaicism for trisomy 21 was detected in all the foetal ovaries with frequencies varying between 0.2 and 0.88 % (average 0.54 %), affecting all cell types. This is not an unexpected finding considering that levels of chromosomal mosaicism in human cleavage stage embryos reach 60–70 % even in those of good quality [53]. Interestingly, when investigated by the same group using the same strategy, testicular cells from four normal male foetuses failed to show a single example of trisomy 21 mosaicism [54]. The difference is again likely to be due to the stringent control of the cell cycle during spermatogenesis; testicular cells with additional (unpaired) chromosomes are likely to be arrested in development and to undergo apoptosis [55].
Mechanisms Leading to Aneuploidy in Cases of Germinal Mosaicism
An extra chromosome in a proportion of premeiotic germ cells can lead to aneuploidy via two mechanisms. During prophase of meiosis I, to accommodate the extra chromosome either a trivalent is formed with all three chromosomes associated, or a bivalent plus a univalent [56]. After trivalent formation, secondary (or inevitable) nondisjunction follows when one daughter cell receives a single chromosome and the other receives the remaining two bodies. There is thus a 50 % aneuploidy risk from a trivalent. In the other situation of a bivalent plus univalent, the evidence is that the aneuploidy risk is 100 %. Observations on mammalian meiosis show that the univalent positions itself at MI in a way that the kinetochore of each sister chromatid is orientated towards opposite spindle poles [57, 58]. The chromosome then splits into two chromatids, and one passes to each daughter cell, either the MII oocyte or the first PB, as observed in cases of germinal mosaicism in humans [50]. The combined effect of these two mechanisms is that even a low percentage of premeiotic trisomic cells significantly elevates the risk of an aneuploid conception and consequent embryonic or foetal death or disability, independently of maternal age.
Aneuploidy of Post-zygotic Origin
One day after fertilisation and under the control of the maternal genome, the zygote starts dividing mitotically into smaller cells called blastomeres. To begin with, these cells are spherical and totipotent, but very soon the mitosis becomes asynchronous and totipotency is lost [59]. Embryonic genome activation takes place approximately 3 days after fertilisation, during the cleavage stage of preimplantation development [60]. Compaction follows soon afterwards, which eventually causes the embryo to undergo its first cellular differentiation into TE which will form the extra-embryonic tissues, and ICM which will form the embryo proper. The embryo is now called a blastocyst and this stage of preimplantation development is reached 5–6 days after fertilisation. Prior to implanting, the blastocyst releases enzymes which open a hole in the membrane surrounding it (zona pellucida), causing it to hatch and begin the process of implantation to the maternal uterus [61, 62].
Chromosome abnormalities occur very frequently after fertilisation and have an important contribution to embryonic aneuploidy. The cleavage and blastocyst stages of preimplantation development have been examined cytogenetically and the following sections will summarise the findings obtained during such investigations.
Cleavage Stage
Following fertilisation on day 0, the embryo enters the cleavage stage that lasts until day 3, when it should contain about 8 cells. Cell division during this time interval is thus under the control of stored maternal mRNA transcripts. The development and application of molecular cytogenetic methods for single cell analysis from 1993 onwards allowed the acquisition of data that revealed the full extent of chromosomal anomalies at the cleavage stage. In stark contrast to the situation in postnatal life, 60 % of an average set of embryos created by IVF will contain at least one aneuploid cell by day 3 but the majority will consist mainly of abnormal cells. Prior to the application of molecular methods, traditional karyotyping had been used to analyse cleavage embryos [22]. Abnormality rates between 25 and 40 % were seen with diploid/aneuploid mosaics the most common type. Jamieson et al., found that full chromosome aneuploidy at that stage mirrored the situation in spontaneous abortions, with selective involvement of the smaller chromosomes [61].
As soon as interphase FISH analysis was developed for use in PGD, the karyotyping data was confirmed; aneuploidy and chromosomal mosaicism were common in the spare embryos from PGD cases [4]. Application of the technique to donated IVF embryos in general showed that abnormally developing or arrested embryos are frequently uniformly aneuploid or polyploid while an equal number are mosaic [63, 64]. More unexpected was the finding that normally developing embryos are also often abnormal; although rarely aneuploid throughout, 30–50 % were mosaics [65, 66]. Further data from PGD cycles showed that even fertile couples produced embryos that had highly abnormal chromosome complements, with anomalies affecting several chromosomes, varying in a random fashion from cell to cell, designated chaotic mosaics. Moreover, it became clear that certain couples were prone to the production of such embryos while others produced none of this type [14]. In general, IVF couples will have 5 % of their embryos in the chaotic category but in those with a history of RIF the proportion is much higher [67, 68].
Essentially, it was found that the greater the number of FISH probes used the higher the rate of mosaicism detected, raising the possibility that by day 3 all embryos created by IVF are mosaic. The answer came with another technical advance, that of single cell analysis by CGH. The whole genome had first to be amplified (WGA) to provide sufficient DNA for analysis. The great advantage of CGH is that the copy number of every chromosome is determined and any imbalance detected. WGA and CGH were then applied to all the single cells from a series of good quality cleavage embryos—12 in each series [28, 69]. There was good agreement between the two sets of data; in both sets 25 % were completely euploid with no anomalies detected. Overall, five embryos were aneuploid throughout, with seven the result of a meiotic error. Two thirds were mosaic in both sets of data; the majority were diploid/aneuploid but half of these had at least 50 % of cells abnormal. The new application of CGH allowed the detection of partial aneuploidy, the product of chromosome breakage, confirmed by the appearance of reciprocal products in sister cells in some cases. This was observed in about 10 % of embryos in these series and also later in a much larger diagnostic cohort [70]. If the chromosomal fragments are acentric they will be lost at the next cell division unless they become translocated to another chromosome; the resulting haploinsuffiency will be highly detrimental to the embryo.
When Does Mosaicism Originate and What Is Its Effect?
The evidence indicates that the first three cleavage divisions are the most prone to mitotic errors that lead to mosaicism. Bielanska et al. analysed 216 embryos available after routine IVF treatment and found that the frequency of mosaics increased from 15 % (2–4 cell stage) to 49 % (5–8 cells) and 58 % by day 4 (morula stage) [71]. They further concluded that mosaicism affecting a high proportion of cells, or of a chaotic type, interferes with development of the embryo to blastocyst. Various other publications support this finding. Katz-Jaffe et al. made use of single cell multiplex fluorescent PCR to distinguish between meiotic and mitotic errors in blastomeres from embryos diagnosed with chromosome 21 aneuploidy [16]. Twenty-five embryos were identified to be carrying an error of meiotic origin, while the remaining 13 were mosaic. The timing of the errors was 38 % first division, 31 % second division, and 19 % third division with the others (12 %) an error at MI. Most of the mosaics in fact were derived from diploid zygotes. Overall, evidence suggests that mosaicism interferes with development to a greater degree than uniform aneuploidy. Developmental arrest may affect embryos with fully chaotic or widespread mosaicism but in those with anomalies confined to one chromosome there is likely to be death or a reduction in proliferation of the aneuploid cells. Depending upon the initial proportion of abnormal cells, the rapid growth of the embryo between days 3 and 5 may allow normal cells to predominate and an embryo diagnosed as aneuploid from a single cell on day 3 may be able to lead to a viable pregnancy [72].
Correlation of Aneuploidy Mechanisms with Types of Infertility
Most of the recently published information on the chromosome status of the cleavage stage embryo has come from the analysis of large series of aneuploidy screening (PGS) cases (e.g. Munne et al.) [73]. However, few of these include full follow up of the abnormal embryos after diagnosis. As a result, embryos are designated ‘aneuploid’ based upon the result from a single cell that cannot provide information about the origin of the aneuploidy. This seems to lead to an overestimate of the contribution of meiotic errors. An interesting variation in the proportion of embryos with meiotic errors according to referral reason was found in a comprehensive follow-up study after PGS carried out by our group (Mantzouratou et al. and unpublished data) [67]. A total of 700 embryos that had been diagnosed as abnormal by single cell analysis on day 3 was followed up by applying the diagnostic probe set for six chromosomes (13,15,16,18,21 and 22) to individual cells in the remainder of the embryos on day 5. The recurrent miscarriage group had the highest proportion of meiotic errors, 24 % for an average maternal age of 37.6 years, with the lowest proportion in the RIF group, 8.9 %, average maternal age 36 years. An intermediate value of 20.4 % was found for the advanced maternal age group (over 40 years, average 42). The embryos from couples with RIF were more prone to post-zygotic errors, especially complex errors of the chaotic type. This latter finding was confirmed by another study [68].
Cytogenetic Mechanisms Leading to Mosaicism
Since interphase FISH analysis has a known error rate, the most efficient studies to investigate the mechanisms leading to mosaic aneuploidy have used sets of two probes per chromosome that hybridise to different loci. An early, unpublished study (Conn and Delhanty 1995) used paired probes for chromosomes 13,18 and 21 on cells from 37 very good quality 3-day-old embryos (6–8 cells). Eight embryos were mosaic for these chromosomes; in four cases this was a result of chromosome loss, in one chromosome gain, and in the other three to mitotic nondisjunction (MND). Two probes for each of chromosomes 1,11, and 18 were used in the study by Daphnis et al. mentioned in the section on blastocysts, below [74]. Chromosome loss was again the most common mechanism. Interestingly, the incidence of MND has been found to be maternal age dependent [15].
Blastocyst Stage
Reaching and surviving the blastocyst stage poses a number of potential challenges to the developing embryo. Hence, embryos that successfully achieve this developmental milestone are thought to be of very good quality and have an excellent implantation potential. However, studies using various cytogenetic methods to examine the chromosome complement of blastocysts have shown that both aneuploidy and the phenomenon of mosaicism persist to this final stage of development before implantation.
Clouston and colleagues attempted to analyse all 23 pairs of chromosomes of 438 human blastocysts with the use of classical G-banding [75, 76]. The results obtained showed that in their majority (68 %) these embryos were euploid. The remaining blastocysts were classified as aneuploid, and abnormalities of meiotic and post-zygotic origin were scored. A variety of chromosomes, including 2, 3, 4, 5, 6, 7, one acrocentric chromosome (13, 15), 17 and 22 were seen to participate in aneuploidy events [75, 76].
As with the cleavage stage though, most of the data concerning the chromosome status of blastocysts, have been obtained with the use of FISH. Two different investigations used a combination of immunosurgery to separate TE and ICM from a total of 131 embryos which were classified as aneuploid during day-3 PGS, but went on to form blastocysts [77, 78]. FISH was used to target chromosomes 13, 16, 18, 21, X and Y. The analysis of these embryos showed that both ICM and TE could tolerate a variety of aneuploidies, including tetraploidy, mosaicism and complex abnormalities involving multiple chromosomes, trisomies and monosomies. It was also evident from both studies that there was no preferential allocation of chromosome errors to the TE cells [77, 78].
Many other studies used FISH to examine non-transferred blastocysts, which were either diagnosed as aneuploid via day-3 PGS or had not undergone screening and were donated for research [63, 74, 79–81]. All these studies confirmed that chromosome abnormalities and mosaicism occur at the blastocyst stage, but their incidence is not as frequent as it is during the earlier cleavage stage. The main mosaicism-causing mechanism was identified to be chromosome loss most likely via anaphase lag. This was followed by chromosome gain, whereas MND (leading to monosomic and trisomic cell lines within the same embryo) was not as frequently observed [63, 74].
The use of FISH enabled the accurate identification of ploidy changes, such as triploidy and haploidy, as well. Interestingly, the most prevalent form of ploidy error seen were mosaic diploid-tetraploid blastocysts [63, 74, 82]. The presence of tetraploid cells is considered to represent the start of trophoblast differentiation, and for this reason mosaic diploid-tetraploid blastocysts are not generally considered abnormal [80].
The recent development of more comprehensive molecular cytogenetic methodology such as CGH (metaphase and array) or SNP microarrays enabled an accurate and reliable analysis of the blastocyst chromosomes and provided definitive evidence on the different types and frequency of anomalies that survive to this stage of preimplantation development. During one such study, a combination of metaphase and array CGH and also FISH took place to analyse three different parts obtained from the same blastocyst [17]. Forty-two percent of the 52 blastocysts which were investigated were euploid in every cell. Of the ones characterised as abnormal, 25 % carried the same chromosome error in all of the cells, suggesting a meiotic origin, whereas the remaining embryos had varying degrees of mosacism. Most of these mosaics contained different abnormalities in every cell, but we did observe a few (10 %) diploid-aneuploid mosaic embryos, which contained a majority of normal cells. The fate of such embryos, as far as their ability to implant is concerned is not currently known [17].
Another investigation which used SNP microarrays to analyse 50 blastocysts reported very similar findings to our study [72]. Specifically, mosaicism was also observed in various degrees in 24 % of the embryos, with the remaining 76 % being either completely normal or uniformly abnormal [72]. Combination of the data from both the abovementioned studies, indicate that approximately 30 % of blastocyst embryos are mosaic, but only a small proportion of these contain a diploid cell line as a majority.
The use of more comprehensive methods for the analysis of blastocysts demonstrated that aneuploidy affects chromosomes of all sizes and groups, with 22, 16, 15, 21 and X being the ones to malsegregate most frequently. Additionally, monosomies seemed to be in excess of trisomies (514 vs. 494 respectively), confirming FISH observations about chromosome loss being the main aneuploidy-causing mechanism during this stage of development [83]. It was also clear that even though most (60 %) aneuploid blastocysts tend to carry a single chromosome error in them, there are others (15 %) with multiple errors, which are capable of reaching the final stage of preimplantation development. Similarly to findings obtained for oocytes and cleavage stage embryos, CGH of blastocysts detected several partial abnormalities, mostly affecting pieces of the larger autosomes (i.e. 1–11 and X), but also the smaller 16 and 20 [83].
Analysis of TE and ICM’s coming from the same embryos showed an identical chromosome complement for both parts [84, 85]. These findings agreed with previous FISH investigations, and suggest that TE biopsy, undertaken in a clinical PGS setting, can generally be relied upon to provide a good indication of the chromosomal status of the ICM and therefore of the foetus [77, 78]. As a result, screening of embryos on day-5 to identify ones that are chromosomally normal has been increasingly used by many IVF clinics, and has shown some very encouraging clinical outcomes [86, 87]. Data obtained from such analysis suggest that the expected blastocyst aneuploidy rate for an average female age of 38 years is 56 % (Fragouli et al. and unpublished) [31]. In agreement with observations during female meiosis and cleavage stage (to an extent), advancing female age seems to be closely related to increasing aneuploidy rates at the blastocyst stage as well. Data obtained for 191 women who underwent CGH screening of blastocysts revealed that the aneuploidy rate in younger patients (age range: 29–34 years, average age: 32.3 years) was 46.5 %, whereas in an older group (age range: 35–50, average age: 40) it was 60.6 %, and this difference was found to be highly statistically significant (Fisher’s exact test, P = 0.0007) [83]. Table 4 illustrates results obtained during the cytogenetic analysis of embryos at the final stage of preimplantation development, using different methods.
Conclusions
The exceptionally high incidence of chromosomal anomalies in human preimplantation embryos is the outcome of the accumulated risk of errors that may occur at various stages during gametogenesis and embryogenesis. In rare cases an error may exist or arise in the premeiotic germ cells; much more commonly it may arise during the first or second meiotic division in the male or female. More efficient cell cycle checkpoints in the male ensure that the incidence of aneuploidy in the male gamete is low compared to that in the female. Hence most errors of meiotic origin come from the female. Chromosomal mosaicism affects the majority of 3-day-old embryos created by IVF and this persists to a lesser degree to the blastocyst stage on day 5. With the exception of women over the age of 40 and some that are particularly prone to meiotic errors, embryos from all other groups will have predominantly post-zygotic mitotic errors. Couples experiencing repetitive implantation failure are particularly likely to produce highly abnormal (chaotic) embryos by post-zygotic mechanisms.
References
Edwards RG, Brody SA. Principles and practice of assisted human reproduction. Philadelphia: Saunders; 1995. p. 65–80.
Bonde JP, Ernst E, Jensen TK, et al. Relation between semen quality and fertility: a population based study of 430 first-pregnancy planners. Lancet. 1998;352:1172–7.
Centers for Disease Control and Prevention. 2006 Assisted reproductive technology success rates: preliminary data national summary and fertility clinic reports. 2008. Available at http://www.cdc.gov/art/art2006/index.htm.
Delhanty JD, Griffin DK, Handyside AH, et al. Detection of aneuploidy and chromosomal mosaicism in human embryos during preimplantation sex determination by fluorescent in situ hybridisation (FISH). Hum Mol Genet. 1993;2:1183–5.
Munne S, Lee A, Rosenwaks Z, et al. Diagnosis of major chromosome aneuploidies in human preimplantation embryos. Hum Reprod. 1993;8:2185–91.
Sandalinas M, Marquez C, Munne S. Spectral karyotyping of fresh, non-inseminated oocytes. Mol Hum Reprod. 2002;8:580–5.
Kuliev A, Cieslak J, Ilkevitch Y, et al. Chromosomal abnormalities in a series of 6,733 human oocytes in preimplantation diagnosis for age-related aneuploidies. Reprod Biomed Online. 2003;6:54–9.
Pellestor F, Andreo B, Arnal F, et al. Maternal aging and chromosomal abnormalities: new data drawn from in vitro unfertilized human oocytes. Hum Genet. 2003;112:195–203.
Gutierrez-Mateo C, Benet J, Wells D, et al. Aneuploidy study of human oocytes first polar body comparative genomic hybridization and metaphase II fluorescence in situ hybridization analysis. Hum Reprod. 2004;19:2859–68.
Fragouli E, Wells D, Thornhill A, et al. Comparative genomic hybridization analysis of human oocytes and polar bodies. Hum Reprod. 2006;21:2319–28.
Hassold T, Hall H, Hunt P. The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet. 2007;16:R203–8.
Fragouli E, Alfarawati S, Goodall NN. The cytogenetics of polar bodies: insights into female meiosis and the diagnosis of aneuploidy. Mol Hum Reprod. 2011;17:286–95.
Harton GL, Tempest HG. Chromosomal disorders and male infertility. Asian J Androl. 2012;14:32–9.
Delhanty JD, Harper JC, Ao A, et al. Multicolour FISH detects frequent chromosomal mosaicism and chaotic division in normal preimplantation embryos from fertile patients. Hum Genet. 1997;99:755–60.
Munne S, Sandalinas M, Escudero T, et al. Chromosome mosaicism in cleavage- stage human embryos: evidence of a maternal age effect. Reprod Biomed Online. 2002;4:223–32.
Katz-Jaffe MG, Trounson AO, Cram DS. Mitotic errors in chromosome 21 of human preimplantation embryos are associated with non-viability. Mol Hum Reprod. 2004;10:143–7.
Fragouli E, Alfarawati S, Daphnis DD, et al. Cytogenetic analysis of human blastocysts with the use of FISH, CGH and aCGH: scientific data and technical evaluation. Hum Reprod. 2011;26:480–90.
Delhanty JD. Mechanisms of aneuploidy induction in human oogenesis and early embryogenesis. Cytogenet Genome Res. 2005;111:237–44.
Angell RR. Predivision in human oocytes at meiosis I: a mechanism for trisomy formation in man. Hum Genet. 1991;86:383–7.
Angell RR, Xian J, Keith J, et al. First meiotic division abnormalities in human oocytes: mechanism of trisomy formation. Cytogenet Cell Genet. 1994;65:194–202.
Pellestor F. Frequency and distribution of aneuploidy in human female gametes. Hum Genet. 1991;86:283–8.
Zenzes MT, Casper RF. Cytogenetics of human oocytes, zygotes and embryos after in vitro fertilisation. Hum Genet. 1992;88:367–75.
Kamiguchi Y, Rosenbusch B, Sterzik K, et al. Chromosomal analysis of unfertilized human oocytes prepared by a gradual fixationair drying method. Hum Genet. 1993;90:533–41.
Lim AS, Ho AT, Tsakok MF. Chromosomes of oocytes failing in-vitro fertilization. Hum Reprod. 1995;10:2570–5.
Mahmood R, Brierley CH, Faed MJ, et al. Mechanisms of maternal aneuploidy: FISH analysis of oocytes and polar bodies in patients undergoing assisted conception. Hum Genet. 2000;106:620–6.
Cupisti S, Conn CM, Fragouli E, et al. Sequential FISH analysis of oocytes and polar bodies reveals aneuploidy mechanisms. Prenat Diagn. 2003;23:663–8.
Wells D, Sherlock JK, Handyside AH, et al. Detailed chromosomal and molecular genetic analysis of single cells by whole genome amplification and comparative genomic hybridisation. Nucleic Acids Res. 1999;27:1214–8.
Wells D, Delhanty JD. Comprehensive chromosomal analysis of human preimplantation embryos using whole genome amplification and single cell comparative genomic hybridisation. Mol Hum Reprod. 2000;11:1055–62.
Wells D, Escudero T, Levy B, et al. First clinical application of comparative genomic hybridisation and polar body testing for preimplantation genetic diagnosis of aneuploidy. Fertil Steril. 2002;78:543–9.
Pellestor F, Anahory T, Hamamah S. The chromosomal analysis of human oocytes. Hum Reprod Update. 2005;11:15–32.
Fragouli E, Katz-Jaffe M, Alfarawati S, et al. Comprehensive chromosome screening of polar bodies and blastocysts from couples experiencing repeated implantation failure. Fertil Steril. 2010;94:875–87.
Kuliev A, Verlinsky Y. Meiotic and mitotic nondisjunction: lessons from preimplantation genetic diagnosis. Hum Reprod Update. 2004;10:401–7.
Geraedts J, Montag M, Magli MC, et al. Polar body array CGH for prediction of the status of the corresponding oocyte. Part I: clinical results. Hum Reprod. 2011;26:3173–80.
Fisher JM, Harvey JF, Morton NE, et al. Trisomy 18: studies of the parent and cell division of origin and the effect of aberrant recombination on nondisjunction. Am J Hum Genet. 1995;56:669–75.
Hassold T, Merrill M, Adkins K, et al. Recombination and maternal age-dependent nondisjunction: molecular studies of trisomy 16. Am J Hum Genet. 1995;57:867–74.
Templado C, Vidal F, Estop A. Aneuploidy in human spermatozoa. Cytogenet Genome Res. 2011;133:91–9.
Uroz L, Rajmil O, Templado C. Meiotic chromosome abnormalities in fertile men: are they increasing? Fertil Steril. 2011;95:141–6.
Burgoyne PS, Mahadevaiah SK, Turner JM. The consequences of asynapsis for mammalian meiosis. Nat Rev Genet. 2009;10:207–16.
Uroz L, Rajmil O, Templado C. Premature separation of sister chromatids in human male meiosis. Hum Reprod. 2008;23:982–7.
Uroz L, Templado C. Meiotic non-disjunction mechanisms in human fertile males. Hum Reprod. 2012;27(5):1518–24.
Fonseka KGL, Griffin DK. Is there a paternal age effect for aneuploidy? Cytogenet Genome Res. 2011;133:280–91.
Miharu N. Chromosome abnormalities in sperm from infertile men with normal somatic karyotypes: oligospermia. Cytogenet Genome Res. 2005;111:347–51.
Benet J, Oliver-Bonet M, Cifuentes P, et al. Segregation of chromosomes in sperm of reciprocal translocation carriers. Cytogenet Genome Res. 2005;111:281–90.
Roux C, Tripogney C, Morel F, et al. Segregation of chromosomes in sperm of Robertsonian translocation carriers. Cytogenet Genome Res. 2005;111:291–6.
Iwarsson E, Malmgren H, Inzunza J, et al. Highly abnormal cleavage divisions in preimplantation embryos from translocation carriers. Prenat Diagn. 2000;20:1038–47.
Simopoulou M, Harper JC, Fragouli E, et al. Preimplantation genetic diagnosis of chromosome abnormalities: implications from the outcome for couples with chromosomal rearrangements. Prenat Diagn. 2003;23:652–62.
Robinson WP, McFadden DE, Stephenson MD. The origin of abnormalities in recurrent aneuploidy/polyploidy. Am J Hum Genet. 2001;69:1245–54.
Kovaleva NV, Tahmasebi-Hesari M. Gonadal mosaicism. Down Syndr News. 2007;14:23.
Somprasit C, Aguinaga M, Cisneros PL, et al. Paternal gonadal mosaicism detected in a couple with recurrent abortions undergoing PGD: FISH analysis of sperm nuclei proves valuable. Reprod Biomed Online. 2004;9:225–30.
Cozzi J, Conn CM, Harper J, et al. A trisomic germ cell line and precocious chromatid separation leads to recurrent trisomy 21 conception. Hum Genet. 1999;104:23–8.
Pujol A, Boiso I, Benet J, et al. Analysis of nine chromosome pairs in first polar bodies and metapahse II oocytes for the detection of aneuploidies. Eur J Hum Genet. 2003;11:325–36.
Hultén MA, Patel SD, Tankimanova M, et al. On the origin of trisomy 21 Down syndrome. Mol Cytogenet. 2008;1:21.
Mantzouratou A, Delhanty JD. Aneuploidy in the human cleavage stage embryo. Cytogenet Genome Res. 2011;133:141–8.
Hultén MA, Patel SD, Westgren M, et al. On the paternal origin of trisomy 21 Down Syndrome. Mol Cytogenet. 2010;3:4.
Morelli M, Cohen P. Not all germ cells are created equal: aspects of sexual dimorphism in mammalian meiosis. Reproduction. 2005;130:761–81.
Speed RM. Meiotic configurations in female trisomy 21 foetuses. Hum Genet. 1984;66:176–80.
LeMaire-Adkins R, Radke K, et al. Lack of checkpoint control at the metaphase/anaphase transition: a mechanism of meiotic nondisjunction in mammalian females. J Cell Biol. 1997;139:1611–9.
Kourznetsova A, Lister L, Nordenskjöld M, et al. Bi-orientation of achiasmatic chromosomes in meiosis I oocytes contributes to aneuploidy in mice. Nat Genet. 2007;39:966–8.
Heikinheimo O, Gibbons WE. The molecular mechanisms of oocyte maturation and early embryonic development are unveiling new insights into reproductive medicine. Mol Hum Reprod. 1998;4:745–56.
Braude P, Bolton V, Moore S. Human gene expression occurs between the four- and eight-cell stages of preimplantation development. Nature. 1988;332:459–61.
Edwards RG, Beard HK. Oocyte polarity and cell determination in early mammalian embryos. Mol Hum Reprod. 1997;3:863–905.
Jamieson ME, Coutts JRT, Conner JM. The chromosome constitution of human preimplantation embryos fertilised in vitro. Hum Reprod. 1994;9:709–15.
Coonen E, Derhaag JG, Dumoulin JC, et al. Anaphase lagging mainly explains chromosomal mosaicism in human preimplantation embryos. Hum Reprod. 2004;19:316–24.
Munné S, Grifo J, Cohen J. Chromosome abnormalities in human arrested preimplantation embryos fertilized in vitro: a multiprobe FISH study. Am J Hum Genet. 1994;55:150–9.
Harper JC, Coonen E, Handyside AH, et al. Mosaicism of autosomes and sex chromosomes in morphologically normal, monospermic preimplantation human embryos. Prenat Diagn. 1995;15:41–9.
Munné S, Sultan KM, Weier HU, Grifo JA, Cohen J, Rosenwaks Z. Assessment of numeric abnormalities of X, Y, 18, and 16 chromosomes in preimplantation human embryos before transfer. Am J Obstet Gynecol. 1995;172:1191–9.
Mantzouratou A, Mania A, Fragouli E, et al. Variable aneuploidy mechanisms in embryos from couples with poor reproductive histories undergoing preimplantation genetic screening. Hum Reprod. 2007;22:1844–53.
Voullaire L, Collins V, Callaghan T, et al. High incidence of complex chromosome abnormality in cleavage embryos from patients with repeated implantation failure. Fertil Steril. 2007;87:1053–8.
Voullaire L, Slater H, Williamson R, et al. Chromosome analysis of blastomeres from human embryos by using comparative genomic hybridisation. Hum Genet. 2000;106:210–7.
Voullaire L, Wilton L, McBain J, et al. Chromosome abnormalities identified by comparative genomic hybridization in embryos from women with repeated implantation failure. Mol Hum Reprod. 2002;8:1035–41.
Bielanska M, Tan SL, Ao A. Chromosomal mosaicism throughout human preimplantation development in vitro: incidence, type, and relevance to embryo outcome. Hum Reprod. 2002;17:413–9.
Northrop LE, Treff NR, Levy B, et al. SNP microarray based 24 chromosome aneuploidy screening demonstrates that cleavage stage FISH poorly predicts aneuploidy in embryos that develop to morphologically normal blastocysts. Mol Hum Reprod. 2010;16:590–600.
Munné S, Bahçe M, Sandalinas M, et al. Differences in chromosome susceptibility to aneuploidy and survival to first trimester. Reprod Biomed Online. 2004;8:81–90.
Daphnis DD, Delhanty JD, Jerkovic S, et al. Detailed FISH analysis of day 5 human embryos reveals the mechanisms leading to mosaic aneuploidy. Hum Reprod. 2005;20:129–37.
Clouston HJ, Fenwick J, Webb AL, et al. Detection of mosaic and non-mosaic chromosome abnormalities in 6- to 8-day old human blastocysts. Hum Genet. 1997;101:30–6.
Clouston HJ, Herbert M, Fenwick J, et al. Cytogenetic analysis of human blastocysts. Prenat Diagn. 2002;22:1143–52.
Evsikov S, Verlinsky Y. Mosaicism in the inner cell mass of human blastocysts. Hum Reprod. 1998;13:3151–5.
Magli MC, Jones GM, Gras L, et al. Chromosome mosaicism in day 3 aneuploid embryos that develop to morphologically normal blastocysts in vitro. Hum Reprod. 2000;15:1781–6.
Ruangvutilert P, Delhanty JD, Serhal P, et al. FISH analysis on day 5 post-insemination of human arrested and blastocyst stage embryos. Prenat Diagn. 2000;20:552–60.
Sandalinas M, Sadowy S, Alikani M, et al. Developmental ability of chromosomally abnormal human embryos to develop to the blastocyst stage. Hum Reprod. 2001;16:1954–8.
Santos MA, Teklenburg G, Macklon NS, et al. The fate of the mosaic embryo: chromosomal constitution and development of Day 4, 5 and 8 human embryos. Hum Reprod. 2010;25:1916–26.
Bielanska M, Jin S, Bernier M, et al. Diploid- aneuploid mosaicism in human embryos cultured to the blastocyst stage. Fertil Steril. 2005;84:336–42.
Fragouli E, Wells D. Aneuploidy in the human blastocyst. Cytogenet Genome Res. 2011;133:149–59.
Fragouli E, Lenzi M, Ross R, et al. Comprehensive molecular cytogenetic analysis of the human blastocyst stage. Hum Reprod. 2008;23:2596–608.
Johnson DS, Cinnioglu C, Ross R, et al. Comprehensive analysis of karyotypic mosaicism between trophectoderm and inner cell mass. Mol Hum Reprod. 2010;16:944–9.
Schoolcraft WB, Fragouli E, Stevens J, et al. Clinical application of comprehensive chromosomal screening at the blastocyst stage. Fertil Steril. 2010;94:1700–6.
Schoolcraft WB, Treff NR, Stevens JM, et al. Live birth outcome with trophectoderm biopsy, blastocyst vitrification, and single-nucleotide polymorphism microarray-based comprehensive chromosome screening in infertile patients. Fertil Steril. 2011;96:638–40.
Ruangvutilert P, Delhanty JD, Serhal P, Simopoulou M, Rodeck CH, Harper JC. FISH analysis on day 5 post-insemination of human arrested and blastocyst stage embryos. Prenat Diagn. 2000;20:552–560.
Additional Reading
Jones KT. Meiosis in oocytes: predisposition to aneuploidy and its increased incidence with age. Hum Reprod Update. 2008;14:143–58.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Fragouli, E., Delhanty, J. (2013). The Origins of Aneuploidy in Human Embryos. In: Gardner, D., Sakkas, D., Seli, E., Wells, D. (eds) Human Gametes and Preimplantation Embryos. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6651-2_10
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6651-2_10
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6650-5
Online ISBN: 978-1-4614-6651-2
eBook Packages: MedicineMedicine (R0)