Abstract
In normal cells, autophagy prevents tumorigenesis through selective cleanup of damaged organelles and certain specific proteins such as p62. In contrast, autophagy provides tumor cells, which require enormous amounts of nutrients, with amino acids, fatty acids, and glucose. Therefore, autophagy represents something of a double-edged sword in cancer: it functions as a tumor suppressor, but can also satisfy metabolic demands once tumors are established. In this chapter, we review the tumor-suppressive and oncogenic effects of autophagy which have been characterized using several approaches including transgenic mice and introduce the involvement of selective autophagy.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Autophagy Autophagy
Macroautophagy (hereafter “autophagy”) is the best characterized form of autophagy, in which isolation membranes, or phagophores, engulf a portion of the cytoplasm and form double-membrane vesicles called autophagosomes. The resulting autophagosomes fuse with lysosomes to form autolysosomes, wherein the contents are degraded. Since the discovery of autophagy-related (ATG) genes in the yeast Saccharomyces cerevisiae, the autophagy field has expanded and it has become apparent that autophagy is involved in multiple physiological events in various species (Xie and Klionsky 2007; Nakatogawa et al. 2009; Mizushima et al. 2011). There is general agreement that a major function of autophagy is to supply amino acids for protein synthesis during starvation (Mizushima and Levine 2010; Mizushima and Komatsu 2011). Several lines of evidence also point to the importance of basal autophagy, which occurs at a low, constitutive rate even in nutrient-rich conditions, in quality control of cellular components including organelles (Mizushima and Komatsu 2011; Komatsu and Ichimura 2010).
1.1 Selective Autophagy
The autophagosome is a globular organelle with a diameter of approximately 1 μm and with volume of 0.5 × 10−18 m3. Given these large dimensions relative to proteins of average molecular weight, one autophagosome can entrap hundreds of thousands of proteins. Therefore, autophagy has been generally considered a bulk protein degradation process that does not select individual cargo proteins. However, recent studies provide evidence that autophagy can specifically eliminate aggregated proteins, unnecessary or damaged organelles, and invading bacteria (Mizushima and Komatsu 2011). This selective type of autophagy is mediated by cargo receptor proteins (e.g., Atg19, Atg32, Atg30, SEPA-1, p62, Nbr1, Nix, NDP52) that interact with autophagosomal membrane proteins, such as Atg8/LC3, and/or adaptor proteins, for instance Atg11 (Fig. 6.1 and Table 6.1), to define selectivity in various physiological situations (Komatsu and Ichimura 2010; Noda et al. 2010; Johansen and Lamark 2011). Huge aggregates containing ubiquitinated proteins are unable to be degraded by the relatively small proteasome; rather these ubiquitinated aggregates are sequestered into autophagosomes by autophagy receptors, such as p62 and NBR1, and degraded by selective autophagy (Fig. 6.1a) (Komatsu et al. 2007; Kirkin et al. 2009). Impairment of selective autophagy leads to the disrupted tissue homeostasis and results in life-threatening diseases, indicating that this system is physiologically important. Defective autophagy is usually accompanied by accumulation of p62-containing aggregates, which enhances the function of p62 as a scaffold protein in several signaling cascades such as NF-κB signaling, apoptosis, and Nrf2 activation (Fig. 6.1b) (Duran et al. 2008; Komatsu et al. 2010). Such abnormalities might be involved in tumorigenesis (see Sect. 3.2) and Paget’s disease of bone.
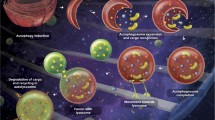
Selective autophagy. (a) Scheme of selective autophagy. p62–Nbr1 aggregates containing ubiquitinated proteins are selectively trapped in autophagosomes by interaction with LC3. (b) Accumulation of p62 following defective autophagy induces cell proliferation or apoptosis because p62 acts as a scaffold protein in several signaling pathways, such as NF- κB signaling, apoptosis, and Nrf2 activation. (c) Nix/Bnip3L is a mitochondrial outer membrane protein that interacts with GABARAP and triggers mitophagy during erythroid differentiation. Parkin translocates to damaged mitochondria in a PINK1-dependent manner, leading to ubiquitination of mitochondrial outer membrane proteins, which marks mitochondria for autophagic sequestration. p62 may be an adaptor in mitophagy. (d) Phosphorylated Atg30, a receptor protein on peroxisomes, interacts with Atg11 and Atg17, causing a peroxisome to be surrounded by an autophagosomal membrane. (e) Cytosolic bacteria are ubiquitinated and removed by p62-, NDP52-, or OPTN-mediated selective autophagy. (f) Autophagy selectively degrades lipid droplets to supply fatty acids and inhibits the intracellular accumulation of lipids
Autophagy is the only known intracellular system capable of degrading the whole organelles. The autophagic degradation of mitochondria is referred to as mitophagy. During erythroid differentiation, mitochondria are selectively eliminated by mitophagy. In this process, Nix/Bnip3L localized to mitochondrial membranes induces mitochondrial depolarization, which promotes mitophagy (Sandoval et al. 2008; Schweers et al. 2007). Loss of Nix/Bnip3L causes retention of mitochondria and results in erythroid maturation arrest, which is associated with severe anemia (Sandoval et al. 2008; Schweers et al. 2007). When mitochondrial membrane potential is lost due to damage, an uncoupling agent, Parkin, translocates from the cytosol to the depolarized mitochondria in a PINK1-dependent manner. Consequently, Parkin ubiquitinates mitochondrial outer membrane proteins and targets the mitochondria for mitophagy (Fig. 6.1c) (Narendra et al. 2008; Matsuda et al. 2010; Narendra et al. 2010; Vives-Bauza et al. 2010). A subset of Parkinson’s disease-related mutations in the Parkin and PINK1 genes result in defective mitophagy, suggesting that mitophagy has a role in preventing the pathogenesis of the Parkinson’s disease.
Autophagy also acts in the selective clearance of peroxisomes called pexophagy. Peroxisomes proliferate in mouse liver in response to phthalate esters, and excess peroxisomes are removed by pexophagy in the recovery process following withdrawal of such agents (Fig. 6.1d) (Iwata et al. 2006).
Autophagy also eliminates bacteria which have invaded mammalian cells. Bacteria in the cytosol are ubiquitinated and recognized by autophagy receptors such as p62, NDP52, and optineurin (OPTN) (Thurston et al. 2009; Wild et al. 2011). This system allows specific sequestration of microbes into autophagosomes, thereby inhibiting their replication and enabling their destruction (Fig. 6.1e).
Lipid droplets are also likely selectively degraded by autophagy. This type of autophagy, called lipophagy, supplies free fatty acids, which can be used for energy generation through β-oxidation (Fig. 6.1f) (Singh et al. 2009). Impairment of lipophagy results in accumulation of lipid droplets in hepatocytes and reduced production of Agouti-related peptide (AgRP) in neurons (Kaushik et al. 2011).
2 Tumor-Suppressive Role of Autophagy
2.1 mTORC1
Mutation or deletion of oncogenes and tumor-suppressor genes involved in the insulin-signaling pathway suppresses autophagy due to persistent activation of mammalian target of rapamycin complex 1 (mTORC1), which serves as a checkpoint kinase in autophagy. Reduced autophagy as a result of constantly activated mTORC1 may participate in neoplasia. In fact, liver-specific Pten or Tsc1 knockout mice exhibit decreased autophagy accompanying constitutively activated mTORC1 and develop hepatocellular carcinoma (Horie et al. 2004; Menon et al. 2012). However, the specific contribution of impaired autophagy to neoplasia is unclear because mTORC1 controls multiple physiologic processes, including protein translation and gene transcription in addition to autophagy.
2.2 Beclin 1 and Its Binding Partners
In 1999, Levine’s group reported that Beclin1 (homologue of yeast ATG6/VPS30) is a tumor-suppressor gene, implicating autophagy in tumor suppression (Liang et al. 1999). Autophagy requires phosphatidylinositol 3-phosphate (PI3P) generated by class III phosphatidylinositol-3 kinase (PI3K). The class III PI3K complex is composed of p150, Vps34, Beclin1, and Atg14 (complex I), and is involved in early autophagosome formation (Itakura et al. 2008; Matsunaga et al. 2010). Another PI3K complex comprising p150, Vps34, Beclin1, and UV irradiation resistance-associated gene (UVRAG) (complex II) facilitates autophagosome and endosome maturation (Liang et al. 2006, 2008; Matsunaga et al. 2009). In contrast, complex III, which consists of RUN domain- and cysteine-rich domain-containing, Beclin 1-interacting protein (Rubicon) associated with complex II (the UVRAG-containing PI3K complex), negatively regulates a later step of autophagy and the endocytic pathway (Matsunaga et al. 2009; Zhong et al. 2009). Bcl-2, an anti-apoptotic protein, interacts with Beclin1 and inhibits starvation-induced autophagy, whereas mutants of Beclin1 that cannot bind to Bcl-2 promote autophagy. It has been proved that Beclin1 is bound to Bcl-2 on the endoplasmic reticulum (ER), while it is released from Bcl-2 through phosphorylation by c-Jun N-terminal kinase 1 under starvation conditions (Wei et al. 2008). Beclin 1 +/− mice show significantly reduced autophagic activity and increased risk for cancer: These mice develop spontaneous tumors, including hepatocellular carcinoma, and more frequently develop liver tumors after hepatitis B virus infection (Qu et al. 2003; Yue et al. 2003). The Beclin1-interacting protein UVRAG enhances autophagy and suppresses the proliferation and tumorigenicity of human colon cancer (Liang et al. 2006). Furthermore, loss of BAX-interacting factor 1 (Bif-1), which activates PI3K complex II through interaction with UVRAG, causes lymphoma, hepatocarcinoma, and colorectal adenocarcinoma in mice (Takahashi et al. 2007). These reports suggest inactivation of class III PI3K in neoplasia. However, it is unclear whether the phenotypes of Beclin1, UVRAG, and Bif-1 knockout mice are entirely the result of defective autophagy since the class III PI3K is involved not only in autophagy but also in the endocytic pathway.
2.3 Atg Proteins
In 2011, it became clear that deletion of Atg5 or Atg7 also causes multiple types of hepatic tumors (Takamura et al. 2011; Inami et al. 2011). Both Atg5 and Atg7 are involved in two ubiquitin-like conjugation systems essential for autophagosome formation. The ubiquitin-like modifiers Atg12 and LC3 (homologue of yeast Atg8) are activated by the E1-like enzyme Atg7 and transferred to two different E2-like enzymes, Atg10 and Atg3, respectively (Mizushima et al. 1998; Ichimura et al. 2000). Whereas Atg12 forms an isopeptide bond with Atg5, LC3 forms an amide bond with phosphatidylethanolamine (PE) dependent on Atg12–Atg5 conjugation (Mizushima et al. 1998; Ichimura et al. 2000). Atg16L forms a high-molecular-weight complex with Atg12–Atg5 (Mizushima et al. 2003). The Atg12–Atg5–Atg16L complex functions as an E3-like enzyme, determining the site of LC3 lipidation (Fujita et al. 2008a). While the Atg12–Atg5–Atg16L complex is required for elongation of the isolation membrane (Mizushima et al. 2001), PE-bound LC3 (LC3-II) is thought to be important for membrane biogenesis and/or closure of the autophagosome (Kabeya et al. 2000; Fujita et al. 2008b; Sou et al. 2008). The Atg5 and Atg7 proteins have a specialized function for autophagy, although recent studies have revealed novel roles of Atg12–Atg5–Atg16L and Atg7 distinct from autophagosome formation in specialized cells such as macrophages and neuroendocrine cells (Zhao et al. 2008; Lee et al. 2012). Microtumors form in the livers of mosaic Atg5-deficient mice and hepatocyte-specific Atg7 knockout mice when the mice are 7–9 months old (Takamura et al. 2011). Importantly, Atg5 mosaic knockouts develop tumors only in liver tissues (Takamura et al. 2011). The number and size of the tumors in mouse livers increased with aging, with livers being almost covered by tumors when the mice are 16–19 months old (Takamura et al. 2011). Tumors in mutant livers are monoclonal with regular arrangements and patterns and metastasis is not observed in other tissues, suggesting that these tumors are benign adenomas (Takamura et al. 2011).
3 Mechanism of Tumorigenesis in Autophagy-Deficient Tissues
3.1 Organelle Homeostasis
Basal autophagy constitutively catabolizes cytoplasmic components and prevents the accumulation of aggregating proteins and organelles. Such autophagic surveillance of cytoplasmic material in hepatocytes might be more important under stressful or pathologic conditions. Excess peroxisomes are thought to initiate neoplastic transformation of hepatocytes by DNA damage associated with increasing levels of intracellular reactive oxygen species (ROS), suggesting that pexophagy has an antitumorigenic effect (Iwata et al. 2006; Warren et al. 1982). Mitochondria generate ROS concomitantly with energy production through oxidative phosphorylation, causing protein, lipid, and DNA oxidation, and excess ROS can induce cell death. Therefore, quality control of mitochondria by mitophagy is also essential for cellular homeostasis and tumor suppression (Youle and Narendra 2011). Atg5- or Atg7-deficient hepatocytes show enlarged mitochondria and excess peroxisomes (Komatsu et al. 2005, 2007; Hara et al. 2006; Komatsu et al. 2006), likely due to impaired mitophagy and pexophagy. As a result, cells in autophagy-deficient livers are thought to accumulate ROS, resulting in spontaneous tumorigenesis followed by genomic instability (Fig. 6.2) (Takamura et al. 2011; Inami et al. 2011). DNA damage in autophagy-deficient cells also activates the DNA damage-sensing, ataxia telangiectasia mutated (ATM)-dependent kinase Chk2, which phosphorylates Ser20 of tumor-suppressor protein p53, which then induces expression of a series of proapoptotic genes (e.g., BAX, PUMA, NOXA) and cell death (Lee et al. 2012). Indeed, the phosphorylation of p53 and enhanced cell death observed in Atg7 −/− MEFs are restored by simultaneous depletion of Chk2 (Lee et al. 2012). Considering the ability of hepatic cells to regenerate, loss of autophagy in liver could give rise to an imbalance between cell proliferation and death (Fig. 6.2).
Mechanism of tumorigenesis in autophagy deficiency. Autophagy defects result in accumulation of damaged mitochondria and peroxisomes, increasing ROS. ROS cause oxidative damage to DNA, proteins, and lipids, thereby inducing tumorigenesis. Further, accumulation of p62 in autophagy-deficient cells leads to Nrf2 activation, apoptosis, and dysregulation of NF-κB signaling. ROS also activate the phosphorylation of p53 mediated by the ATM–Chk2 pathway, leading to apoptosis. In addition, Atg7 deficiency results in impairment of p53-mediated cell cycle arrest
3.2 Autophagic Turnover of p62
Liver adenoma growth in mice with liver-specific knockout of Atg7 is remarkably suppressed when p62 is simultaneously knocked out, because loss of autophagy results in marked accumulation of p62, which leads to dysregulation of NF-κB signaling, apoptosis, and Nrf2-activation (Fig. 6.2) (Duran et al. 2008; Jin et al. 2009; Komatsu et al. 2010). NF-κB is a transcription factor regulating cell survival and proliferation, and it is frequently activated in tumor cells. Increased p62 in autophagy-incompetent cells leads to dysregulation of NF-κB signaling, which may predispose to tumors in mutant mice (Mathew et al. 2009). In agreement with this hypothesis, suppressing NF-κB signaling by p62 knockout prevents growth and development of Ras-induced lung adenocarcinoma (Duran et al. 2008). It is also known that induction of p62 expression via constitutive activation of Kras contributes to the development of pancreatic ductal adenocarcinoma (Ling et al. 2012). Furthermore, it has been reported that Nrf2 activation participates in tumorigenesis and/or tumor development. The transcription factor Nrf2 is responsible for the expression of a battery of genes encoding antioxidant proteins and detoxification enzymes. Nrf2 is usually interacted and ubiquitinated by the Keap1–Cul3 ligase complex, and degraded by the proteasome. Somatic mutations in either Keap1 or Nrf2 have been identified in patients with lung, head, neck, and gallbladder cancers (Taguchi et al. 2011; Hayes and McMahon 2009). These mutations result in the loss of interaction between Keap1 and Nrf2 and are accompanied by persistent activation of Nrf2, which makes tumor cells resistant to oxidative damage and anticancer agents (Taguchi et al. 2011; Hayes and McMahon 2009). Nrf2 is activated in certain types of cancer even in the absence of these somatic mutations. For instance, in type 2 papillary renal cell carcinomas that carry mutations in fumarate hydratase, Keap1 is succinated, leading to hyperactivation of Nrf2 (Adam et al. 2011; Kinch et al. 2011). In addition, oncogene-driven Nrf2 transcription serves as an early tumorigenic event (DeNicola et al. 2011). Furthermore, Nrf2 redirects glucose and glutamine into anabolic pathways, specially under the sustained activation of PI3K–Akt signaling (Mitsuishi et al. 2012). The active PI3K–Akt pathway augments the nuclear accumulation of Nrf2 and enables Nrf2 to promote metabolic activities that support cell proliferation in addition to enhancing cytoprotection (Mitsuishi et al. 2012). These studies suggest that activation of Nrf2 is involved in tumorigenesis and tumor development. Interestingly, p62-positive aggregates are often detected in human cancers, including as hepatocellular carcinoma (Zatloukal et al. 2002), and expression of Nrf2 target genes has been observed in most of these tumors, suggesting that persistent activation of Nrf2 in response to increased levels of p62 contributes to hepatoma development.
4 Advanced Cancer and Autophagy
Although autophagy functions as a tumor suppressor in non-tumor cells and during the early stages of tumor cell development, autophagy becomes important for cancer cell survival once tumors are established. Cancer cells have an increased metabolic demand (in terms of both energy sources and building blocks), and they often need to grow under hypoxic conditions until angiogenesis can establish blood flow to the tumor (Kimmelman 2011). Therefore, cancer cells, particularly those with Ras mutations such as pancreatic cancer, rely heavily on autophagy and are “addicted” to autophagy (Guo et al. 2011). Though the precise molecular mechanism remains unclear, blockade of autophagy is sufficient to inhibit proliferation of pancreatic cancer cells (Guo et al. 2011). In support of this, treatment with a combination of a leucine-free diet, which activates caspases and triggers apoptosis, and an autophagy inhibitor synergistically suppresses tumor growth in xenograft models of human melanoma tumors (Sheen et al. 2011). Autophagy suppression has also been shown to reduce growth in Myc-induced lymphoma and polyomavirus middle T-induced mammary tumor cells (Amaravadi et al. 2007). Accumulation of ROS and genomic instability are observed in pancreatic cancer cells with defective autophagy. Further, loss of autophagy in pancreatic cancer cells is accompanied by impaired oxidative phosphorylation, likely due to decreased supply of intermediates from the tricarboxylic acid cycle (Karantza-Wadsworth et al. 2007). Hence, it is plausible that cancer cells are addicted to autophagy as it is critical to both quality control of organelles such as mitochondria and peroxisomes and supply of amino acids to support their survival and proliferation under metabolic stress conditions (Fig. 6.3). However, the involvement of Ras is not simple: Ras-induced autophagy contributes to tumor suppression by inducing autophagic cell death and senescence (Elgendy et al. 2011). Ras-mediated autophagy might have different roles in tumor growth dependent on cellular context or cancer stage.
5 Concluding Remarks
Because inhibition of autophagy has been shown to have a suppressive effect on cancer proliferation in many types of cancer cells, clinical trials with autophagy repressors combined with antineoplaston therapy or radiotherapy have already started. However, several issues remain before inhibition of autophagy is ready for clinical application. First, because the molecular mechanism and the physiological role of autophagy in cancer are still incompletely understood, unexpected side effects might occur. Moreover, neither biomarkers in blood or urine judging the effect of the drug nor probes that can visualize autophagic activity in vivo have been established to date. Therefore, we cannot determine whether it is truly suppression of autophagy which is responsible for response to treatment even if certain drugs are effective. Further, the development of autophagy-specific inhibitors is necessary because the most common drugs used to inhibit autophagy are chloroquine and its derivative hydroxychloroquine, which both function by obstructing lysosomal acidification. Finally, it is crucial to develop a drug which suppresses autophagy in autophagy-addicted tumor cells but not in normal cells, because autophagy is indispensable for homeostasis in almost all normal tissues.
References
Adam J, Hatipoglu E, O’Flaherty L, Ternette N, Sahgal N, Lockstone H et al (2011) Renal cyst formation in Fh1-deficient mice is independent of the Hif/Phd pathway: roles for fumarate in KEAP1 succination and Nrf2 signaling. Cancer Cell 20:524–537
Amaravadi RK, Yu D, Lum JJ, Bui T, Christophorou MA, Evan GI et al (2007) Autophagy inhibition enhances therapy-induced apoptosis in a Myc-induced model of lymphoma. J Clin Invest 117:326–336
DeNicola GM, Karreth FA, Humpton TJ, Gopinathan A, Wei C, Frese K et al (2011) Oncogene-induced Nrf2 transcription promotes ROS detoxification and tumorigenesis. Nature 475:106–109
Duran A, Linares JF, Galvez AS, Wikenheiser K, Flores JM, Diaz-Meco MT et al (2008) The signaling adaptor p62 is an important NF-kappaB mediator in tumorigenesis. Cancer Cell 13:343–354
Elgendy M, Sheridan C, Brumatti G (2011) Oncogenic Ras-induced expression of Noxa and Beclin-1 promotes autophagic cell death and limits clonogenic survival. Mol Cell 42:23–35
Fujita N, Itoh T, Omori H, Fukuda M, Noda T, Yoshimori T (2008a) The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell 19:2092–2100
Fujita N, Hayashi-Nishino M, Fukumoto H, Omori H, Yamamoto A, Noda T et al (2008b) An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol Biol Cell 19:4651–4659
Guo JY, Chen HY, Mathew R, Fan J, Strohecker AM, Karsli-Uzunbas G et al (2011) Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev 25:460–470
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441:885–889
Hayes JD, McMahon M (2009) NRF2 and KEAP1 mutations: permanent activation of an adaptive response in cancer. Trends Biochem Sci 34:176–188
Horie Y, Suzuki A, Kataoka E, Sasaki T, Hamada K, Sasaki J et al (2004) Hepatocyte-specific Pten deficiency results in steatohepatitis and hepatocellular carcinomas. J Clin Invest 113:1774–1783
Ichimura Y, Kirisako T, Takao T, Satomi Y, Shimonishi Y, Ishihara N et al (2000) A ubiquitin-like system mediates protein lipidation. Nature 408:488–492
Inami Y, Waguri S, Sakamoto A, Kouno T, Nakada K, Hino O et al (2011) Persistent activation of Nrf2 through p62 in hepatocellular carcinoma cells. J Cell Biol 193:275–284
Itakura E, Kishi C, Inoue K, Mizushima N (2008) Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol Biol Cell 19:5360–5372
Iwata J, Ezaki J, Komatsu M, Yokota S, Ueno T, Tanida I et al (2006) Excess peroxisomes are degraded by autophagic machinery in mammals. J Biol Chem 281:4035–4041
Jin Z, Li Y, Pitti R, Lawrence D, Pham VC, Lill JR et al (2009) Cullin3-based polyubiquitination and p62-dependent aggregation of caspase-8 mediate extrinsic apoptosis signaling. Cell 137:721–735
Johansen T, Lamark T (2011) Selective autophagy mediated by autophagic adapter proteins. Autophagy 7:279–296
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al (2000) LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 19:5720–5728
Karantza-Wadsworth V, Patel S, Kravchuk O, Chen G, Mathew R, Jin S et al (2007) Autophagy mitigates metabolic stress and genome damage in mammary tumorigenesis. Genes Dev 21:1621–1635
Kaushik S, Rodriguez-Navarro JA, Arias E, Kiffin R, Sahu S, Schwartz GJ et al (2011) Autophagy in hypothalamic AgRP neurons regulates food intake and energy balance. Cell Metab 14:173–183
Kimmelman AC (2011) The dynamic nature of autophagy in cancer. Genes Dev 25:1999–2010
Kinch L, Grishin NV, Brugarolas J (2011) Succination of Keap1 and activation of Nrf2-dependent antioxidant pathways in FH-deficient papillary renal cell carcinoma type 2. Cancer Cell 20:418–420
Kirkin V, Lamark T, Sou YS, Bjorkoy G, Nunn JL, Bruun JA et al (2009) A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell 33:505–516
Komatsu M, Ichimura Y (2010) Selective autophagy regulates various cellular functions. Genes Cells 15:923–933
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I et al (2005) Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol 169:425–434
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441:880–884
Komatsu M, Waguri S, Koike M, Sou YS, Ueno T, Hara T et al (2007) Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131:1149–1163
Komatsu M, Kurokawa H, Waguri S, Taguchi K, Kobayashi A, Ichimura Y et al (2010) The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nat Cell Biol 12:213–223
Lee IH, Kawai Y, Fergusson MM, Rovira II, Bishop AJ, Motoyama N et al (2012) Atg7 modulates p53 activity to regulate cell cycle and survival during metabolic stress. Science 336:225–228
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al (1999) Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 402:672–676
Liang C, Feng P, Ku B, Dotan I, Canaani D, Oh BH et al (2006) Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol 8:688–699
Liang C, Lee JS, Inn KS, Gack MU, Li Q, Roberts EA et al (2008) Beclin1-binding UVRAG targets the class C Vps complex to coordinate autophagosome maturation and endocytic trafficking. Nat Cell Biol 10:776–787
Ling J, Kang Y, Zhao R, Xia Q, Lee DF, Chang Z et al (2012) KrasG12D-induced IKK2/beta/NF-kappaB activation by IL-1alpha and p62 feedforward loops is required for development of pancreatic ductal adenocarcinoma. Cancer Cell 21:105–120
Mathew R, Karp CM, Beaudoin B, Vuong N, Chen G, Chen HY et al (2009) Autophagy suppresses tumorigenesis through elimination of p62. Cell 137:1062–1075
Matsuda N, Sato S, Shiba K, Okatsu K, Saisho K, Gautier CA et al (2010) PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J Cell Biol 189:211–221
Matsunaga K, Saitoh T, Tabata K, Omori H, Satoh T, Kurotori N et al (2009) Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat Cell Biol 11:385–396
Matsunaga K, Morita E, Saitoh T, Akira S, Ktistakis NT, Izumi T et al (2010) Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L. J Cell Biol 190:511–521
Menon S, Yecies JL, Zhang HH, Howell JJ, Nicholatos J, Harputlugil E et al (2012) Chronic activation of mTOR complex 1 is sufficient to cause hepatocellular carcinoma in mice. Sci Signal 5(27):ra24
Mitsuishi Y, Taguchi K, Kawatani Y, Shibata T, Nukiwa T, Aburatani H et al (2012) Nrf2 redirects glucose and glutamine into anabolic pathways in metabolic reprogramming. Cancer Cell 22:66–79
Mizushima N, Komatsu M (2011) Autophagy: renovation of cells and tissues. Cell 147:728–741
Mizushima N, Levine B (2010) Autophagy in mammalian development and differentiation. Nat Cell Biol 12:823–830
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD et al (1998) A protein conjugation system essential for autophagy. Nature 395:395–398
Mizushima N, Yamamoto A, Hatano M, Kobayashi Y, Kabeya Y, Suzuki K et al (2001) Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J Cell Biol 152:657–668
Mizushima N, Kuma A, Kobayashi Y, Yamamoto A, Matsubae M, Takao T et al (2003) Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12–Apg5 conjugate. J Cell Sci 116:1679–1688
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132
Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y (2009) Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol 10:458–467
Narendra D, Tanaka A, Suen DF, Youle RJ (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183(5):795–803
Narendra DP, Jin SM, Tanaka A, Suen DF, Gautier CA, Shen J et al (2010) PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol 8:e1000298
Noda NN, Ohsumi Y, Inagaki F (2010) Atg8-family interacting motif crucial for selective autophagy. FEBS Lett 584:1379–1385
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A et al (2003) Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 112:1809–1820
Sandoval H, Thiagarajan P, Dasgupta SK, Schumacher A, Prchal JT, Chen M et al (2008) Essential role for Nix in autophagic maturation of erythroid cells. Nature 454:232–235
Schweers RL, Zhang J, Randall MS, Loyd MR, Li W, Dorsey FC et al (2007) NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc Natl Acad Sci USA 104:19500–19505
Sheen JH, Zoncu R, Kim D, Sabatini DM (2011) Defective regulation of autophagy upon leucine deprivation reveals a targetable liability of human melanoma cells in vitro and in vivo. Cancer Cell 19:613–628
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M et al (2009) Autophagy regulates lipid metabolism. Nature 458:1131–1135
Sou YS, Waguri S, Iwata J, Ueno T, Fujimura T, Hara T et al (2008) The Atg8 conjugation system is indispensable for proper development of autophagic isolation membranes in mice. Mol Biol Cell 19:4762–4775
Taguchi K, Motohashi H, Yamamoto M (2011) Molecular mechanisms of the Keap1-Nrf2 pathway in stress response and cancer evolution. Genes Cells 16:123–140
Takahashi Y, Coppola D, Matsushita N, Cualing HD, Sun M, Sato Y et al (2007) Bif-1 interacts with Beclin 1 through UVRAG and regulates autophagy and tumorigenesis. Nat Cell Biol 9:1142–1151
Takamura A, Komatsu M, Hara T, Sakamoto A, Kishi C, Waguri S et al (2011) Autophagy-deficient mice develop multiple liver tumors. Genes Dev 25:795–800
Thurston TL, Ryzhakov G, Bloor S, von Muhlinen N, Randow F (2009) The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat Immunol 10:1215–1221
Vives-Bauza C, Zhou C, Huang Y, Cui M, de Vries RL, Kim J et al (2010) PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc Natl Acad Sci USA 107:378–383
Warren JR, Lalwani ND, Reddy JK (1982) Phthalate esters as peroxisome proliferator carcinogens. Environ Health Perspect 45:35–40
Wei Y, Pattingre S, Sinha S, Bassik M, Levine B (2008) JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol Cell 30:678–688
Wild P, Farhan H, McEwan DG, Wagner S, Rogov VV, Brady NR et al (2011) Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333:228–233
Xie Z, Klionsky DJ (2007) Autophagosome formation: core machinery and adaptations. Nat Cell Biol 9:1102–1109
Youle RJ, Narendra DP (2011) Mechanisms of mitophagy. Nat Rev Mol Cell Biol 12:9–14
Yue Z, Jin S, Yang C, Levine AJ, Heintz N (2003) Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA 100:15077–15082
Zatloukal K, Stumptner C, Fuchsbichler A, Heid H, Schnoelzer M, Kenner L et al (2002) p62 Is a common component of cytoplasmic inclusions in protein aggregation diseases. Am J Pathol 160:255–263
Zhao Z, Fux B, Goodwin M, Dunay IR, Strong D, Miller BC et al (2008) Autophagosome-independent essential function for the autophagy protein Atg5 in cellular immunity to intracellular pathogens. Cell Host Microbe 4:458–469
Zhong Y, Wang QJ, Li X, Yan Y, Backer JM, Chait BT et al (2009) Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat Cell Biol 11:468–476
Acknowledgement
We would like to thank Dr. S. Kageyama (Tokyo Metropolitan Institute of Medical Science) for illustrating figures.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Ichimura, Y., Komatsu, M. (2013). Selective Autophagy and Cancer. In: Wang, HG. (eds) Autophagy and Cancer. Current Cancer Research, vol 8. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6561-4_6
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6561-4_6
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6560-7
Online ISBN: 978-1-4614-6561-4
eBook Packages: MedicineMedicine (R0)