Abstract
Prostate Cancer (PCa) genetic risk has recently been defined in numerous genome-wide association studies (GWAS), which revealed more than 50 disease-associated single nucleotide polymorphisms (SNPs), known as tagSNPs, each at a different locus. More than 80% of these tagSNPs are located in noncoding regions of the genome for which functionality remains unknown. We and others hypothesize that at least some of these SNPs affect noncoding genomic regulatory signatures such as enhancers. Many research laboratories including ours have profiled the genomic distribution of androgen receptor (AR) and the dynamic state of the PCa genome for active chromatin regions (H3K9,14ac), open chromatin regions (DNaseI), enhancers (H3K4me1/2), and active/engaged enhancers (H3K27ac). In order to identify candidate functional SNPs, which may confer risk associated with PCa, we recently developed an open-source (R/Bioconductor) package, called FunciSNP (Functional Integration of SNPs), which systematically integrates SNPs from the 1000 genomes project with these biologically active chromatin features. Here we report several potential AR enhancers, defined by genome-wide data and from chromatin biofeatures, which may be functionally involved in PCa risk.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Enhancer
- Androgen Receptor Occupied Regions (ARORs)
- Chromatin
- Genome
- Single Nucleotide Polymorphism
- Post-GWAS function
5.1 Introduction
Within the past 5 years, prostate cancer (PCa) genome-wide association studies (GWAS) have identified more than 50 risk loci, with more than 80% found in noncoding genomic regions (Fig. 5.1) [1–4]. Integrating data associated with genomic profiles of androgen receptor (AR) occupancy and other enhancers to these known risk regions will inevitably provide an informative genomic landscape that may yield insight and understanding into the etiology and ultimately treatment of PCa. Characterizing and understanding the role of the AR in prostate cancer and the dynamic interactions between AR and nucleosomes is an important area of PCa research [5, 6] and consequently is hypothesized to play “driver” roles at GWAS risk loci. The aim of this chapter is to describe recent research in profiling the AR interactions with genomic features. In addition, we will present and describe the use of a novel bioinformatic tool (developed by our group), which integrates several large-scale data types to identify well-demarcated new genomic risk regions at known risk loci that may have important implications for PCa genetic predisposition.
Summary of all known GWAS SNPs across a number of different cancer types. GWAS SNPs were annotated using known genomic features supplied by HOMER (version 3.9) [43], using build hg19 as reference. GWAS SNPs were extracted from the UCSC Genome table browser, track name “gwasCatalog” with a P-value cut-off of 9e−06 [2]. Figure and legend reproduced from Coetzee SG. et al. 2012 [3] with permission from Oxford University Press
5.1.1 Normal Function of AR in Prostate
Understanding the role of AR in PCa and the regulation of AR activity remains an intense and important area of research. In normal prostate development and maintenance, AR drives differentiation and regulates the expression of select genes for the production of proteins that mainly function in seminal fluid development, similar to the role estrogen receptor provides in the regulation and production of milk in the breast [7]. Therefore, AR is not normally classified as an oncogene during prostate development and maintenance, whereas its role in prostate tumorigenesis is indisputably as a driver and as such may be viewed as an oncogene during the progression of the disease.
5.1.2 AR Significance in PCa
As early as the 1940s, Dr. Charles B. Huggins (a Nobel laureate) discovered that steroid hormones such as androgen could be used to control the spread of prostate and breast cancer. Orchiectomy, the removal of the male testes, was found to be an effective surgical procedure for the treatment of advanced PCa, later replaced with GnRH agonist [8, 9]. In the 1950s, it was found that patients who relapse after orchiectomy respond better to andrenalectomy, which indicates that the tumors may be stimulated by residual androgens from adrenal glands [8, 9]. From the mid-1990s, multiple two-arm trials looked at the effects of castration versus castration + AR antagonist (such as flutamide, milutamide, cyproterone acetate, and bicalutamide) to block effects of residual androgen expression [8, 9]. The hypothesis at the time was to further block AR, which would provide an alternative to castration procedures. After hundreds of millions of dollars invested into researching the effect of these antagonists, it was found that there was minimal improvement, with modest responses to AR antagonists given at relapse after castration, with a general conclusion that more complete AR blockade was no more effective then castration alone [9].
During the same time, basic research labs found that AR and AR regulated genes were highly expressed in castration recurrent/resistant prostate cancer (CRPC). The AR gene was found to be amplified in ∼30% of CRPC and AR antagonists selected for AR mutations [7, 10]. By 2000, work by several research groups found that high intra-tumoral androgen levels in CRPC were synthesized from weak adrenal androgens, and this led to the hypothesis that androgens were being developed in a de novo fashion from cholesterol products [9, 11, 12]. These observations provided important clinical application, and as recent as 2011, survival advantage from post- and pre-chemotheraphy for Abiraterone (CYP17A1 inhibitor, an enzyme for androgen synthesis) in CRPC was found to decrease serum testosterone from ∼20–50 ng/dl to <1 ng/dl [11, 12]. These clinical studies provided evidence that AR to some degree is driving PCa progression [11, 12]. In 2012, survival advantage for a novel AR antagonist, MDV3100, was found to be highly selective [13–15]. Despite progressive AR decreasing during tumor progression, tumors have found many genetic means of maintaining AR mediated signaling. We know that most tumors that have become castration resistant remain dependent on AR mediated activity, which has been shown with the advancement of drugs such as MDV3100, which continue to target AR pathway axis [10, 15]. Even more recently, exome sequencing of 50 CRPCs revealed many somatic mutations in the androgen-signaling pathway, DNA repair, and histone/chromatin modifier genes [16]. These and other important findings shed both clinical and biological importance in determining the mechanisms mediating intrinsic and acquired resistance to androgen and AR mediated pathways.
5.1.3 Advancement in Sequencing Technology Allows for Comprehensive Profiling of Specific Genomic and Epigenomic Marks
The last 15 years of biomedical research has been fueled in part by development of new technologies, techniques, and protocols. The recent advent of next-generation sequencing has not only spawned many new exciting applications and opportunities but also challenges for cancer researchers. ChIP-seq is a technique to systematically detect genome-wide protein:DNA interactions and histone modifications. In a given landscape of the genome, there are a variety of diverse proteins that interact with our genome in a sequence and tissue-specific manner. During a typical ChIP-seq experiment, the cells are treated with formaldehyde to stably cross-link all protein:DNA complexes. The chromatin is then sheared and fragmented to approximately 200–1,000 bp by sonication. Generally, such an experiment is done with about 2 million cells to enrich the amount of protein:DNA complex using a specific antibody targeting a protein of interest. Once the purified and enriched DNA fragments are obtained, high-throughput sequencing is performed and fragments of 50–100mers are mapped back to the reference genome. Currently, deep sequencing technology can sequence up to 100 million reads for a given experiment, and this number is very likely to increase exponentially while the overall cost of sequencing will drop over the next 5–10 years [17, 18]. Analyzing deep sequencing data for a particular ChIP experiment requires several important steps, which include initial quality control, peak calling, peak quality assessment, motif analysis, and large prediction and annotations of the data [17]. There are many different peak calling algorithms available, including MACS2 and FindPeaks [17].
In Sect. 5.2, we will describe the current understanding of AR occupancy regions (ARORs) in prostate cancer cell lines and associated enhancer elements in promoters and non-promoters of genes profiled using deep sequencing technologies. Section 5.3 will introduce the current state of GWAS in PCa and the idea of a novel bioinformatic tool we developed to integrate data profiled from deep sequencing, GWAS, and the 1000 genomes project to identify candidate regulatory elements involved in PCa progression and/or risk.
5.2 AROR Identification and Genome-wide Distribution of Enhancer Elements
Myles Brown and his research group was among the first to systematically identify thousands of AR binding sites in PCa cells after DHT treatment by AR chromatin immunopreciptation (ChIP) followed chip hybridization (initially) or by deep sequencing (more recently) (ChIP-seq; described in detail in Sect. 5.1.3) [19]. In addition, we and others profiled AROR in DHT-treated LNCaP cells. We reported that about 20% of histone H3 acetylated at Lys-9 and Lys-14 (H3K9,14ac) enrichment surrounds ARORs [20]. We also observed a high number of AROR peaks distributed in noncoding regions, while a significant enrichment of ARORs was found in promoters of known genes only with overlapping H3K9,14ac (Fig. 5.2a, b). If ARORs overlapping and not overlapping known H3K9,14ac peaks are separated, we observe that the genomic distribution to all known transcription start sites are significantly depleted at promoters for AROR peaks not overlapping H3K9,14ac (Fig. 5.2c, d). Interestingly, AROR peaks overlapping H3K9,14ac are enriched at promoters (Fig. 5.2e). This genomic profile of AROR has given researchers an unprecedented view of the wide distribution of AR binding to the human genome in PCa after DHT stimulation. It has been determined that AR binding sites have AR element (ARE) motif and are generally associated with a consensus motif for FOXA1 [19, 21]. In our own study, we performed de novo motif analysis on our identified AROR peaks. Significantly identified AROR peaks are highly enriched for the known ARE motif, while ARORs located in distant or non-promoters were also enriched for the FOXA1 motif (Fig. 5.3). It has been shown that FOXA1 binds with AR and other steroid receptors in a diverse set of tissues [19]. Among other factors, FOXA1 has been shown to function as a pioneer factor to open the locus prior to AR binding [22]. Further work by the Brown lab and others have found that sites occupied by FOXA1 (and the recruitment of FOXA1) are highly associated with enhancers, since these regions usually have nucleosomes with enhancer histone marks such as H3K4me1 and H3K4me2 [19, 20, 23] (Table 5.1). Prior to androgen stimulation and AR binding, ARE is covered with loosely associated nucleosome that contains a variant of H2A.Z which is displaced by AR [24]. At the chromatin level, AR was found enriched at ARE sites associated with H3K4me1 and H3K4me2. These distinct epigenetic marks are highly associated with FOXA1, which provides recruitment in CRPC [25]. This process is reversed by LSD1 (lysine-specific demethylase 1), which mediates demethylation of H3K4me1 and H3K4me2 [25]. Thus it appears the sequence of events start with the epigenetic alterations of histones, specifically methylation of K4 on the H3 complex, and recruitment of FOXA1, thereby allowing for an open conformational change of the nucleosome, thereby allowing for poised interactions for AR stimulated by DHT to incorporate with ARE, and thereby activate targeted genes responsible for progression and differentiation of the prostate.
Complete genomic distribution of AROR and H3K9,14ac. (a–b) raw counts and log2 ratio (peaks versus background) for H3K9,14ac (AcH) and AR distribution across known genomic regions, 3′UTR, 5′UTR, exon, intergenic, intron, promoter-TSS (promoter transcription start site), TTS (transcription termination site). Each column of plots indicates differing sets of peaks, All—total peaks per type (AcH or AR), No overlap (distinct peak files), overlap (peak files in common between AcH and AR). Black color indicates observed peaks, while gray colors indicate background as computed 1,000 times (randomly generated regions selected genome wide). (c–e) Genomic profile of AROR as a function of distance to the nearest canonical gene’s TSS. ARORs were divided into (c) total, (d) AROR not overlapping AcH, (e) AROR overlapping AcH. Red color indicates observed peaks; while gray colors indicate background as computed 1,000 times (randomly generated regions selected genome wide)
Further subsequent findings in advanced castration resistant PCa using LNCaP-abl cells have shown that AR regulates the expression of specific G2/M genes (CDK1, UBE2C, others) in advanced castration resistant but not in androgen sensitive LNCaP cells [23]. In summary, the spectrum of AR regulated genes may change during tumor development and progression.
Recently, it was discovered that FOXA1 suppressive sites are enriched for insulator protein binding sites, which are sites of compact chromatin (CTCF-FOXA1-Groucho family complexes) [26, 27]. They observed, by knocking down FOXA1 followed by ChIPseq of AR binding sites, hundreds of AREs are dependent of FOXA1 while if FOXA1 is deleted, 3,500 AR binding sites are revealed and are independent of FOXA1. These sites are enriched for insulator protein binding sites, CTCF–FOXA1, and set of corepressive elements [26, 27]. The overall physiological importance is unclear, but these studies illustrate that the dynamic integrity of AR binding to the DNA is quite flexible and provides a complex environment with a multitude of diverse functions yet to be fully characterized.
DNaseI hypersensitivity (HS) is a method to assay chromatin accessibility [28]. It allows for a comprehensive profile of distinct genomic regions, which are preferentially unbound by nucleosomes and thus are characterized as open chromatin region. DNaseI sequencing results are aligned to the human genome reference, compared to a background control experiment and peaks are called as described in Sect. 5.1.3, to identify candidate open chromatin regions in LNCaP cells. Most studies have shown that DNaseI hypersensitive sites correlate very well with regulatory elements and enhancer activity in a variety of tissues and cell types [22, 29]. We put together a table, which summarizes the total number of AR and available enhancer peaks across a number of different studies, as well as the overlap with data downloaded from the ENCODE project [30] (Table 5.1).
5.3 Identification of Significant Risk Variants Associated with PCa
5.3.1 Statistical Analysis and Bioinformatics Tool to Identify Candidate Functional Enhancer Elements
GWAS have yielded numerous single nucleotide polymorphisms (SNPs) associated with many phenotypes. The main goals for genetic risk factor identification have been to allow clinicians and epidemiologist a factor to predict risk for a disease or treatment such as chemotherapy or surgical prevention. It has also been important to allow researchers an opportunity to understand the biology of cancer by identifying genes or loci involved in the development of the cancer and finally to identify novel genetic regions or loci for drug targeting and treatment.
The first GWAS study published in 2005 identified a risk region for age-related macular degeneration disease [31], and GWAS have been constantly evolving and expanding their reach in large part due to the advancement of genotyping technology. During the last 6 years, it is apparent that GWAS efforts fostered by these advancements have made significant discoveries, for example, more than 1,449 tagSNPs have been published for association with well over 200 different diseases or traits at a p-value of 5 × 10−8 [2, 3]. It is also apparent that as new methods and stratifications by populations emerge, these studies could yield even higher numbers of variations associated to a particular disease or trait. Interestingly, approximately 200 genomic variants have been reported for more than 20 different cancer types, and it is expected that this number of associated risk loci will likely exceed 300 by the end of this year [32]. Remarkably, we and others have noted that the vast majority of these variants associated with cancer, in particular with prostate and breast, are enriched in noncoding regions (Table 5.2, Fig. 5.1) [3]. Despite the many novel and insightful biological discoveries GWAS have provided to the scientific community, we have a long way to go before we can fully understand genetic heritability and risk. In the case of breast cancer, there are very rare allele frequencies in a given population with very high effect size, many identified through family-based studies, candidate gene re-sequencing, and regions with candidate risk association. The rare variance contributes on total, over 25% to understanding the familial risk of most cancer whereas only 10–15% contributes to common risk with low to modest effect size [33]. However, the field is currently looking to see if the risk variant for low-to-common allele frequency can be identified with intermediate to large effect size, which is believed to be largely due to genetic factors. Currently, with the advancement in sequencing technology and methodology, we are at the forefront of making discoveries of these low to common variants with measurable effect size with suitable power. And as the advancement of sequencing technology increases and the cost to perform such high-throughput deep sequencing reaches a critical cost–benefit level, the major bottleneck will be computational methods and tools needed in order to tease out the genetic variants and risk associated with many of these critically debilitating human diseases.
At the core of all association studies dominated during the past 6–7 years is the idea of linkage disequilibrium (LD). LD is the observation that two or more alleles in a population segregating together more often than one would expect by chance [3, 34]. What this definition means to researchers interested in identifying genetic basis or identify candidate functional variants in a particular disease is that the identity of the functional variant is not required. As an example, a causal or putative functional variant is located on the ancestral allele, and this variant will segregate successfully through the population over time. If we are able to visualize this variant, then we can measure LD and therefore indirectly identify the variant using markers or tags surrounding the causal or putative functional variant [35, 36].
SNPs are the most common form of variation we have measured in the human population, and therefore SNP markers provide the best opportunity to identify the ancestral causing or functional allele in a given population exhibiting a shared or common disease or trait. However, as the ancestral causing allele passes through time in a population (over numerous generations), the LD breaks down significantly. This breakdown in LD structure then makes identification of variant causing allele more difficult. Therefore, a systematic identification of all known SNPs or variants is required and to this end, the HapMap project evolved. The HapMap project aim was to identify SNPs using four different human populations to put together a phased haplotype block map and identifiable LD blocks of the entire human genome [37]. This project was successful at the time, identifying 4–5 million SNPs in the profiled population. Although these blocks identified regions of interest, it did not successfully identify the candidate or functional variant [38]. Therefore, the 1000 genomes project (1000 gp) emerged to tackle the issue by harnessing the obvious observation that the world’s population is quite diverse than the original four populations profiled from the HapMap project [39]. The 1000 gp set out to sequence more than 1,000 individuals across a larger number of different populations. Currently, the 1000 gp have identified more than 60 million SNPs, indels, somatic mutations, and other genomic variants across the population [39]. This has now allowed us to identify a larger number of variants among a given haploblock containing the identified risk allele. Some have argued this increase in variants coupled with unidentified functional variant provides an even bigger challenge in identifying the causal or functional variant. However, we and others have noted that this increase in number of identified variants provides a deeper resolution of the heritability and thus allows us to harness more informative variants than previously identified [3, 40]. We developed a bioinformatic tool called FunciSNP (Functional Identification of SNP), which allows us to harness the information provided by the 1000 gp, the location of the previously reported risk allele (tagSNP), and the chromatin features to identify the causal and functional regions associated with PCa [3] (Fig. 5.4b) (for results, see Sect. 5.3.3–5.3.4).
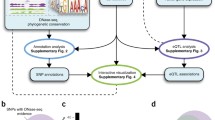
GWAS integration with biological features. (a) Circos plot illustrating genomic distribution of the 51 known GWAS tagSNPs as of June 2012 (red lines indicate the position of the tagSNPs with the length reflecting the number of independent tagSNPs within a 10 MB window. The white concentric circles are markers for 1, 2, and 3 SNPs; outside to inside). From outermost to innermost, the colored bands describe the concentration (red, yellow, green, blue; high to low) DNase1 sensitivity, H3K27ac, H3K4m31, AROR, and genes, respectively. The chromatin marks were obtained from LNCaP cells. On the ideograms the locations of the centromeres are depicted in green and tagSNPs in red. (b) Schematic flowchart to describe FunciSNP. Purple boxes represent process before integration with biofeature. Red boxes represent information after integration with biofeature (Figure and legend reproduced from Coetzee SG. et al. 2012 [3] with permission from Oxford University Press.). (c) Schematic diagram indicating total number of candidate SNPs pooled from the 1000 gp that overlap one or more biological genomic features and which are in linkage disequilibrium to the original GWAS SNP. Total number of SNPs in each section is identified by a blue box. (d) Overall distribution of R 2 values for all identified 1000 gp SNPs overlapping at least one biological feature. Each identified bin reports total number of candidate 1000 gp SNPs
5.3.2 GWAS PCa
PCa genetic risk has recently been defined in a number of GWAS, which have revealed 51 disease-associated SNPs, known as tagSNPs, each at a different locus (Table 5.2, Figs. 5.1 and 5.4a). More than 80% of these tagSNPs are in intergenic or intron regions of the genome for which functionality remains unknown. Interestingly, the tagSNPs are distributed among many chromosomes (Fig. 5.4a) with the exception being that chromosomes 1, 14, 15, 16, 18, 20, and 21 do not contain any tagSNPs. They occur at relatively gene rich areas (inner circle of Fig. 5.4a), with the notable exceptions of the tagSNPs at chromosome 3 (middle SNP), 8q, and the distal SNP at 12q. Many tagSNPs track closely with chromatin biofeatures (outer four circles of Fig. 5.4a, and see below).
Taking the 1000 genomes data into consideration, more than 10,000 correlated SNPs are revealed, which each define risk due to linkage disequilibrium (correlated to the tagSNP with an R 2>0.8, Fig. 5.4c). The above therefore makes the identification of functional and/or causal SNPs not a trivial task. Therefore, we hypothesize that at least some of these SNPs affect noncoding genomic regulatory signatures, such as enhancers or insulators. As we described in Sect. 6.2, there are distinct regions in the genome profiled in PCa cells, which we and others have identified and have significant biological relevance in PCa.
5.3.3 FunciSNP Integrates AROR, Enhancer, GWAS and 1000gp to Identify Candidate Functional Elements
In order to reduce the number of candidate functional SNPs profiled in the 1000gp linked to the associated tagSNP in PCa, we developed an open-source (R/Bioconductor) package, called FunciSNP, which systematically integrates the 1000gp SNP data with chromatin features of interest [3]. To define functionality in noncoding DNA associated with PCa risk, we extracted open chromatin and enhancer features generated by next-generation sequencing technologies. The open chromatin state (DnaseI, H3K9,14ac), enhancers (H3K4me1), and active/engaged enhancers (H3K27ac) were generated either by our lab [20], or harvested from the ENCODE project [41], or retrieved from recent publications [22, 26] (Table 5.1 and visualized in Fig. 5.4a). All chromatin features were identified in the same PCa cell line (LNCaP). We identified 113 PCa risk correlated SNPs at androgen receptor occupied regions (ARORs), 1,545 at DNase1 sensitivity sites, and 403 at histone modified regions [H3K4me1 (160 SNPs), H3K9,14 ac (121 SNPs), H3K27ac (122 SNPs)]; all features excluded transcription start sites of known annotated genes (Figs. 5.4d, 5.5, 5.6a). ARORs coinciding with DNase1 sites revealed four novel SNPs correlated with four GWAS tagSNPs. Of the four novel surrogate SNPs, two are located 4 kb upstream from KLK3, one is located within the 3’UTR of NKX3.1, and one is located within an intron of RUVBL1 (Fig. 5.6a; http://goo.gl/cYCl7). DNase1 sites coinciding with any histone modification provided 12 novel risk regions correlated with 10 GWAS tagSNPs (Tables 5.3 and 5.4). Four are located in 8q24 genomic region and each is more than 100 kb away from a known annotated gene.
FunciSNP heatmap of the number of 1000 gp between tagSNP and biofeature for PCa. Total number of candidate SNP is listed by color within each quadrant to represent the number of potential candidate functional SNPs overlapping a biofeature (y-axis), which are in linkage disequilibrium to the original GWAS risk SNP (x-axis)
Genomic plots with FunciSNP results and independent validation on eight enhancer elements harboring a candidate functional SNP, performed in three different cell types. (a) UCSC genome browser tracks are ordered in the following manner: dbSNP135, FunciSNP result, biofeatures, refseq genes, and known lincRNA. TagSNP is highlighted in the FunciSNP result track, and each candidate SNP is color coded to reflect the number of biofeatures which it overlaps. The color ranges from blue (low number of biofeature overlap) to red (high number of overlap). Each candidate function SNP is identified by its known rsID and the calculated R 2 value to the known GWAS tagSNP. The results are saved in a UCSC genome session: http://goo.gl/cYCl7. (b) Enhancer activities were evaluated using a dual luciferase reporter assay by cloning eight candidate enhancer regions harboring a candidate functional SNP in three different prostate cell types [LNCaP, PC3 (+AR), and PREC (normal primary prostate epithelial cells) (+AR)]. In order to stimulate AR, prostate cells were treated with 10 nM DHT. Orange–pink color represents cells treated with DHT and green–blue represents cells treated without DHT. Enhancer activities of positive control region (PSA enhancer) (n = 1) and negative control regions (n = 2) are presented to the left. Dashed horizontal line indicates the average enhancer activities of the two negative controls. Error bar indicates the standard deviation from the mean of luciferase activity values from four independent experimental replicates
5.3.4 Enhancer Validation in Two PCa Cell Lines and in Primary Prostate Epithelial Cells
To measure potential enhancer activities in the eight chosen potential enhancer regions, we employed a firefly luciferase reporter (driven by a basal tk-promoter) into which ∼1.2 kb potential enhancers coinciding with PCa risk correlated SNPs were cloned. These vectors, along with renilla luciferase controls, were transfected into two PCa cell lines: LNCaP cells, PC3 cells along with an AR expression vector, and normal primary prostate epithelial cells (PREC) along with an AR expression vector and stimulated with DHT. Enhancer activities were measured as previously described [42] and compared with two negative controls (regions with no enhancer histone marks) and a positive control (PSA enhancer). DHT-stimulated enhancer activity for seven of the eight potential enhancers was evident in at least one cell type (the exception is F26), indicating that the identification of enhancers using our protocol is both high and specific (Fig. 5.6b). Further analyses are required to determine the role(s) of these enhancers in PCa risk.
5.4 Future Perspective/Discussion
During the past decade three major findings in human genetics/genomics have caused a dramatic change in our appreciation of our genome and how it functions. First, the human genome project revealed that humans have only some 22,000 genes within the 3 billion-nucleotide genome. Second, many transcription factors (such as the AR discussed above) do not preferentially occupy gene promoters preferentially but are rather scattered among the many noncoding stretches of DNA in introns and intergenic regions. Third, as pointed out above, GWAS signals for complex diseases are preferentially found in introns and intergenic regions. These three novel insights have pointed to the importance of so-called noncoding DNA (previously even referred to as junk DNA). Transcription factor occupancy, histone modification marks, and nucleosome-depleted regions in chromatin are presently used to annotate functionality within most of our genome. Ultimately this will yield important insight to understand gene expression, genetic risk of complex diseases, and genetic associations with most human phenotypes.
Abbreviations
- AR:
-
Androgen receptor
- DHT:
-
Dihydrotestosterone
- ARE:
-
Anrogen responsive element
- ARORs:
-
Androgen receptor occupied regions
- GWAS:
-
Genome-wide association studies
- 1000gp:
-
1000 genomes project
- SNP:
-
Single nucleotide polymorphisms
- FunciSNP:
-
Functional Identification of SNPs
- LD:
-
Linkage disequilibrium
References
Visscher PM et al (2012) Five years of GWAS discovery. Am J Hum Genet 90(1):7–24
Hindorff LA et al (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 106(23):9362–9367
Coetzee SG et al (2012) FunciSNP: an R/bioconductor tool integrating functional non-coding data sets with genetic association studies to identify candidate regulatory SNPs. Nucl Acids Res 40:e139
Coetzee GA (2012) The usefulness of prostate cancer genome-wide association studies. J Urol 187(1):9–10
Coetzee GA et al (2010) A systematic approach to understand the functional consequences of non-protein coding risk regions. Cell Cycle 9(2):47–51
Heinlein CA, Chang C (2004) Androgen receptor in prostate cancer. Endocr Rev 25(2):276–308
Bluemn EG, Nelson PS (2012) The androgen/androgen receptor axis in prostate cancer. Curr Opin Oncol 24(3):251–257
Denmeade SR, Isaacs JT (2002) A history of prostate cancer treatment. Nat Rev Cancer 2(5):389–396
Cai C, Balk SP (2011) Intratumoral androgen biosynthesis in prostate cancer pathogenesis and response to therapy. Endocr Relat Cancer 18(5):R175–R182
Lonergan PE, Tindall DJ (2011) Androgen receptor signaling in prostate cancer development and progression. J Carcinog 10:20
Montgomery RB et al (2008) Maintenance of intratumoral androgens in metastatic prostate cancer: a mechanism for castration-resistant tumor growth. Cancer Res 68(11):4447–4454
Mostaghel EA et al (2010) Variability in the androgen response of prostate epithelium to 5alpha-reductase inhibition: implications for prostate cancer chemoprevention. Cancer Res 70(4):1286–1295
(2011) Triple-acting drug boosts prostate cancer survival. Cancer Discov 1(7): OF1
Tran C et al (2009) Development of a second-generation antiandrogen for treatment of advanced prostate cancer. Science 324(5928):787–790
Mukherji D, Pezaro CJ, De-Bono JS (2012) MDV3100 for the treatment of prostate cancer. Expert Opin Investig Drugs 21(2):227–233
Grasso CS et al (2012) The mutational landscape of lethal castration-resistant prostate cancer. Nature 487:239–243
Chen Y et al (2012) Systematic evaluation of factors influencing ChIP-seq fidelity. Nat Methods 9:609–614
Qin B et al (2012) CistromeMap: a knowledgebase and web server for ChIP-Seq and DNase-Seq studies in mouse and human. Bioinformatics 28(10):1411–1412
Lupien M et al (2008) FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132(6):958–970
Andreu-Vieyra C et al (2011) Dynamic nucleosome-depleted regions at androgen receptor enhancers in the absence of ligand in prostate cancer cells. Mol Cell Biol 31(23):4648–4662
Wang Q et al (2009) Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell 138(2):245–256
Wang D et al (2011) Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474(7351):390–394
Zhang C et al (2011) Definition of a FoxA1 Cistrome that is crucial for G1 to S-phase cell-cycle transit in castration-resistant prostate cancer. Cancer Res 71(21):6738–6748
Dryhurst D et al (2012) Histone H2A.Z prepares the prostate specific antigen (PSA) gene for androgen receptor-mediated transcription and is upregulated in a model of prostate cancer progression. Cancer Lett 315(1):38–47
Cai C et al (2011) Androgen receptor gene expression in prostate cancer is directly suppressed by the androgen receptor through recruitment of lysine-specific demethylase 1. Cancer Cell 20(4):457–471
Sahu B et al (2011) Dual role of FoxA1 in androgen receptor binding to chromatin, androgen signalling and prostate cancer. EMBO J 30(19):3962–3976
Taslim C et al (2012) Integrated analysis identifies a class of androgen-responsive genes regulated by short combinatorial long-range mechanism facilitated by CTCF. Nucleic Acids Res 40(11):4754–4764
Song L et al (2011) Open chromatin defined by DNaseI and FAIRE identifies regulatory elements that shape cell-type identity. Genome Res 21(10):1757–1767
He HH et al (2012) Differential DNase I hypersensitivity reveals factor-dependent chromatin dynamics. Genome Res 22(6):1015–1025
Myers RM et al (2011) A user’s guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol 9(4):e1001046
Rivera A et al (2005) Hypothetical LOC387715 is a second major susceptibility gene for age-related macular degeneration, contributing independently of complement factor H to disease risk. Hum Mol Genet 14(21):3227–3236
Hosking FJ, Dobbins SE, Houlston RS (2011) Genome-wide association studies for detecting cancer susceptibility. Br Med Bull 97:27–46
Park JH et al (2010) Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat Genet 42(7):570–575
Kruglyak L (2008) The road to genome-wide association studies. Nat Rev Genet 9(4):314–318
Hirschhorn JN, Daly MJ (2005) Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 6(2):95–108
Hardy J, Singleton A (2009) Genomewide association studies and human disease. N Engl J Med 360(17):1759–1768
International HapMap Consortium (2003) The International HapMap Project. Nature 426(6968):789–796
Pennisi E (2010) Genomics. 1000 Genomes Project gives new map of genetic diversity. Science 330(6004):574–575
Consortium TGP (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073
Freedman ML et al (2011) Principles for the post-GWAS functional characterization of cancer risk loci. Nat Genet 43(6):513–518
Rosenbloom KR et al (2011) ENCODE whole-genome data in the UCSC Genome Browser: update 2012. Nucl Acids Res 40:D912–D917
Jia L et al (2009) Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet 5(8):e1000597
Heinz S et al (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38(4):576–589
Takata R et al (2010) Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat Genet 42(9):751–754
Eeles RA et al (2009) Identification of seven new prostate cancer susceptibility loci through a genome-wide association study. Nat Genet 41(10):1116–1121
Kote-Jarai Z et al (2011) Seven prostate cancer susceptibility loci identified by a multi-stage genome-wide association study. Nat Genet 43(8):785–791
Schumacher FR et al (2011) Genome-wide association study identifies new prostate cancer susceptibility loci. Hum Mol Genet 20(19):3867–3875
Murabito JM et al (2007) A genome-wide association study of breast and prostate cancer in the NHLBI’s Framingham Heart Study. BMC Med Genet 8(Suppl 1):S6
Gudmundsson J et al (2009) Genome-wide association and replication studies identify four variants associated with prostate cancer susceptibility. Nat Genet 41(10):1122–1126
Eeles RA et al (2008) Multiple newly identified loci associated with prostate cancer susceptibility. Nat Genet 40(3):316–321
Thomas G et al (2008) Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet 40(3):310–315
Zheng SL et al (2007) Association between two unlinked loci at 8q24 and prostate cancer risk among European Americans. J Natl Cancer Inst 99(20):1525–1533
Yeager M et al (2007) Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet 39(5):645–649
Gudmundsson J et al (2007) Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat Genet 39(5):631–637
Duggan D et al (2007) Two genome-wide association studies of aggressive prostate cancer implicate putative prostate tumor suppressor gene DAB2IP. J Natl Cancer Inst 99(24):1836–1844
Liu H, Wang B, Han C (2011) Meta-analysis of genome-wide and replication association studies on prostate cancer. Prostate 71(2):209–224
Chung CC et al (2011) Fine mapping of a region of chromosome 11q13 reveals multiple independent loci associated with risk of prostate cancer. Hum Mol Genet 20(14):2869–2878
Zheng SL et al (2009) Two independent prostate cancer risk-associated Loci at 11q13. Cancer Epidemiol Biomarkers Prev 18(6):1815–1820
Bonilla C et al (2011) Prostate cancer susceptibility Loci identified on chromosome 12 in African Americans. PLoS One 6(2):e16044
Sun J et al (2008) Evidence for two independent prostate cancer risk-associated loci in the HNF1B gene at 17q12. Nat Genet 40(10):1153–1155
Haiman CA et al (2011) Genome-wide association study of prostate cancer in men of African ancestry identifies a susceptibility locus at 17q21. Nat Genet 43(6):570–573
Hsu FC et al (2009) A novel prostate cancer susceptibility locus at 19q13. Cancer Res 69(7):2720–2723
Sun J et al (2009) Sequence variants at 22q13 are associated with prostate cancer risk. Cancer Res 69(1):10–15
Acknowledgments
The authors thank Charles Nicolet at the USC Epigenome Center for library construction and high throughput sequencing.
Funding
Original work reported here was funded by the National Institutes of Health (NIH) [CA109147, U19CA148537 and U19CA148107 to G.A.C.; 5T32CA009320-27 to H.N.] and David Mazzone Awards Program (G.A.C). Additionally, some of the scientific development and funding of this project were supported by the Genetic Associations and Mechanisms in Oncology (GAME-ON): a NCI Cancer Post-GWAS Initiative.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Noushmehr, H., Coetzee, S.G., Rhie, S.K., Yan, C., Coetzee, G.A. (2013). The Functionality of Prostate Cancer Predisposition Risk Regions Is Revealed by AR Enhancers. In: Wang, Z. (eds) Androgen-Responsive Genes in Prostate Cancer. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6182-1_5
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6182-1_5
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6181-4
Online ISBN: 978-1-4614-6182-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)