Abstract
DNA repair pathways maintain the integrity of the genome, reducing the onset of cancer, disease, and aging. The majority of anticancer therapeutics (radiation and chemotherapy) function as genotoxins, eliciting genomic DNA damage in an attempt to induce cell death in the tumor. However, cellular DNA repair proteins counteract the effectiveness of these therapeutic genotoxins by repairing and removing the cell death-inducing DNA lesions, implicating DNA repair proteins as prime targets for improving response to currently available anticancer regimens. To trigger a tumor-specific cell death response (with minimal normal cell toxicity), the level of genomic DNA damage must therefore surpass the DNA repair capacity of the tumor without overwhelming the DNA repair potential of normal tissue. Interestingly, cancer-specific DNA repair defects offer novel approaches for tumor-selective therapy. This has become highly relevant as it is suggested that most cancer cells are likely to be defective in some aspect of DNA repair. Herein, we describe the molecular pathways that participate in the repair of DNA damage induced by radiation- and chemotherapeutics and discuss strategies that are being developed to target DNA repair for cancer treatment and highlight key DNA repair inhibitors that can enhance response. Further, we present novel therapeutic strategies being considered to exploit inherent weaknesses in tumor cells such as defects in one or more DNA repair pathways or related processes that may provide the opportunity to selectively increase tumor-specific cell death.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Homologous Recombination
- Nucleotide Excision Repair
- Base Excision Repair
- Fanconi Anemia
- Replication Stress
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
6.1 Role of DNA Repair Pathways for Cancer Treatment
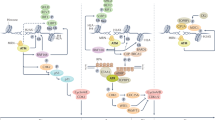
Human cells must repair tens of thousands of DNA lesions per day [1]. If they are not repaired, these lesions lead to mutations or genome aberrations that threaten cell survival and genomic integrity. To combat these threats, cells have evolved multiple DNA repair and DNA damage response (DDR) mechanisms that signal the presence of lesions and promote their repair or regulate cellular processes in response to the DNA damage (Fig. 6.1) [2]. Defects in these repair and response pathways can promote tumorigenesis and, indeed, are common in human cancers [3, 4]. On the other hand, current therapy options for cancer patients exploit the DNA-damaging properties of certain drugs and agents. The success of radiation exposure during radiotherapy and the success of most chemotherapy agents rely on the destructive nature that these agents have on cellular DNA, ultimately resulting in death and hopefully eradication of the tumor cells. Hence, DNA damage and repair mechanisms play a crucial role in determining treatment outcome. On a cellular level, resistance to treatment is profoundly determined by the capacity of the cancer cell to respond to and repair the individual DNA lesions that are induced by the chemotherapeutic agents or radiation.
Our increasing knowledge of these processes has led to the development of new concepts that target and exploit the cancer cell DDR [5]. Counteracting resistance to chemotherapy by targeting the appropriate DNA repair pathway is a promising strategy in cancer treatment. Another is to exploit the DNA repair and response defects that are present in cancer cells thereby specifically targeting tumor cells while sparing healthy cells from a high load of unrepaired DNA damage [6, 7]. Together, these strategies might provide promising avenues in the conquest against cancer. Here, we will briefly describe essential cellular DNA repair and response mechanisms and illustrate novel concepts and promising strategies to exploit and target DNA repair for cancer treatment.
6.1.1 DNA Damage Response
The initial DDR of a cell involves the recognition of the DNA damage followed by the propagation of a series of signals ranging from alterations in RNA or protein expression and modification of protein function or stability through post-translation modification, among other signals. The cell’s defense to genotoxic lesions is triggered and accomplished by a series of events that mediate and regulate proliferation, cell death, or DNA repair crucial to its survival [2, 4]. The initial steps for an appropriate response require detection of the lesion, signaling of its presence and promotion of repair. Cells act upon DNA damage not only by promoting and executing repair but also respond by halting the cell cycle or by promoting cell death mechanisms in order to prevent propagation of the damage. The DDR therefore has an impact on transcription, cellular metabolism, cell cycle regulators, as well as cell death, via apoptosis and senescence.
One of the most prominent members of the DNA damage signaling pathway that links DNA damage with cell cycle checkpoints is the protein ataxia telangiectasia-mutated (ATM) [8], a protein kinase that is recruited to DNA double-strand breaks (DSB) such as those induced by ionizing radiation. The formation of DSBs triggers the activation of ATM and activated ATM then phosphorylates a wide range of downstream substrate proteins thereby signaling the presence of the damage throughout the cell to facilitate repair [9]. Initial activation of ATM is promoted by its autophosphorylation that initiates a signaling cascade of further phosphorylation events that constitute the DDR [10]. With excessive unrepaired DNA damage present, this cellular response can culminate in an apoptotic response in which p53 is central but not necessarily always required.
Another key DDR signaling component is the ataxia telangiectasia-RAD3-related (ATR) kinase that gets activated after replication stress-induced DNA damage. Replication stress, caused, for example, by exposure to hydroxyurea (HU), results in the formation of large stretches of single-stranded DNA coated with replication protein A (RPA) that triggers activation of ATR. Similar to processes in ATM-mediated signaling, ATR signaling is promoted by regulatory proteins. ATRIP and TopBP1, together with RAD17-mediated 9-1-1 (Rad9-Rad1-Hus1) clamp loading, “sense” the damage and trigger the activation of ATR.
Further downstream of these initial events, the cell cycle checkpoint-regulating protein kinases CHK1 and CHK2 are among the most important targets (substrates) of ATM and ATR. Supported by the activation of p53 and mediated via multiple paths, this signaling cascade ultimately results in the reduction of cyclin-dependent kinase activity that drives cell cycle progression. The halt in cell cycle progression is thought to allow time for repair and, most importantly, if not successfully repaired, to prevent propagation of the DNA damage. Cell cycle checkpoints are in place at the border from G1 to S, within S and at the G2/M border. The prevalent blocks and their extent depend on the damaging agent and the number and type of lesion. Another important downstream target of ATM and ATR is p53, an essential player in the induction of apoptosis upon DNA damage. Thus, DDR mechanisms have a crucial role in the protection against genome instability and chemo/radiotherapy response.
ATM/ATR signaling also enhances repair by recruiting repair factors to the site of the lesion and activating DNA repair proteins through phosphorylation or indirectly, by modulating acetylation, ubiquitylation, SUMOylation, or DNA repair gene transcription. These kinases also influence chromatin structure through phosphorylation of the histone H2A variant (γH2AX). Thereby, they facilitate recruitment of DDR factors and expedite DNA repair while amplifying DSB signaling that is crucial to cellular survival following exposure to DNA-damaging agents.
The role of the DDR is broad with respect to the type of cancer therapeutic. DDR activity is involved upon exposure of a whole range of chemotherapeutic agents and upon radiation. Indeed, DDR and cell cycle blocks are induced by radiation, topoisomerase I and II poisons, anthracyclines, alkylating drugs including platinum analogues and antitumor antibiotics. Interference in DDR by the use of inhibitors will likely affect the response and cellular survival in most cancer treatment options.
6.1.2 Direct-Reversal Repair and Mismatch Repair
One of the first DNA repair proteins to be considered as a viable target for improving chemotherapy was O6-alkylguanine-DNA alkyltransferase (MGMT or AGT), a protein encoded by the O6-methylguanine-DNA methyltransferase gene (MGMT) [11]. In depth, discussion on the function of MGMT and its role in cancer and chemotherapy can be found in many excellent reviews [12–14]. MGMT falls within the category of direct-reversal (DR) DNA repair proteins that also include the AlkB family of proteins [13, 15]. Unlike most other DNA repair pathways that correct lesions by removing the base containing the lesion [base excision repair (BER), see below], removing a short oligonucleotide containing the lesion [nucleotide excision repair (NER), see below] or removing long tracts of DNA (mismatch repair, MMR) followed by a DNA synthesis step (repair-directed DNA synthesis), DR proteins such as MGMT or the AlkB proteins reverse the damage to the DNA base directly, and the mechanism of repair does not involve a DNA synthesis step. This section will focus on the role of MGMT in DNA lesion repair and the subsequent role that the MMR proteins play in the cellular response when MGMT is unable to repair the O6-alkylguanine lesion. Further discussion on the mechanism of action of AlkB proteins can be found elsewhere [13].
Like many DNA repair proteins, MGMT repairs lesions from both carcinogenic compounds and from chemotherapeutic agents. As such, MGMT acts to suppress cancer formation by removing lesions induced by carcinogens such as methylnitrosourea (MNU), the tobacco smoke lung carcinogen 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone (NKK), and the colon carcinogen azoxymethane. Conversely, many chemotherapeutic agents, including Temozolomide (Temodar, TMZ), dacarbazine, streptozotocin, procarbazine, BCNU (camustine), CCNU (lomustine), and gliadel trigger cell death by inducing the formation of an alkyl lesion (methyl- or chloroethyl-) on the O6 position of guanine bases in DNA [12–14, 16]. Upon MGMT binding to the DNA containing the alkyl lesion, the O6-alkyl group is transferred from the guanine base onto a Cysteine (Cys) residue (amino acid residue Cys145 in humans) in the MGMT protein [12, 14]. Upon transfer of the alkyl group to MGMT, the protein undergoes a conformational change that both releases the protein from the repaired DNA and promotes the ubiquitylation and subsequent proteasome-mediated degradation. This suicide mechanism of MGMT has been taken advantage of clinically, as will be described below regarding the development and evaluation of MGMT inhibitors (see Sect. 6.3.2).
The chemotherapeutic agents mentioned above induce the formation of methyl or chloroethyl adducts on the O6 position of guanine bases in DNA. If not repaired, the majority of the chloroethyl lesions are converted to G-C interstrand DNA cross-links. Such lesions are primarily repaired by a concerted effort of the NER, HR, and fanconi anemia (FA) pathways (see below). In general, interstrand DNA cross-links are highly genotoxic, inducing cell death. Conversely, if the methyl lesion (O6-MeG) is not removed by MGMT, during cellular replication, the mispairing of O6-MeG with thymine leads to the formation of a O6-MeG:T mispair, a substrate for MMR. Repair mediated by the MMR pathway facilitates the removal of the DNA strand containing the newly synthesized “T” base. However, re-synthesis of the DNA in the process of MMR will regenerate the O6-MeG:T mispair, perpetuating the O6-MeG lesion and the presence of the mispair. As such, in the absence of MGMT-mediated repair, O6-MeG is suggested to initiate a futile cycle of MMR or alternately to trigger ATR protein kinase activation through the action of several MMR proteins [17], leading to apoptosis and cell death [18–20]. Details on the MMR pathway can be found elsewhere [21]. However, for the purpose of this discussion, we should consider the MMR pathway as an essential sensor to trigger cell death from chemotherapeutic agents that induce the O6-MeG lesion. Briefly, recognition of the O6-MeG:T mispair by the MMR protein MSH2 induces recruitment and activation of ATR and subsequently CHK1 and CHK2 to activate an apoptotic response [22, 23]. In fact, much of the resistance to agents such as TMZ observed clinically is due to high expression of MGMT (and subsequent repair of the lesion) or loss of MMR (therefore preventing the initiation of apoptotic signaling) [24–26]. Currently, TMZ along with radiation and surgery are the standard of care for glioblastoma multiforme (GBM), the most common and aggressive primary brain tumor [27]. Median survival is less than two years [28–30], and unfortunately, almost all patients eventually recur with the disease and the large majority of recurrent tumors are resistant to chemotherapy [31, 32]. Inhibition of MGMT-mediated repair has been taken advantage of experimentally and in many clinical trials [33] since MGMT can be inhibited with the O6-MeG analogue O6-benzylguanine [34] (Sect. 6.3.2). Improved prognosis has also been reported in tumors with loss of MGMT expression due to promoter methylation [35] whereas poor prognosis is observed when MGMT expression levels are high or MMR capacity is compromised. Hence, elevated expression of MGMT and/or a non-functional MMR pathway contribute much of the observed resistance to TMZ in many tumor cell lines and in clinical trials.
6.1.3 Base Excision Repair
As suggested in its namesake, the BER pathway is the primary mechanism to remove and repair base lesions. A special sub-pathway of BER, single-strand break repair (SSBR), is also essential for the repair of single-strand DNA breaks [13, 36, 37]. The types of base lesions repaired by the proteins of the BER pathway range from base deamination products (e.g., conversion of C to U or 5 meC to T) to oxidative modification of bases (8-oxo-7,8-dihydro-2′- deoxyguanosine; 8-oxodG), alkylation products such as N7-meG and N3-meA that are induced by chemotherapeutic agents such as TMZ [38] and many others [36, 37]. These and many other lesions are induced in genomic (and mitochondrial) DNA by a multitude of anticancer treatments including radiation, monofunctional alkylators such as TMZ, cisplatin, and 5FU, among others. There are over 20 proteins brought to bear to facilitate the complete process of BER [36]. Repair is initiated following recognition of the base lesion by one of the eleven DNA glycosylases in humans. This group of proteins is further subdivided into two classes: Bifunctional and monofunctional DNA glycosylases. A β-bifunctional glycosylase such as OGG1 excises the modified base and hydrolyzes the DNA backbone (via a β-elimination step) 3′ to the incised base, leaving a 3′ unsaturated aldehyde (after β-elimination) and a 5′ phosphate at the termini of the repair gap. Alternatively, a β,δ-bifunctional glycosylase such as NEIL1 hydrolyzes the glycosidic bond to release the lesion and then cleave the DNA backbone 3′ to the resulting apurinic/apyrimidinic (AP or abasic) site via β-elimination and 5′ to the abasic site via δ-elimination. More detail on the mechanism of these bifunctional DNA glycosylases and the specific BER proteins involved in processing the resulting repair gaps can be found elsewhere [36, 37].
For the purpose of describing the complete BER pathway, we will focus on the initiation of BER by monofunctional DNA glycosylases, with an emphasis on the methylpurine DNA glycosylase (MPG) (also called AAG or ANPG). MPG is the primary glycosylase for the repair of the chemotherapy-induced DNA lesions such as N7-meG and N3-meA. These lesions are removed by hydrolysis of the glycosidic bond, producing an abasic site, a substrate for AP-Endonuclease 1 (APE1). Given the highly toxic nature of the intermediates in BER [13], it has been suggested that the product of each BER reaction “hands off” the toxic BER intermediate to the next enzyme in the pathway likening the complete reaction to the hand-off of a baton in a relay race [39]. Such a process or hand-off mechanism has the advantage of eliminating or avoiding the accumulation of free BER intermediates that are prone to induce cell death [13]. Once formed by the glycosylase, the resulting abasic site is then handed off to APE1 to be hydrolyzed on the 5′ end. The resulting single-nucleotide repair gap contains a 3′OH and a 5′deoxyribose-phosphate (5′dRP) moiety at the margins. It has been suggested that this BER intermediate (a single-strand break with a 5′dRP moiety) recruits poly(ADP)ribose polymerase (PARP)1 to the lesion site. Recruitment then triggers activation of PARP1. Activated PARP1 polymerizes NAD+ to yield the polymer poly (ADP) ribose (PAR), an essential posttranslational modification. The first protein to be modified by PAR is PARP1 itself (auto-modification). Subsequently, it has been observed that XRCC1 and many other proteins are modified [40]. Once modified, activated PARP1 then facilitates chromatin relaxation (likely to provide access to the lesions for repair) [41, 42] and recruitment of the remaining BER proteins required to complete repair, including XRCC1, DNA Ligase III, and DNA polymerase β (Polβ). Whereas XRCC1 is a scaffold protein, Polβ carries out two essential enzymatic functions in BER. First, the repair gap is tailored by the 5′dRP lyase activity of Polβ. Next, Polβ fills the single-nucleotide gap, preparing the strand for ligation by either DNA ligase I (LigI) or a complex of DNA ligase III (LigIII), and XRCC1 [36].
Although some BER substrates (base lesions) induced by chemotherapeutic agents are cytotoxic [43], most are found to be mutagenic. However, essentially, every intermediate throughout the BER pathway (abasic sites, 5′dRP lesions, and single-strand DNA breaks) is toxic [13] and as such, there has been considerable interest in developing BER inhibitors to enhance the accumulation of the cytotoxic repair intermediates following chemotherapy or radiation treatment. This is discussed further in the sections below.
6.1.4 Nucleotide Excision Repair
Another multi-protein, highly complex DNA repair pathway is the NER pathway. NER plays an important role in the repair of DNA lesions induced by many genotoxins and chemotherapeutics including DNA cross-linking agents such as chloroethylating agents (see Sect. 6.1.2), cisplatin, carboplatin, and lesions induced by photodynamic therapy (PTD). Put simply, NER facilitates the removal of bulky DNA adducts that grossly distort the DNA double helix and those that cause a block to transcription. Molecular details on the proteins involved in NER can be found in several excellent reviews [44–47]. Overall, the pathway consists of two complementary sub-pathways that have some overlap. The two sub-pathways are distinct regarding the lesion recognition step but converge and utilize the same proteins to remove the oligonucleotide containing the lesion and for the steps involving new DNA synthesis.
The global genomic NER (GG-NER) pathway surveys the entire genome for DNA helix distorting lesions whereas the transcription-coupled repair NER (TC-NER) pathway is recruited to facilitate removal of DNA lesions that block the elongating RNA polymerase and stall transcription. The GG-NER pathway utilizes the DDB1/DDB2 heterodimer, part of the DDB1-Cul4A-DDB2 E3 ubiquitin ligase, to facilitate lesion recognition and repair [48]. As such, targeting the proteasome (Sect. 6.4.2) or deubiquitinating enzymes (DUBs) (Sect. 6.5.4) would therefore indirectly impact NER function. The TC-NER pathway partners with the TFIIH transcription complex to recognize and repair lesions that halt transcription. Upon lesion recognition, XPG mediates cleavage of the DNA strand containing the lesion on the 3′ side of the lesion, and subsequently, the ERCC1/XPF heterodimer hydrolyzes the DNA strand containing the lesion on the 5′ side. Replication factors then facilitate DNA synthesis and ligation. Of all the proteins in this pathway, ERCC1 has emerged as a valuable biomarker of response to chemotherapeutic agents that induce DNA damage repaired by NER (e.g., cisplatin) and is under consideration as a drug target [49, 50]. Currently, biomarker measurements have included both mRNA and protein analysis. However, it is not yet clear whether protein levels of ERCC1 are a valid biomarker [51, 52].
6.1.5 Non-Homologous-End-Joining
One of the most cytotoxic lesions is a DNA double-strand break (DSB). If not repaired, DSBs lead to chromosome breaks, loss of genetic material, and gross genomic rearrangements. Whereas tolerance to the presence of DSBs might vary in different cell types and cellular states, only a few DSBs will cause cell death or prevent clonogenicity in most cells including cancer cells [53]. These lesions are induced by multiple agents such as ionizing radiation, bleomycin, and topoisomerase II inhibitors, but can also be induced indirectly at replication forks when converting DNA single-strand breaks (SSBs) induced by topoisomerase I inhibitors (camptothecin).
Two major cellular pathways deal with the repair of DNA DSBs. The use of homologous DNA for repair distinguishes those repair pathways. As indicated by the name, the non-homologous-end-joining (NHEJ) repair pathway does not require any homologous sequences. Proteins of the NHEJ pathway can repair the two ends in a DSB by simple end joining while the homologous recombination (HR) repair pathway (see Sect. 6.1.6) requires homologous DNA stretches as templates for DNA synthesis and repair.
After the initial recognition of the DSB that is held in place and stabilized by the binding of the MRN complex (MRE11, RAD50, NBS) and promoted by ATM (see above), DSB repair is executed by the DNA-dependent protein kinase (DNA-PK). DNA-PK is comprised of the catalytic subunit DNA-PKcs and the relatively small Ku proteins (Ku70/80). They promote the simple ligation of the two broken DNA ends. Damage-induced DSBs, in particular after ionizing radiation, are rarely re-ligateable, and some end processing might be required that is accomplished by other enzymes such as Artemis. The DNA ligase IV-XRCC4 complex finally re-ligates the two ends. DNA PK–independent DSB end-joining activity has been observed in cells that have impaired NHEJ activity, the so-called alternative or B-NHEJ pathway. PARP and Ligase III activity appears to be implicated in this cellular DSB repair option [54, 55].
The role of ATM seems to be of particular importance in the repair of a certain proportion of DSBs, namely those in heterochromatic regions of the genome [56, 57]. These, judging from the repair kinetics, require more time to repair but influence survival substantially as indicated by the hypersensitivity to radiation of cells with impaired ATM function.
Based on the cytotoxic nature of DSBs, cells impaired in any step of the NHEJ process are highly sensitive to ionizing radiation. Genetic defects in or inhibition of NHEJ also profoundly affects survival of cells by other DNA-damaging agents that cause DSBs (directly or indirectly) such as DNA cross-linkers, bleomycin, and topoisomerase inhibitors.
6.1.6 Homologous Recombination DSB Repair and the Fanconi Pathway
The HR repair pathway, in contrast to NHEJ (see above), requires homologous DNA stretches as templates for DNA synthesis and repair [44, 58]. To assure accurate repair, HR tends to use the sister chromatid as a template, restricting this pathway to the S and G2 phase of the cell cycle. Together with the FA pathway, it has a crucial role in the surveillance of replication fork progression. As anticipated by its requirement for a homologous template, HR mainly determines survival of S- and G2-phase cells. HR’s involvement upon radiation is not only required at directly induced DSBs but also at secondarily induced DSBs that result from replication attempts on nicked DNA [59]. In agreement with such a function, HR has been shown to determine radiosensitivity in a cell cycle phase-dependent manner. HR and the FA pathway are also crucial in resolving blocked replication forks [60]. Such blocks are, for example, caused by interstrand cross-links (ICLs) that tether the two DNA strands together and prevent separation during replication. Survival upon other cancer therapeutic agents that induce replication-blocking lesions such as alkylating agents and topoisomerase inhibitors is strongly determined by the functionality of HR. These blocks must be repaired or bypassed to allow cells to survive.
A complex multi-member process assures HR-driven repair. In brief, an ordered assembly of nucleoprotein filaments of RAD52, RAD51, and RAD54 upon RPA coating of the resected DNA promotes and catalyzes homologous DNA pairing. The extent of the resection at the break site is mediated by the MRN complex and appears to partly define the use of HR instead of NHEJ. Strand exchange is assisted by the RAD51 paralogs RAD51B, RAD51C, RAD51D, XRCC2, and XRCC3. In concert, these proteins direct and provide the recombinase activity, that is, crucial to resolve the complex-branched structures that arise in this process. Notably, the products of the breast cancer susceptibility genes BRCA1 and BRCA2 are involved in the FA and HR repair pathways, assisting DSB and cross-link repair.
To allow the resolution of blocked replication fork structures, in particular following exposure to DNA cross-linking agents, another replication-associated repair process is required, the FA pathway. Its members were discovered while analyzing FA patients, victims of a human genetic disease that is characterized, among other features [61], by extreme cellular sensitivity to drugs that produce ICLs. Subsequently, their role and actions in cellular cross-link repair was revealed.
The products of at least 15 genes have been currently implicated in this pathway [61–63]. This pathway constitutes a major signaling cascade upon replication fork stalling: the FA “core complex” consists of at least 8 FA elements (FANCA, FANCB, FANCC, FANCE, FANCF, FANCG, FANCL, and FANCM) and acts by realizing the mono-ubiquitylation of the FA ID complex (FANCD2 and FANCI). This activation of the ID complex allows chromatin binding and is thought to facilitate DNA repair, in particular HR. The FA-mediated recruitment of the RAD51 recombinase and the BRCA1-FANCJ helicase activity allows re-establishment of the replication fork. Resolution of stalled replication forks appears to be also supported by the translocase activity of FANCM that can remodel branched DNA structures. The FA core complex is regulated by ATR and cell cycle checkpoint elements (CHK1) allowing the activation of the pathway [64]. Importantly, any mutation upstream of the FA pathway that will disrupt the mono-ubiquitylation of FANCD2 will result in the cellular ICL hypersensitivity phenotype.
As illustrated above, upon exposure to DNA-damaging agents, it is the multitude of cellular repair capacities that ultimately determines survival. HR and FA have been shown to determine the cellular sensitivity to a wide range of cancer therapeutics. HR-defective cells are hypersensitive to cross-linkers, IR, topoisomerase inhibitors, and alkylators as they induce DSBs and replication stalling.
6.2 Strategies Targeting DNA Repair for Cancer Treatment
The requirement for DNA repair and genome maintenance in response to radiation and genotoxic chemotherapeutics implicates DNA repair proteins as prime targets for improving response to currently available anticancer regimens. In addition, frequent cancer-specific DNA repair and DDR defects offer tumor-selective therapy options. Thus, strategies targeting DNA repair pathways represent promising new avenues to improve outcome in cancer treatment.
Targeting DNA repair pathways in cancer treatment has been proposed in several settings (Fig. 6.2). Most evidently, inhibition of cellular DNA repair will cause increased sensitivity to chemotherapeutic agents or radiotherapy [65]. As illustrated above, cellular death upon exposure to most chemotherapeutic agents and ionizing radiation partly depends on the repair capacity for the respective lesions. HR, NHEJ, BER, and DR processes are responsible for the resistance to these agents. Hence, inhibitors to DNA repair elements might be useful as dose intensifiers, augmenting the cell-killing properties of many if not all therapeutic agents. This can be particularly useful in a setting in which cancer cells obtained resistance to certain agents due to an improved repair capacity. Thus, counteracting marked cancer cell resistance is one strategy in which DNA repair inhibitors are proposed to act as dose intensifiers. By lowering the tolerance and inhibiting alternative repair routes while augmenting cell kill, the application of “intelligent” dose intensification by DNA repair inhibition can also prevent the development of chemotherapy resistance.
Strategies to improve cancer treatment by targeting DNA damage response and repair. The overall goal is to increase tumor cell kill (y-axis) while sparing normal tissue by increasing tumor specificity (x-axis) of the cancer treatment. While some strategies will achieve increased cell death and thereby an increased probability to control the tumor (for example by applying DNA-PK inhibitors in combination with radiotherapy or by re-sensitizing chemotherapy-resistant tumors), others (such as those based on the exploitation of tumor-specific defects) will increase tumor specificity of cancer therapies. However, in the manner in which current chemo- and radiotherapy regimens are largely applied, each of the targeted approaches will be only beneficial in a small fraction of patients with tumors that harbor the respective DNA repair defects. Nevertheless, considering the wide range of combination possibilities and the high number of targets and exploitation opportunities, together they represent a promising avenue in cancer treatment
Dose intensification will not be, in general, however, well tolerated, since chemotherapeutic drugs and radiotherapy doses are often administered at maximum-tolerated levels. Non-cancerous cells, with few exceptions, are as much exposed to the chemotherapeutic agents as are the cancer cells. Indeed, it is the normal tissue response that defines the dose level and use of dose intensifiers. Some tumor properties could, however, provide a tumor-specific effect when targeting DNA repair. Further, in line with the rationale of most classical cancer therapeutics, the proliferative nature of tumors is exploitable. For example, targeting replication-associated repair pathways such as HR with novel targeted agents could be beneficial in radiotherapy regimens in which the healthy cells in the irradiated area are nonproliferative. A gain could also be expected if chemotherapy dose limits are not defined by the proliferative cells, allowing dose intensification by targeting replication-associated DNA repair. Other tumor-specific properties such as exposure to hypoxic conditions or the altered metabolic status in cancer cells can offer opportunities to achieve tumor-specific dose intensification and will be discussed in the following paragraphs.
Another implication of DNA repair-targeting strategies has, however, become highly relevant. It is suggested that most cancer cells are likely to be defective in some aspect of DNA repair. Considering the multitude of repair options of healthy cells, DNA repair defects in cancer cells can be exploited by targeting the remaining repair processes. The combined lethal effect of two genetic variations that are otherwise non-lethal is termed “synthetic lethality” [66]. In compliance with the synthetic lethality concept, cancer cells defective in the primary repair pathway are viable but rely heavily on secondary backup repair for survival. As this is not only restricted to repair of endogenously produced lesions, this concept also applies to cells exposed to exogenous damage by exacerbating the effects of chemo- and radiotherapy in the defective cancer cells only (also may be called synthetic sickness). Despite mutations and genetic defects, the differential expression of DNA repair proteins or the altered engagement of DNA repair sub-pathways can be a base of tumor-specific activities. Hence, these tumor-specific DDR and repair defects offer promising novel approaches to tumor-selective therapy.
Investigation of the functionality of the individual DNA repair pathways in cancer cells and the knowledge on which pathways are implicated upon the inhibition of DNA repair drug targets are necessary to combine these cancer therapy options in an intelligent manner while focusing on a differential effect in the cancer versus normal cells.
6.3 DNA Repair Targets
The recognition that DNA repair processes are prime targets for chemo- and radiosensitization has driven the development of specific inhibitors to elements of DDR, NHEJ, HR, DR, and BER. The more recent discovery of tumor-specific targeting opportunities by the inhibition of DNA repair processes fueled such attempts and has yielded a multitude of DNA repair inhibitors [5]. Some of these novel agents are currently being evaluated in the clinic while others are being tested preclinically. Novel DNA repair targets have been identified, and compounds that specifically inhibit their activity are sought in order to apply tumor-specific anticancer strategies.
We will list currently explored DNA repair targets and some of the most advanced compounds according to their developmental stage (Table 6.1). Rationales and applied strategies will be discussed while pointing to opportunities on combinations and other DNA repair targets.
6.3.1 Poly(ADP)Ribose Polymerase
One of the most advanced and applied DNA repair target inhibitors to date are the PARP inhibitors [67–69]. Since the discovery that cells with defects in the BRCA genes are selectively killed by the inhibition of PARP, PARP inhibitors have rapidly made their way into the clinic [70, 71]. As tumors from carriers of mutations in the breast cancer susceptibility genes BRCA1 and BRCA2 are almost exclusively composed of such BRCA-defected cells while normal cells of these carriers still carry a functional allele, hence are HR proficient, these PARP inhibitors achieve tumor-specific kill with little normal cell toxicity. The proposed mechanism that causes such selectivity points to the dependence of BER-inhibited cells on HR due to secondarily induced cytotoxic DSBs (Fig. 6.3a) [60]. Indeed, early synthetic lethality screens in yeast indicated such an opportunity revealing a crucial link between BER and HR for cellular survival [72]. Other hypotheses assume a direct role of PARP inhibitors in replication fork stalling [73]. Most compounds with PARP inhibitory activity target PAR generation by blocking the catalytic activity of the enzyme. In principle, these compounds compete with NAD+ for the PARP catalytic site and are therefore not necessarily specific to PARP1 and could impact the activity of the other PARP isoforms [74]. Several PARP inhibitors are in clinical development. To date, the leading compounds Olaparib (AZD2281, AstraZeneca; originally developed by KuDos) and Veliparib (ABT-888, Abbott) are probably the two most extensively studied in the clinic whereas at least one compound, namely Iniparib (BSI-201; Sanofi-Aventis), has been reported to lack effective PARP inhibitory activity [75, 76].
Cellular functions of PARP and opportunities for cancer treatment. a The mode of action of PARP inhibitors is based primarily on inhibition of the poly(ADP)ribosylation activity of the enzyme PARP1. As a result, BER or SSBR cannot be executed. In addition, the lack of poly(ADP)ribosylation is likely to negatively impact chromatin and lesion accessibility. Auto-poly(ADP)ribosylation of PARP1 is thought to promote its repulsion from DNA. A failure to recruit downstream BER elements or an increase in lesion shielding by trapping PARP1 on the DNA further inhibits BER. BER intermediates accumulated upon chemo- and radiotherapy, however, will then cause replication problems that in turn will induce DSBs. Those will ultimately lead to cell death via apoptosis and/or mitotic catastrophe in particular if not repaired by HR. b PARP activity has several cellular roles that can be taken advantage of in cancer treatment. PARP promotes DNA repair and its inhibition, when combined with chemotherapy or with tumor-specific defects, enhances tumor cell kill. NAD + depletion, as a consequence of chemo- or radiotherapy-induced PARP activity, also induces cell kill. Agents depleting cellular NAD+ levels could indirectly inhibit PARP. Conversely, PARP inhibitors can have the effect of lowering chemotherapy-induced NAD + depletion, thereby altering the mode of cell death. Lastly PARP also has regulatory functions regarding DNA damage–induced gene expression that is often connected to an inflammatory or fibrotic response. PARP inhibitors have been reported to exhibit anti-inflammatory properties
Although registration of these drugs is still awaiting approval, several studies have shown their beneficial application [77, 78]. One obstacle could be that these PARP inhibitors were expected to act in a fraction of tumors, those exhibiting HR defects due to BRCA1 & BRCA2 mutations only. It should be noted that those early clinical trials revealed that not all BRCA mutation carriers benefit, indicating that a certain degree and type of HR defect is required to be exploitable with PARP inhibition. The impact of individual BRCA mutations with respect to HR functionality and/or PARP inhibitor sensitivity could be variable [79]. The status and propensity to use the remaining DSB repair mechanism NHEJ, for example via 53BP1 channeling, also influences the extent of PARP inhibitor toxicity [60, 80]. In addition, other general drug resistance mechanisms such as increased compound rejection by drug transporters, decreased tumor perfusion, cellular metabolism, or simply pharmacogenetics of a certain compound might also be responsible for a lack of benefit.
However, new studies indicate that other defects that supposedly result in impaired HR or DSB repair, such as when ATM is mutated, result in PARP inhibitor-mediated selective kill [81, 82]. Mantle cell lymphoma harbors ATM defects and preclinical studies demonstrate sensitivity of these tumors to PARP inhibition [83, 84]. Other genetic analyses indicated frequent ATM mutations in human cancer. These studies therefore warrant the application of PARP inhibitors for cancer treatment in a much larger patient population. In agreement with a synthetic lethal interaction with HR-driven processes, cells with an impaired FA pathway are hypersensitive to PARP inhibition, thus further enlarging the potential patient population that might benefit from such therapies [81]. Some studies indicated an impact of PTEN deletion on HR via a reduction of Rad51 or Rad51 paralogs, a condition that could be exploited by PARP inhibition [85, 86]. However, this could only be partly confirmed in a separate study [87]. HR impairment was observed when cells experience hypoxia [88, 89] which sensitized them to PARP inhibition. PARP inhibitor sensitivity in hypoxic cells or those with defects in FA, ATM, or PTEN is, however, generally less pronounced than when BRCA-defective. Yet, a benefit, in particular when combined with radio- or chemotherapy, can be anticipated since PARP inhibitors appear to enlarge the therapeutic window and could spare healthy cells from chemo- to radiosensitization.
Historically, prior to the discovery of specific killing of tumors with defective HR, PARP inhibitors have been actively studied in preclinical and clinical investigations to potentiate the cytotoxic effects of chemo- and radiotherapy (Fig. 6.3a). PARP inhibitors, due to their DNA repair-inhibiting properties, are radiation sensitizers and are highly effective in sensitizing cells to chemotherapeutic agents such as DNA alkylators (e.g., TMZ) and topoisomerase I inhibitors (irinotecan and topotecan). Combining PARP inhibitors with chemo-/radiotherapy could be beneficial in a large patient population with the added advantage of single-agent activity in a fraction of patients that happen to have tumors with exploitable DNA repair defects. As noted above, in such dose-intensification strategies, the benefit of tumor-specific killing needs to be evaluated against normal tissue effects that are connected to the use of DNA repair inhibitors. To exemplify, since the radiosensitizing effect of PARP inhibitors is most effective in proliferating cells, PARP inhibitors have been proposed in clinical radiotherapy settings in which normal tissue toxicity within the radiation field is not defined by a highly proliferating (stem) cell fraction such as in the treatment of lung cancer and glioblastoma [90, 91]. Radiation-induced lung toxicity is strongly determined by inflammatory and fibrotic processes. Therefore, PARP inhibition might impact lung toxicity in a beneficial way by altering the inflammatory DDR [92, 93].
Despite crude chemosensitization and synthetic lethal activity, other strategies exploit the BER inhibitory properties of PARP inhibitors. These strategies aim to prevent the quick development of resistance to alkylators such as temozolomide (TMZ). Resistance to TMZ is associated with MGMT expression and MMR defects (see above). PARP inhibitors, however, are able to (re-) sensitize MMR-defective cells to the anti-tumor effect of TMZ [94–98] thereby counteracting the resistance to TMZ.
Other PARP inhibitor combinations have been pursued. Assuming that BRCA defects do limit HR functions but do not fully impair HR, BER inhibition and consequently increased engagement of HR might be the basis for the observed synergistic cytotoxicity of Olaparib and Cisplatin [99, 100]. Depending on the cytotoxic agents and genetic background, BRCA-defected cells die by apoptosis or mitotic catastrophe upon PARP inhibition.
DNA damage–induced and PARP-activation-mediated consumption of NAD+ has been implicated in the increase in genotoxin-induced cell death [74, 101]. The PARP1 and PARP2 proteins [102] act as sensors of DNA damage such as DNA single-strand breaks and become hyperactivated, consuming NAD+ as a substrate to synthesize PAR [103]. Consumption of NAD+ after DNA damage leads to ATP depletion, likely due to continued re-synthesis of NAD+ as well as ongoing cellular utilization of NAD+ and ATP for metabolic functions [74, 101]. Following up on the observation that cell death due to BER inhibition and the accumulation of BER intermediates results in PARP hyperactivation [103], it was shown that the combination of BER and NAD+ biosynthesis inhibition significantly sensitizes glioma cells to TMZ. Dual targeting of these two interacting pathways (DNA repair and NAD+ biosynthesis) may prove to be an effective treatment combination for patients with resistant and recurrent GBM. Thus, in summary, several distinct roles of PARP including the promotion of DNA repair and its impact on damage-induced gene expression as well as the relationship to NAD+ levels can be exploited for cancer therapy (Fig. 6.3b).
6.3.2 O6-Methylguanine-DNA Methyltransferase
As described in Sect. 6.1 above, MGMT is the sole protein responsible for the repair of O6-alkylguanine lesions (formed by chemotherapeutic agents such as TMZ). Tumors with elevated MGMT expression are resistant to TMZ and related chemotherapeutic agents, and so, an active area of investigation has been the development of MGMT inhibitors. If MGMT is inhibited (or MGMT is not expressed), the tumor becomes highly sensitive to the agent (provided the tumor cell is proficient in MMR—see Sect. 6.1.2). There have been multiple methodologies proposed to inhibit or overcome resistance mediated by MGMT expression in the tumor as well as to prevent sensitivity of normal tissue (primarily hematopoietic cells) [12]. As we alluded in Sect. 6.1.2, the mechanism of action of MGMT in the repair or de-alkylation of guanine suggested O6-benzylguanine as an ideal inhibitor [104]. This inhibitor, also called BG, is an analogue of the O6-alkylguanine base and contains a benzyl ring instead of an alkyl group. The BG compound readily reacts with MGMT, and the benzyl moiety is transferred to Cys145 as shown (Fig. 6.4), releasing free guanine and rendering MGMT inactive. In some cases, this has been shown to trigger ubiquitylation and proteasome-mediated destruction of MGMT [105]. Further details on BG and related analogues to inhibit or regulate MGMT can be found elsewhere [106]. A second inhibitor with greater inhibitory activity that has been widely tested is the alkylguanine analogue 6-[4-bromo-2-thienyl]methoxypurin-2-amine [107], also called lomeguatrib [108, 109].
Schematic representation depicting the mechanism of action of the MGMT inhibitor O6-benzylguanine (BG). The benzyl moiety of BG is transferred to the Cys145 residue in MGMT, releasing free guanine, rendering MGMT inactive. In some cases, this has been shown to trigger ubiquitylation and proteasome-mediated destruction of MGMT
A significant challenge with all or most chemosensitizers is the observed increase in normal cell/tissue sensitivity or cell death. This has been addressed with regard to MGMT by the use of an ex vivo gene therapy approach to express a mutant of MGMT in the bone marrow or hematopoietic stem cells (cells are modified ex vivo and re-delivered to the patient), as described [12]. This is feasible since the G156A or P140K mutants of MGMT are 60-fold and 500-fold more resistant to BG than the wild-type protein, respectively [110].
6.3.3 Cell Cycle Checkpoints
DNA damage activates checkpoints to arrest proliferation. Normal cells have intact G1, S, and G2 checkpoints that are mediated by the ATM/CHK2 and ATR/CHK1 pathways. Owing to mutations in the p53 or pRB tumor suppressor genes, cancer cells, however, lack a G1 checkpoint and as a result, rely on G2 checkpoints to prevent cell division and propagation of the damage. Thus, the S/G2 checkpoint is an attractive target for cancer-specific sensitization to DNA-damaging agents [111]. In order to exploit the cancer-specific defects, inhibitors have been developed that abrogate the G2 checkpoint. CHK1 has been a prime target for such attempts, as activated CHK1 mediates the arrest by phosphorylating Cdc25A and Cdc25C leading to their degradation and inactivation that otherwise promote S-phase progression and entry into mitosis. Loss of the G2 cell cycle checkpoint, despite the presence of unrepaired damage, is thought to provoke mitotic catastrophe and ultimately cell kill. Consistent with this idea, loss of intra-S or G2/M checkpoints increases the cytotoxicity of DNA-damaging agents such as ionizing radiation and cisplatin. CHK1 knockdown sensitizes cells to 5-fluorouracil, doxorubicin, and etoposide. Despite the proposed mechanism that CHK1 inhibition causes sensitization in p53-defective cancer cells due to the G2 block abrogation in a G1 block-deficient background, the data to support this are contradictory [112–114]. Xeno-transplant studies on human triple-negative breast cancer demonstrated the benefit of combining irinotecan with CHK1 inhibitors, inducing checkpoint bypass and apoptosis. A role of p53 was supported by the gain of CHK1 sensitization after p53 knockdown in the resistant tumors [115]. A more complicated rationale, however, argues that a series of DDR defects in tumors should be considered and could be exploited by the inhibition of CHK1. These are based on the secondary effects of CHK1 inactivation on replication and are discussed below.
A large battery of CHK1 inhibitors is available to support cancer treatment in combination with radio- and chemotherapy [116]. Older CHK1 inhibitors such as UCN01 were not very selective but did demonstrate potent chemosensitization to cisplatin and camptothecin. More potent and specific CHK1 inhibitors were developed; however, concomitant CHK2 inhibition to some degree is common to most CHK1 inhibitors. In general, cells appear to depend on a functional G2 checkpoint when exposed to agents that cause replication stress. Hence, potentiation is greatest to cross-linkers, topoisomerase I poisons and nucleoside analogues such as gemcitabine. Consistent with replication stress hypersensitivity when G2 arrest is abrogated, PARP inhibitors also cause problems. PARP inhibition can cause a G2 checkpoint dependence and the combination of PARP inhibitors with CHK1 inhibitors is synthetic lethal [117]. These cellular sensitivity features are the basis for the clinical trials testing the combination of older (UCN01) or later generation compounds (for example AZD7762, AstraZeneca; PF477736, Pfizer; LY2606368, Eli Lilly and SCH900776, Schering Plough) with cisplatin, topotecan, and gemcitabine. For a more detailed review, see [116]. Unfortunately, safety requirements as assessed in these initial studies were not met in at least one compound (AZD7762) [118]. Similar to chemotherapy regimens, CHK1 inhibitors prevent ionizing radiation (IR) induced S and G2 arrest and demonstrated some potential to radiosensitize in a p53-dependent manner. CHK1 is upregulated in Myc-overexpressing lymphomas, and single-agent activity is expected since CHK1 inhibition is cytotoxic to these cells.
Interestingly, recent data indicate that CHK1 activity might prevent replication-induced DNA damage or is implicated in DNA repair [119, 120]. A conversion of halted replication forks (induced by the chemotherapeutics) into persistent and cytotoxic DSBs has been postulated. The CDK-driven unscheduled initiation of replication origins can be accounted for with regard to this DSB induction and explains chemosensitization beyond G2/M checkpoint abrogation [120]. The notion that CHK1 inhibition promotes replication stress merits their evaluation in synthetic lethal strategies exploiting tumor-specific DNA repair defects [121].
6.3.4 AP-Endonuclease 1
Targeted knockout (KO) of AP(apurinic/apyrimidinic)-endonuclease 1 (Ape1 or Ref1) in mice is lethal [122], and deletion of the Ape1 gene in mouse cells induces apoptosis within 24 h [123]. Interestingly, granzyme A(GzmA)-mediated cell death is enhanced by GzmA-mediated cleavage of Ape1. It is suggested that the proteolysis of APE1 enhances GzmA-mediated cell death by promoting apoptosis [124]. In human cells, RNA interference–mediated depletion of APE1 suppresses cell and tumor growth [125]. These and many other studies therefore support the development of APE1 inhibitors to enhance radiation and chemotherapy response [126, 127].
The first such compound to be tested clinically that impacts APE1 function in BER is methoxyamine (TRC102; Tracon Pharmaceuticals). Methoxyamine hydrochloride (MX) was first suggested to be a mutator, inducing the formation of 5,6-dihydro-6-methoxyaminecytosine residues in DNA [128]. It was subsequently determined that MX traps aldehyde groups, forming a stable intermediate [129]. The AP site in DNA is not a chemically unique species but exists as an equilibrium mixture of the ring-closed cyclic hemiacetals and open-chain aldehyde, and hydrate forms [130]. The transient open-chain aldehyde form is reactive with aldehyde-specific reagents such as methoxyamine, allowing the trapping or quantitative measurement of AP sites in DNA [131]. It was subsequently determined that the reaction between MX and the open-chain aldehyde form of an AP site blocks repair of DNA base lesions by BER. The trapped AP site is a highly cytotoxic intermediate and sensitizes cells to the cytotoxic effects of alkylating agents [132–134] and other DNA-damaging agents that may give rise to spontaneous or enzyme-mediated AP sites, such as TMZ [135], iododeoxyuridin + radiation [136], BCNU [137], manumycin A [138], fludarabine [139], radiation [140, 141], and pemetrexed [142]. Interestingly, elevated expression of the downstream BER protein DNA polymerase β (Polβ) reverses MX sensitization of alkylating agents, suggesting that the cleaved open-chain aldehyde may be the preferred MX substrate [134]. MX is in the clinic under the brand name TRC102 and is undergoing clinical evaluation in Phase II trials for solid tumors as a sensitizer to Temodar® (TMZ) or Alimta® (Pemetrexed) and in hematologic malignancies with Fludara® (Fludarabine).
In parallel, there are a number of groups developing direct APE1 active site inhibitors [143–146], and an overview of the development of APE1 inhibitors has just been reported [147]. As might be expected, many of these APE1 inhibitors are themselves cytotoxic and enhance the cytotoxicity of DNA-damaging agents such as TMZ [144, 145] or other alkylating agents [143]. Although Ape1 KO or GzmA-mediated Ape1 cleavage in mouse cells induces apoptosis, the cell death mechanism(s) induced by these recently developed APE1 inhibitors has yet to be resolved. Interestingly, the cell death mechanism triggered by some of these APE1 inhibitors may involve the accumulation of DNA DSBs since it was observed that the compounds are more cytotoxic in cells deficient in the HR proteins BRCA1 or BRCA2 [148].
6.3.5 Ataxia Telangiectasia-RAD3 Related
Similar strategies and rationales that apply to the CHK1 target also apply to ATR. A direct link connects ATR with CHK1 [121]. After initial Rad17 binding, Claspin-loaded single-strand DNA mediates the activation of CHK1 by ATR upon replication stress. ATR is essential in the surveillance of replication stress, especially when associated with exposed single-stranded DNA mostly connected to replication problems. Stalled replication forks can collapse which results in the formation of DSBs, a signal that mainly, but not exclusively, triggers ATM whereas the single-stranded DNA-induced damage response appears to be ATM independent. Endogenous and exogenous damage induces replication stress that requires a proficient ATR/CHK1 response for survival. One prevalent rationale to inhibit ATR for cancer therapy is to exacerbate the levels of replication stress that might be augmented in cancer cells due to inherited defects in the DDR and DNA repair pathways. Cancer cells are exposed to a higher load of replication stress compared to normal cells and will suffer most from targeting ATR. In addition, since targeting replication-associated processes, any selective killing property will be augmented in highly proliferative cells. Such a strategy is supported by two observations: (1) the activated DDR found in early stages of tumorigenesis [149, 150] indicates an increased load of “endogenous” DNA damage and (2) the discovery that a wide variety of oncogenes generate such damage.
Based on this proposed endogenous damage (replication stress)-induced killing mechanism, such ATR inhibitors have been proposed to act as single agents. However, combination strategies similar to those for the CHK1 inhibitor are envisioned. As noted before, the response of normal cells should be carefully taken into consideration. A few compounds have been pursued by industry. The ATR inhibitor NU6027 appears to impair HR and enhances the cytotoxicity of PARP inhibitors [151], a theme that is consistent to other DDR inhibitors. One compound (NVP-BEZ235) has been recently discovered and found to trigger preferential cell kill in cyclin-E overexpressing cells [152] and is awaiting entry into clinical trials for cancer therapy.
6.3.6 Ataxia Telangiectasia-Mutated
ATM inhibitors are less advanced in their clinical development than PARP or CHK1 inhibitors. Since involved in DSB repair, NHEJ, and HR, ATM inhibition results in reduced cellular DSB repair activity and cell cycle checkpoint defects. As a result, ATM inhibitors are highly potent radiosensitizers while exhibiting little toxicity on their own. Therefore, they have been proposed to be applied in this context [153]. Several ATM-inhibiting compounds have been identified and pursued in preclinical studies: KU55933 and KU60019 (KuDos/AstraZeneca) and CP466722 (Pfizer).
Combination strategies for ATM inhibitors have been proposed to enhance the cytotoxicity of PARP inhibitors (see above). Interestingly, p53 disruption in normal cells sensitized those cells to the combination of PARP and ATM inhibitors [83]. Due to its crucial role in DSB repair, the inhibition of ATM will radio- and chemosensitize most cells, including normal cells, with no evident DNA repair defects. However, synergistic cytotoxicity can be observed under certain conditions that could be exploited for tumor-targeted strategies. Cells with BER defects for example rely on secondary DSB repair pathways such as HR for survival upon damaging agents. Similar to the PARP inhibitor/HR-defect synthetic lethal interaction, the conversion of unrepaired SSBs to DSBs could be a mechanism that underlies such dependence. BER defects of different kinds have been reported in tumors, and it is suggested that inhibition of DSB repair processes will be beneficial in such a setting [154].
6.3.7 DNA-Dependent Protein Kinase
DNA-PK is crucial to DSB repair as well as cellular survival following radiation and topoisomerase II poisons, and so, DNA-PK has been a long sought target for drug development [155]. Kinase activities are relatively easy to target, and drugs inhibiting DNA-PK activity have been available for some time. One should note that a considerable fraction of first generation tyrosine kinase inhibitors do also inhibit DNA-PK to some extent. The radiosensitization phenotype by some of these agents should therefore be re-evaluated with respect to its origin (see below). Some examples of currently pursued inhibitors are NU7026 and NU7441 [156, 157]. Other targeting strategies apply short modified DNA molecules that are supposed to interfere in DNA-PK signaling (DT01, DNA therapeutics) or short peptides that resemble Ku80 (HNI-38) and disrupt the interaction with DNA-PK [158]. The dual DNA-PK and PI3K inhibitor KU-0060648 takes advantage of the inhibition of two cellular processes central to promoting survival upon chemo- and radiotherapeutic insults. Surprisingly, this inhibitor enhanced etoposide-induced xenograft tumor growth delay substantially without exacerbating etoposide toxicity in mice indicating that the combination provided some tumor specificity [159].
Most DNA-PK-inhibiting agents are very potent radiosensitizers, enhancing clonogenic cell kill markedly [160]. Radiosensitization by one compound, the DNA-PK inhibitor BEZ235, has been shown to be associated with an accelerated p53-dependent senescence [161]. Although proposed as radiation dose intensifiers, DNA-PK inhibitors, similar to ATM inhibitors (though often to a greater extent), will sensitize normal tissue to radiation at least as much as it does sensitize the tumors. However, some DNA repair defects inherent to tumors might provide some therapeutic gain when applying low inhibitor doses that could spare normal tissue. Opportunities exist for example in the treatment of ATM-deficient tumors since DNA-PK inhibition induces tumor cell kill in ATM-deficient cells [162].
Considering that disruption of the catalytic activity of DNA-PK confers severe immunodeficiency, DNA-PK inhibitors will have to be evaluated carefully for toxicities in preclinical studies before entering clinical trial [163]. Tumor-targeted delivery strategies could, however, make use of these highly potent radiation dose intensifiers. Other strategies exploiting the differential expression or use of DNA-PK and Ku-dependent DNA repair in some tissues with respect to the tumors could apply these inhibitors successfully. An example for this is the sensitization of EGFRvIII-overexpressing cells with DNA-PK inhibitors [164].
6.4 Indirect DNA Repair Modulators
6.4.1 Signaling Pathways, EGFR and the PI3K/AKT Pathway
Targeting the cellular signaling pathways via EGFR, PI3K/AKT, or MAPK can modulate the DNA repair status of cells. The effects are multiple and involve different pathways. Cellular signaling influences DNA repair in multiple ways including changes in the expression levels of crucial DNA repair proteins such as RAD51 or the regulation of the activation or translocation of enzymes and kinases such as DNA-PK. For example, the inhibition of the MAPK pathway can lead to reduced DSB repair by both homologous and non-homologous pathways [165, 166]. Links between the AKT, MAPK, and EGFR pathways and DNA repair have been found, particularly with DSB repair by NHEJ or HR [166–171]. Hence, targeting these pathways can result in the concomitant inhibition of DNA repair.
Based on knowledge that the PI3K pathway promotes DNA repair and survival, inhibitors of AKT have been evaluated as chemosensitizers for alkylating agents [172, 173]. AKT inhibitors such as LY294002 and wortmannin radiosensitized and caused enhanced sensitivity to alkylating agents such as TMZ, or the cross-linkers cisplatin in various human tumor cell lines.
The suppression of ATR/CHK1 checkpoints has been observed upon treatment with wortmannin (a fairly non-specific PI3K inhibitor). For more details, see [174] and references within. Note that an influence on Rad51 expression or on NHEJ via DNA-PK has also been observed.
Co-targeting PARP and the PI3K pathway has been shown to synergistically decrease growth and further induce apoptosis [175]. The molecular mechanism of this combination effect is not completely clear but could originate from secondary effects on DNA repair pathways and the DDR from the inhibition of cellular signaling in human cancer cells. Further detail on AKT inhibitors is discussed in more detail in Chap. 13.
6.4.2 Proteasome Inhibitors
Proteasome-mediated destruction and removal of key DNA repair and DDR proteins are an essential aspect of the cellular response to genotoxins [4, 176]. Failure to remove critical DNA repair and DDR proteins in a timely fashion effectively halts the process, triggering a cascade of events leading to cell death [177]. As the endpoint in ubiquitin-mediated protein turnover, targeting the proteasome is a high-profile target to enhance DNA damage–induced cell death [178, 179]. Further detail on the proteasome pathway and proteasome inhibitors is discussed in more detail in Chap. 12.
6.4.3 NAD+ Biosynthesis Inhibitors
The metabolite NAD+ is an essential substrate for all PARPs [74]. Since PARP1, as well as PARP2 and PARP3, is a critical protein involved in DNA repair and the DDR [36, 103], we have included NAD+ biosynthesis inhibitors as an indirect DNA repair modulator. All of the NAD+ biosynthesis inhibitors developed to date target NAMPT, a pivotal and rate-limiting enzyme in the salvage pathway of NAD+ biosynthesis [180]. NAMPT catalyzes the synthesis of nicotinamide mononucleotide (NMN) from nicotinamide to 5-phosphoribosyl-1-pyrophosphate [181]. The resulting NMN is then converted to NAD+ by one of the three isoforms of NMNAT (1, 2 or 3), located in the nucleus, cytosol, or mitochondria, respectively [182]. The most studied inhibitor of NAMPT is FK866 (also referred to as Apo866) although several other NAMPT inhibitors have been reported including GMX1778 [183], CB30865 [184], and CHS-828 [185], and one group is actively screening for NAMPT inhibitors [186]. FK866 binds at the interface of the NAMPT dimer [187] and effectively inhibits NAMPT and depletes cellular NAD+ levels within 24 h, inducing apoptosis [188] although the overall level of cell death may be offset by FK866-mediated induction of autophagy [189]. Further detail on the shift of cell death triggered by NAD+ biosynthesis inhibitors is discussed in Chap. 2.
As an NAMPT inhibitor, FK866 does have a chemo-potentiating effect [190–192] that is more pronounced when combined with a second DNA repair inhibitor [133] or when combined with TRAIL [193]. Interestingly, it has been suggested that some tumors are deficient in the NAPRT1-mediated NAD+ biosynthesis pathway that is responsible for generating NAD+ from nicotinic acid (NA) [183, 194]. In sum, these studies suggest that NAMPT is a potentially valuable target to impact both metabolism and DNA repair and may provide a type of synthetic lethality based on DNA repair and NAD+ biosynthesis status [133, 195].
6.5 Promising Future Targets and Opportunities
6.5.1 ERCC1
An emerging DNA repair target protein is ERCC1, encoded by the excision repair cross-complementing rodent repair deficiency, complementation group 1 gene (ERCC1) [49, 50]. As an essential NER protein, elevated ERCC1 expression enhances the repair of cisplatin-induced genomic DNA damage [15] and decreased expression is suggested to improve response to platinum-based therapies [196]. As described above (Sect. 6.1.4), ERCC1 exists as an obligate heterodimer with the protein XPF to yield an essential NER structure-specific nuclease ERCC1/XPF [15]. The ERCC1/XPF heterodimer is recruited to function in NER by interaction with the NER protein XPA [197] although multiple interaction sites are found to facilitate recruitment of ERCC1/XPF to the NER complex [198]. In this regard, it has been suggested that the function of ERCC1 in NER can be disrupted by interfering with the interaction between ERCC1 and XPA [199]. Alternatively, since heterodimer formation (ERCC1/XPF) is needed to maintain the stability of ERCC1 [200], it might be feasible to interrupt the heterodimer complex and force ERCC1 proteolysis [201]. Finally, one might also consider deregulating expression of ERCC1 through modulation of the RAS kinase or ERK1/2 pathways [202]. Using the Raf kinase and VEGF receptor inhibitor sorafenib reduces ERCC1 levels and effectively enhances the response to radiation and chemotherapy [203]. Of course this treatment would have multiple effects on gene regulation.
6.5.2 DNA Polymerases
There are 15 DNA polymerases in the human cell [204] and several have been suggested to be viable drug targets either because they facilitate repair of DNA damage or may be overexpressed in cancer [205]. DNA polymerase β (Polβ) is a member of the X family of DNA polymerases [204, 206] and is an essential BER protein, as detailed above. Polβ has two active sites and several critical functional domains that may be considered as targets to inhibit Polβ and BER. Further, the central and pivotal role of Polβ in BER implicates this protein as a prime target to enhance the response to chemotherapeutic agents or radiation. Although many inhibitors of Polβ have been developed and characterized very few, if any, show specificity and all are cytotoxic even in Polβ KO cells. Three recent reviews on Polβ inhibitors should be referred to for more detail [195, 207, 208].
Other polymerases have been also identified as potential targets. A siRNA screen and large-scale tumor sample analysis identified DNA polymerase theta (POLQ) as a target that determines radiosensitivity and is overexpressed in tumors. POLQ depletion radiosensitized these cells arguing for the development of POLQ inhibitors [209–211].
6.5.3 DNA Glycosylases
DNA glycosylases are the initiating enzymes in BER (see above). In general, DNA glycosylases probe the DNA helix for base lesions by a base-flipping mechanism, interrogating each base as it is flipped out of the major groove as the enzyme moves along the DNA helix. Identification of the lesion then promotes hydrolysis [212]. As discussed above, radiation and chemotherapeutic agents induce the formation of BER substrates and many of these base lesions block replication and are therefore cytotoxic. For example, the methylated base 3-methyladenine, induced by the chemotherapeutic agent TMZ and repaired by MPG, triggers lesion-induced apoptosis as well as sister chromatid exchange, chromatid and chromosome gaps and breaks, and S-phase arrest [213]. As such, it has been hypothesized that inhibition of MPG would enhance response to TMZ and other alkylators. Similarly, UNG, the primary glycosylase responsible for the removal of deoxyuracil, is reported to govern the efficacy of pemetrexed [142], and therefore, UNG might be considered a potential target to enhance the response to this chemotherapeutic agent. Finally, it was recently demonstrated that the DNA glycosylase OGG1 is in a complex with the BER protein PARP1. The OGG1–PARP1 complex promotes PARP1 activity whereas loss of OGG1 suppressed PARP1 activity [214]. From this study, it might be suggested that an inhibitor designed to disrupt the OGG1–PARP1 interaction may function as a pseudo- or indirect PARP1 inhibitor. However, there are multiple challenges to developing effective DNA glycosylase inhibitors such as the large overlap in enzyme substrate specificity [36, 37] and the requirement for some glycosylases in normal cell survival [215, 216] and immunoglobulin class-switch recombination [217]. To date, we know of no DNA glycosylase inhibitors with the exception of Ugi, a virally encoded UNG inhibitor [218].
6.5.4 DUBs
As with most posttranslational modifications, both the synthesis and removal of the ubiquitin modification is a highly regulated process [219, 220]. Recently, several studies have revealed the potential for targeting DUBs to enhance radiation and chemotherapeutic response [221, 222]. Toward that end, several groups have reported the development of DUB inhibitors that are effective in preclinical studies [223–226].
6.5.5 HR/FA
HR and FA strongly determine cellular survival upon cross-linkers, topoisomerase poisons, alkylators, and radiation. These pathways are mostly engaged in a proliferation-dependent manner with less influence on damage repair and response in quiescent cells. Thus, targeting HR and FA can be an effective means to potentiate radio- and chemotherapy [227].
Novel agents targeting RAD51, albeit indirectly, are now emerging in clinical trials. MP470 (Supergen) is a tyrosine kinase inhibitor with RAD51 protein–suppressing properties. More specific inhibitors are sought in high-throughput screening efforts of which some provided some compounds with convincing specificity and potency [228].
The value of HR inhibition can be deduced from experiments in which elements of the HR or FA pathway such as RAD51 or BRCA2 have been downregulated, supporting the potential of HR/FA-targeting strategies. Downregulation of Rad51 or BRCA2 sensitized glioma cells to the cytotoxic effects of TMZ or nimustine in cells with low MGMT levels [229]. The marked resistance of gliomas to chemo- or radiotherapy regimens that might be partly defined by HR or increased Rad51 levels can be defined by targeting RAD51, indicating a similar strategy option to counteract resistance by HR inhibition [230].
FA- and HR-defected cells have been found to lack the increased radio-resistance that is observed in hypoxic areas of the tumors [231]. Inhibitors targeted to HR or FA will not only preferentially sensitize proliferating cancer cells but also hypoxic tumors to radiation. Owing to less-effective DSB induction under low oxygen concentration, radiotherapy often fails in patients with hypoxic tumors. Tumor specificity of HR-targeting drugs will be increased when applied to BER-defected cancer cells as illustrated above. BRCA2-knockdown radiosensitizes cells that express a dominant-negative Polβ mutant to a greater extent than isogenic controls [232]. Similarly, NHEJ-defected cancer cells are expected to rely on HR for survival upon genotoxic threats. Together, these interactions and data support strategies applying HR- or FA-inhibiting drugs for tumor-specific sensitization.
6.5.6 Poly(ADP)ribose Glycohydrolase
Poly(ADP)ribose glycohydrolase (PARG) [233] is an enzyme involved in BER and is the major factor responsible for removing the PAR molecules synthesized by PARP1 and PARP2, among other PARPs [234]. As seen with PARP1, PARG is also a substrate for caspase-3 [235], supporting the significance of PAR metabolism in cell death processes. Mouse KO studies or RNA interference studies to knockdown PARG in human cells have suggested that blocking PAR degradation is an effective means to modulate chemotherapy and radiation response [134, 236, 237]. Pre-clinical evaluations using early phase small-molecule inhibitors such as gallotannin, tannic acid, and related small molecules have implicated PARG in genotoxin or chemotherapy sensitivity [238–240]. The inhibitor GPI-16552 (N-bis-(3-phenyl-propyl)9-oxo-fluorene-2,7-diamide) has shown early success in cell and mouse models as a chemotherapy sensitizer [97] but has mostly been effective to reduce inflammation [241–245]. Recent drug discovery efforts seem promising and have yielded PARG inhibitors with increased specificity and cell permeability [246, 247].
6.6 Summary and Concluding Remarks
Inducing tumor-specific or selective cell death is a significant challenge in the treatment of cancer. Although radiation and most chemotherapeutic treatments are designed to induce significant genotoxic damage to the tumor cells, a multitude of DNA repair and DDR gene products respond to the presence of genotoxic stress (DNA damage) by orchestrating a massively complex cellular response to survive and repair the genome insult. Most DNA repair pathways focus on specific damage types (e.g., DSBs), but there is considerable overlap and backup capacity when evaluating the overall involvement of DR, BER, MMR, NER, HR, and NHEJ proteins in response to these agents (radiation and chemotherapy). It has been observed that some cancer cells can survive the loss of key DNA repair proteins (e.g., BRCA1/2). In fact, we have come to realize that many cancer cell types are defective in one or more DNA repair or DDR genes/proteins, precipitating the need to identify the key stress nodes in these defective cells in response to radiation and classical chemotherapy treatments. With this in mind, we have described the molecular pathways that participate in the repair of DNA damage induced by radiation and chemotherapeutics. Further, we present novel therapeutic strategies being considered to exploit inherent weaknesses in tumor cells such as defects in one or more DNA repair pathways that may provide the opportunity to selectively increase tumor-specific cell death.
References
Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362(6422):709–715
Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461(7267):1071–1078. Epub 2009/10/23
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. Epub 2011/03/08
Harper JW, Elledge SJ (2007) The DNA damage response: ten years after. Mol Cell 28(5):739–745. Epub 2007/12/18
Ljungman M (2009) Targeting the DNA damage response in cancer. Chem Rev 109(7):2929–2950. Epub 2009/06/24
Alberts B (2009) Redefining cancer research. Science 325(5946):1319. Epub 2009/09/12
Helleday T, Petermann E, Lundin C, Hodgson B, Sharma RA (2008) DNA repair pathways as targets for cancer therapy. Nat Rev Cancer 8(3):193–204
Lavin MF (2008) Ataxia-telangiectasia: from a rare disorder to a paradigm for cell signalling and cancer. Nat Rev Mol Cell Biol 9(10):759–769. Epub 2008/09/25
Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER 3rd, Hurov KE, Luo J et al (2007) ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316(5828):1160–1166
Bakkenist CJ, Kastan MB (2003) DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421(6922):499–506
Wood RD, Mitchell M, Sgouros J, Lindahl T (2001) Human DNA repair genes. Science 291(5507):1284–1289
Gerson SL (2004) MGMT: its role in cancer aetiology and cancer therapeutics. Nat Rev Cancer 4(4):296–307
Fu D, Calvo JA, Samson LD (2012) Balancing repair and tolerance of DNA damage caused by alkylating agents. Nat Rev Cancer 12(2):104–120. Epub 2012/01/13
Kaina B, Christmann M, Naumann S, Roos WP (2007) MGMT: key node in the battle against genotoxicity, carcinogenicity and apoptosis induced by alkylating agents. DNA Repair (Amst) 6(8):1079–1099
Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, Ellenberger T (2006) DNA repair and mutagenesis. 2nd Edn. ASM Press, Washington, p 1164
Wyatt MD, Pittman DL (2006) Methylating agents and DNA repair responses: methylated bases and sources of strand breaks. Chem Res Toxicol 19(12):1580–1594
Wang JY, Edelmann W (2006) Mismatch repair proteins as sensors of alkylation DNA damage. Cancer Cell 9(6):417–418
Friedman HS, Johnson SP, Dong Q, Schold SC, Rasheed BK, Bigner SH et al (1997) Methylator resistance mediated by mismatch repair deficiency in a glioblastoma multiforme xenograft. Cancer Res 57(14):2933–2936
Roos WP, Batista LF, Naumann SC, Wick W, Weller M, Menck CF et al (2007) Apoptosis in malignant glioma cells triggered by the temozolomide-induced DNA lesion O6-methylguanine. Oncogene 26(2):186–197
Caporali S, Falcinelli S, Starace G, Russo MT, Bonmassar E, Jiricny J et al (2004) DNA damage induced by temozolomide signals to both ATM and ATR: role of the mismatch repair system. Mol Pharmacol 66(3):478–491
Jiricny J (2006) The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 7(5):335–346
Pabla N, Ma Z, McIlhatton MA, Fishel R, Dong Z (2011) hMSH2 recruits ATR to DNA damage sites for activation during DNA damage-induced apoptosis. J Biol Chem 286(12):10411–10418. Epub 2011/02/03
Wang Y, Qin J. MSH2 and ATR form a signaling module and regulate two branches of the damage response to DNA methylation. Proceedings Nat Acad Sci 100(26):15387–15392
Hegi ME, Liu L, Herman JG, Stupp R, Wick W, Weller M et al (2008) Correlation of O6-methylguanine methyltransferase (MGMT) promoter methylation with clinical outcomes in glioblastoma and clinical strategies to modulate MGMT activity. J Clin Oncol 26(25):4189–4199
Sarkaria JN, Kitange GJ, James CD, Plummer R, Calvert H, Weller M et al (2008) Mechanisms of chemoresistance to alkylating agents in malignant glioma. Clin Cancer Res 14(10):2900–2908
Pollack IF, Hamilton RL, Sobol RW, Burnham J, Yates AJ, Holmes EJ et al (2006) MGMT expression strongly correlates with outcome in childhood malignant gliomas: results from the CCG-945 cohort. J Clin Oncol 24(21):3431–3437
Holland EC (2000) Glioblastoma multiforme: the terminator. Proc Natl Acad Sci USA 97(12):6242–6244. Epub 2000/06/07
Cohen MH, Johnson JR, Pazdur R (2005) Food and drug administration drug approval summary: temozolomide plus radiation therapy for the treatment of newly diagnosed glioblastoma multiforme. Clin Cancer Res 11(19 Pt 1):6767–6771
Stupp R, Hegi ME, Mason WP, van den Bent MJ, Taphoorn MJ, Janzer RC et al (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5 year analysis of the EORTC-NCIC trial. Lancet Oncol 10(5):459–466
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996
Cahill DP, Levine KK, Betensky RA, Codd PJ, Romany CA, Reavie LB et al (2007) Loss of the mismatch repair protein MSH6 in human glioblastomas is associated with tumor progression during temozolomide treatment. Clin Cancer Res 13(7):2038–2045
Yip S, Miao J, Cahill DP, Iafrate AJ, Aldape K, Nutt CL et al (2009) MSH6 mutations arise in glioblastomas during temozolomide therapy and mediate temozolomide resistance. Clin Cancer Res 15(14):4622–4629
Friedman HS, Keir S, Pegg AE, Houghton PJ, Colvin OM, Moschel RC et al (2002) O6-benzylguanine-mediated enhancement of chemotherapy. Mol Cancer Ther 1(11):943–948
Tserng KY, Ingalls ST, Boczko EM, Spiro TP, Li X, Majka S et al (2003) Pharmacokinetics of O6-benzylguanine (NSC637037) and its metabolite, 8-oxo-O6-benzylguanine. J Clin Pharmacol 43(8):881–893
Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V et al (2000) Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med 343(19):1350–1354
Almeida KH, Sobol RW (2007) A unified view of base excision repair: lesion-dependent protein complexes regulated by post-translational modification. DNA Repair 6(6):695–711
Svilar D, Goellner EM, Almeida KH, Sobol RW (2011) Base excision repair and lesion-dependent sub-pathways for repair of oxidative DNA damage. Antioxid Redox Signal 14(12):2491–2507. Epub 2010/07/24
Sobol RW (2009) Temozolomide. In: Schwab M (ed) Encyclopedia of cancer, 2nd edn. Springer, Berlin Heidelberg, pp 2928–2933
Wilson SH, Kunkel TA (2000) Passing the baton in base excision repair. Nat Struct Biol 7(3):176–178
Gagne JP, Pic E, Isabelle M, Krietsch J, Ethier C, Paquet E et al (2012) Quantitative proteomics profiling of the poly(ADP-ribose)-related response to genotoxic stress. Nucleic Acids Res 40(16):7788–7805. Epub 2012/06/07
Gibson BA, Kraus WL (2012) New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol 13(7):411–424. Epub 2012/06/21
Sousa FG, Matuo R, Soares DG, Escargueil AE, Henriques JA, Larsen AK et al PARPs and the DNA damage response. Carcinogenesis 33(8):1433–1440. Epub 2012/03/21
Roos WP, Kaina B (2012) DNA damage-induced apoptosis: from specific DNA lesions to the DNA damage response and apoptosis. Cancer Lett In Press. Epub 2012/01/21
Hoeijmakers JH (2001) Genome maintenance mechanisms for preventing cancer. Nature 411(6835):366–374
De Laat WL, Jaspers NG, Hoeijmakers JH (1999) Molecular mechanism of nucleotide excision repair. Genes Dev 13(7):768–785. Epub 1999/04/10
Wood RD (1996) DNA repair in eukaryotes. Annu Rev Biochem 65:135–167. Epub 1996/01/01
Shuck SC, Short EA, Turchi JJ (2008) Eukaryotic nucleotide excision repair: from understanding mechanisms to influencing biology. Cell Res 18(1):64–72. Epub 2008/01/02
Kapetanaki MG, Guerrero-Santoro J, Bisi DC, Hsieh CL, Rapic-Otrin V, Levine AS (2006) The DDB1-CUL4ADDB2 ubiquitin ligase is deficient in xeroderma pigmentosum group E and targets histone H2A at UV-damaged DNA sites. Proc Natl Acad Sci USA 103(8):2588–2593
Barakat K, Gajewski M, Tuszynski JA (2012) DNA repair inhibitors: the next major step to improve cancer therapy. Curr Top Med Chem 12(12):1376–1390. Epub 2012/07/17
Postel-Vinay S, Vanhecke E, Olaussen KA, Lord CJ, Ashworth A, Soria JC (2012) The potential of exploiting DNA-repair defects for optimizing lung cancer treatment. Nat Rev Clin Oncol 9(3):144–155. Epub 2012/02/15
Bhagwat NR, Roginskaya VY, Acquafondata MB, Dhir R, Wood RD, Niedernhofer LJ (2009) Immunodetection of DNA repair endonuclease ERCC1-XPF in human tissue. Cancer Res 69(17):6831–6838. Epub 2009/09/03
Niedernhofer LJ, Bhagwat N, Wood RD. ERCC1 and non-small-cell lung cancer. N Engl J Med. 2007;356(24):2538-40; author reply 40-1. Epub 2007/06/15
Lobrich M, Shibata A, Beucher A, Fisher A, Ensminger M, Goodarzi AA et al (2010) GammaH2AX foci analysis for monitoring DNA double-strand break repair: strengths, limitations and optimization. Cell Cycle 9(4):662–669. Epub 2010/02/09
Audebert M, Salles B, Calsou P (2004) Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J Biol Chem 279(53):55117–55126
Mladenov E, Iliakis G (2011) Induction and repair of DNA double strand breaks: the increasing spectrum of non-homologous end joining pathways. Mutat Res 711(1–2):61–72. Epub 2011/02/19
Goodarzi AA, Noon AT, Deckbar D, Ziv Y, Shiloh Y, Lobrich M et al (2008) ATM signaling facilitates repair of DNA double-strand breaks associated with heterochromatin. Mol Cell 31(2):167–177. Epub 2008/07/29
Murray JM, Stiff T, Jeggo PA (2012) DNA double-strand break repair within heterochromatic regions. Biochem Soc Trans 40(1):173–178. Epub 2012/01/21
Moynahan ME, Jasin M (2010) Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat Rev Mol Cell Biol 11(3):196–207. Epub 2010/02/24
Jeggo PA, Geuting V, Lobrich M (2011) The role of homologous recombination in radiation-induced double-strand break repair. Radiother Oncol 101(1):7–12. Epub 2011/07/09
Allen C, Ashley AK, Hromas R, Nickoloff JA (2011) More forks on the road to replication stress recovery. J Mol Cell Biol 3(1):4–12. Epub 2011/02/01
Kim H, D’Andrea AD (2012) Regulation of DNA cross-link repair by the Fanconi anemia/BRCA pathway. Genes Dev 26(13):1393–1408. Epub 2012/07/04
Crossan GP, Patel KJ (2012) The fanconi anaemia pathway orchestrates incisions at sites of crosslinked DNA. J Pathol 226(2):326–337. Epub 2011/10/01
Yuan F, Song L, Qian L, Hu JJ, Zhang Y (2010) Assembling an orchestra: Fanconi anemia pathway of DNA repair. Front Biosci 15:1131–1149. Epub 2010/06/03
Wang W (2008) A major switch for the Fanconi anemia DNA damage-response pathway. Nat Struct Mol Biol 15(11):1128–1130. Epub 2008/11/06
Begg AC, Stewart FA, Vens C (2011) Strategies to improve radiotherapy with targeted drugs. Nat Rev Cancer 11(4):239–253. Epub 2011/03/25
Kaelin WG Jr (2005) The concept of synthetic lethality in the context of anticancer therapy. Nat Rev Cancer 5(9):689–698
Javle M, Curtin NJ (2011) The role of PARP in DNA repair and its therapeutic exploitation. Br J Cancer 105(8):1114–1122. Epub 2011/10/13
Sandhu SK, Yap TA, De Bono JS (2010) Poly(ADP-ribose) polymerase inhibitors in cancer treatment: a clinical perspective. Eur J Cancer 46(1):9–20. Epub 2009/11/21
Peralta-Leal A, Rodriguez MI, Oliver FJ (2008) Poly(ADP-ribose)polymerase-1 (PARP-1) in carcinogenesis: potential role of PARP inhibitors in cancer treatment. Clin Transl Oncol 10(6):318–323
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E et al (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434(7035):913–917
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434(7035):917–921
Hendricks CA, Razlog M, Matsuguchi T, Goyal A, Brock AL, Engelward BP (2002) The S.cerevisiae Mag1 3-methyladenine DNA glycosylase modulates susceptibility to homologous recombination. DNA Repair 1:645–659
Shaheen M, Allen C, Nickoloff JA, Hromas R (2011) Synthetic lethality: exploiting the addiction of cancer to DNA repair. Blood 117(23):6074–6082. Epub 2011/03/29
Hassa PO, Haenni SS, Elser M, Hottiger MO (2006) Nuclear ADP-ribosylation reactions in mammalian cells: where are we today and where are we going? Microbiol Mol Biol Rev 70(3):789–829
Patel AG, De Lorenzo SB, Flatten KS, Poirier GG, Kaufmann SH (2012) Failure of iniparib to inhibit poly(ADP-Ribose) polymerase in vitro. Clin Cancer Res 18(6):1655–1662. Epub 2012/02/01
Liu X, Shi Y, Maag DX, Palma JP, Patterson MJ, Ellis PA et al (2012) Iniparib nonselectively modifies cysteine-containing proteins in tumor cells and is not a bona fide PARP inhibitor. Clin Cancer Res 18(2):510–523. Epub 2011/12/01
Audeh MW, Carmichael J, Penson RT, Friedlander M, Powell B, Bell-McGuinn KM et al (2010) Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: a proof-of-concept trial. Lancet 376(9737):245–251. Epub 2010/07/09
Tutt A, Robson M, Garber JE, Domchek SM, Audeh MW, Weitzel JN et al (2010) Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet 376(9737):235–244. Epub 2010/07/09
Vollebergh MA, Jonkers J, Linn SC (2012) Genomic instability in breast and ovarian cancers: translation into clinical predictive biomarkers. Cell Mol Life Sci 69(2):223–245. Epub 2011/09/17
Bunting SF, Callen E, Kozak ML, Kim JM, Wong N, Lopez-Contreras AJ et al (2012) BRCA1 functions independently of homologous recombination in DNA interstrand crosslink repair. Mol Cell 46(2):125–135. Epub 2012/03/27
McCabe N, Turner NC, Lord CJ, Kluzek K, Bialkowska A, Swift S et al (2006) Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-Ribose) polymerase inhibition. Cancer Res 66(16):8109–8115
Yap TA, Sandhu SK, Carden CP, De Bono JS (2011) Poly(ADP-ribose) polymerase (PARP) inhibitors: exploiting a synthetic lethal strategy in the clinic. CA Cancer J Clin 61(1):31–49. Epub 2011/01/06
Williamson CT, Kubota E, Hamill JD, Klimowicz A, Ye R, Muzik H et al (2012) Enhanced cytotoxicity of PARP inhibition in mantle cell lymphoma harbouring mutations in both ATM and p53. EMBO Mol Med 4(6):515–527. Epub 2012/03/15
Williamson CT, Muzik H, Turhan AG, Zamo A, O’Connor MJ, Bebb DG et al ATM deficiency sensitizes mantle cell lymphoma cells to poly(ADP-ribose) polymerase-1 inhibitors. Mol Cancer Ther 9(2):347–57. Epub 2010/02/04
McEllin B, Camacho CV, Mukherjee B, Hahm B, Tomimatsu N, Bachoo RM et al (2010) PTEN loss compromises homologous recombination repair in astrocytes: implications for glioblastoma therapy with temozolomide or poly(ADP-ribose) polymerase inhibitors. Cancer Res 70(13):5457–5464. Epub 2010/06/10
Mendes-Pereira AM, Martin SA, Brough R, McCarthy A, Taylor JR, Kim JS et al (2009) Synthetic lethal targeting of PTEN mutant cells with PARP inhibitors. EMBO molecular medicine 1(6–7):315–322. Epub 2010/01/06
Fraser M, Zhao H, Luoto KR, Lundin C, Coackley C, Chan N et al (2011) PTEN deletion in prostate cancer cells does not associate with loss of RAD51 function: implications for radiotherapy and chemotherapy. Clin Cancer Res 18(4):1015–1027. Epub 2011/11/25
Chan N, Pires IM, Bencokova Z, Coackley C, Luoto KR, Bhogal N et al (2010) Contextual synthetic lethality of cancer cell kill based on the tumor microenvironment. Cancer Res 70(20):8045–8054. Epub 2010/10/07
Liu SK, Coackley C, Krause M, Jalali F, Chan N, Bristow RG (2008) A novel poly(ADP-ribose) polymerase inhibitor, ABT-888, radiosensitizes malignant human cell lines under hypoxia. Radiother Oncol 88(2):258–268. Epub 2008/05/06
Chalmers AJ (2010) Overcoming resistance of glioblastoma to conventional cytotoxic therapies by the addition of PARP inhibitors. Anti-Cancer Agents Med Chem 10(7):520–533. Epub 2010/10/01
Chalmers AJ, Lakshman M, Chan N, Bristow RG (2010) Poly(ADP-ribose) polymerase inhibition as a model for synthetic lethality in developing radiation oncology targets. Semin Radiat Oncol 20(4):274–281. Epub 2010/09/14
Aguilar-Quesada R, Munoz-Gamez JA, Martin-Oliva D, Peralta-Leal A, Quiles-Perez R, Rodriguez-Vargas JM et al (2007) Modulation of transcription by PARP-1: consequences in carcinogenesis and inflammation. Curr Med Chem 14(11):1179–1187
Giansanti V, Dona F, Tillhon M, Scovassi AI (2010) PARP inhibitors: new tools to protect from inflammation. Biochem pharmacol 80(12):1869–1877. Epub 2010/04/27
De la Lastra CA, Villegas I, Sanchez-Fidalgo S (2007) Poly(ADP-ribose) polymerase inhibitors: new pharmacological functions and potential clinical implications. Curr Pharm Des 13(9):933–962. Epub 2007/04/14
Wedge SR, Porteous JK, Newlands ES (1996) 3-Aminobenzamide and/or O6-benzylguanine evaluated as an adjuvant to temozolomide or BCNU treatment in cell lines of variable mismatch repair status and O6-alkylguanine-DNA alkyltransferase activity. Br J Cancer 74(7):1030–1036. Epub 1996/10/01
Tentori L, Turriziani M, Franco D, Serafino A, Levati L, Roy R et al (1999) Treatment with temozolomide and poly(ADP-ribose) polymerase inhibitors induces early apoptosis and increases base excision repair gene transcripts in leukemic cells resistant to triazene compounds. Leukemia 13(6):901–909
Tentori L, Leonetti C, Scarsella M, D’Amati G, Vergati M, Portarena I et al (2003) Systemic administration of GPI 15427, a novel poly(ADP-ribose) polymerase-1 inhibitor, increases the antitumor activity of temozolomide against intracranial melanoma, glioma, lymphoma. Clin Cancer Res 9(14):5370–5379
Curtin NJ, Wang LZ, Yiakouvaki A, Kyle S, Arris CA, Canan-Koch S et al (2004) Novel poly(ADP-ribose) polymerase-1 inhibitor, AG14361, restores sensitivity to temozolomide in mismatch repair-deficient cells. Clin Cancer Res 10(3):881–889
Rottenberg S, Jaspers JE, Kersbergen A, van der Burg E, Nygren AO, Zander SA et al (2008) High sensitivity of BRCA1-deficient mammary tumors to the PARP inhibitor AZD2281 alone and in combination with platinum drugs. Proc Natl Acad Sci USA 105(44):17079–17084. Epub 2008/10/31
Evers B, Drost R, Schut E, de Bruin M, van der Burg E, Derksen PW et al (2008) Selective inhibition of BRCA2-deficient mammary tumor cell growth by AZD2281 and cisplatin. Clin Cancer Res 14(12):3916–3925 Epub 2008/06/19
Berger NA (1985) Poly(ADP-ribose) in the cellular response to DNA damage. Radiat Res 101(1):4–15
Hottiger MO, Hassa PO, Luscher B, Schuler H, Koch-Nolte F (2010) Toward a unified nomenclature for mammalian ADP-ribosyltransferases. Trends Biochem Sci 35(4):208–219. Epub 2010/01/29
Tang J, Goellner EM, Wang XW, Trivedi RN, St. Croix CM, Jelezcova E et al (2010) Bioenergetic metabolites regulate base excision repair-dependent cell death in response to DNA damage. Mol Cancer Res 8(1):67–79. Epub 2010/01/14
Dolan ME, Pegg AE (1997) O6-Benzylguanine and its role in chemotherapy. Clin Cancer Res 3(6):837–847. Epub 1997/06/01
Xu-Welliver M, Pegg AE (2002) Degradation of the alkylated form of the DNA repair protein, O(6)-alkylguanine-DNA alkyltransferase. Carcinogenesis 23(5):823–30. Epub 2002/05/23
Pegg AE (2011) Multifaceted roles of alkyltransferase and related proteins in DNA repair, DNA damage, resistance to chemotherapy, and research tools. Chem Res Toxicol 24(5):618–639. Epub 2011/04/07
Middleton MR, Kelly J, Thatcher N, Donnelly DJ, McElhinney RS, McMurry TB et al (2000) O(6)-(4-bromothenyl)guanine improves the therapeutic index of temozolomide against A375 M melanoma xenografts. Int J Cancer 85(2):248–252. Epub 2000/01/11
Zhang J, Stevens MF, Bradshaw TD (2012) Temozolomide: mechanisms of action, repair and resistance. Curr Mol Pharmacol 5(1):102–114. Epub 2011/11/30
Khan O, Middleton MR (2007) The therapeutic potential of O6-alkylguanine DNA alkyltransferase inhibitors. Expert Opin Investig Drugs 16(10):1573–1584. Epub 2007/10/10
Davis BM, Roth JC, Liu L, Xu-Welliver M, Pegg AE, Gerson SL (1999) Characterization of the P140 K, PVP(138-140)MLK, and G156A O6-methylguanine-DNA methyltransferase mutants: implications for drug resistance gene therapy. Hum Gene Ther 10(17):2769–2778. Epub 1999/12/10
Chen T, Stephens PA, Middleton FK, Curtin NJ (2011) Targeting the S and G2 checkpoint to treat cancer. Drug Discovery Today 17(5–6):194–202. Epub 2011/12/24
Zenvirt S, Kravchenko-Balasha N, Levitzki A (2010) Status of p53 in human cancer cells does not predict efficacy of CHK1 kinase inhibitors combined with chemotherapeutic agents. Oncogene 29(46):6149–6159. Epub 2010/08/24
Cho SH, Toouli CD, Fujii GH, Crain C, Parry D (2004) Chk1 is essential for tumor cell viability following activation of the replication checkpoint. Cell Cycle 4(1):131–139. Epub 2004/11/13
Sorensen CS, Hansen LT, Dziegielewski J, Syljuasen RG, Lundin C, Bartek J et al (2005) The cell-cycle checkpoint kinase Chk1 is required for mammalian homologous recombination repair. Nat Cell Biol 7(2):195–201
Ma CX, Cai S, Li S, Ryan CE, Guo Z, Schaiff WT et al (2012) Targeting Chk1 in p53-deficient triple-negative breast cancer is therapeutically beneficial in human-in-mouse tumor models. J Clin Invest 122(4):1541–1552. Epub 2012/03/27
Ma CX, Janetka JW, Piwnica-Worms H (2010) Death by releasing the breaks: CHK1 inhibitors as cancer therapeutics. Trends Mol Med 17(2):88–96. Epub 2010/11/23
Mitchell C, Park M, Eulitt P, Yang C, Yacoub A, Dent P (2010) Poly(ADP-ribose) polymerase 1 modulates the lethality of CHK1 inhibitors in carcinoma cells. Mol Pharmacol 78(5):909–917. Epub 2010/08/11
Sausville EA, LoRusso P, Carducci MA, Barker PN, Agbo F, Oakes P et al (2011) Phase I dose-escalation study of AZD7762 in combination with gemcitabine (gem) in patients (pts) with advanced solid tumors. J Clin Oncol 29(suppl):abstr 3058
McNeely S, Conti C, Sheikh T, Patel H, Zabludoff S, Pommier Y et al (2010) Chk1 inhibition after replicative stress activates a double strand break response mediated by ATM and DNA-dependent protein kinase. Cell Cycle 9(5):995–1004. Epub 2010/02/18
Sorensen CS, Syljuasen RG (2012) Safeguarding genome integrity: the checkpoint kinases ATR, CHK1 and WEE1 restrain CDK activity during normal DNA replication. Nucleic Acids Res 40(2):477–486. Epub 2011/09/23
Toledo LI, Murga M, Fernandez-Capetillo O (2011) Targeting ATR and Chk1 kinases for cancer treatment: a new model for new (and old) drugs. Mol Oncol 5(4):368–373. Epub 2011/08/09
Xanthoudakis S, Smeyne RJ, Wallace JD, Curran T (1996) The redox/DNA repair protein, Ref-1, is essential for early embryonic development in mice. Proc Nat Acad Sci 93(17):8919–8923
Izumi T, Brown DB, Naidu CV, Bhakat KK, Macinnes MA, Saito H et al (2005) Two essential but distinct functions of the mammalian abasic endonuclease. Proc Natl Acad Sci USA 102(16):5739–5743
Fan Z, Beresford PJ, Zhang D, Xu Z, Novina CD, Yoshida A et al (2003) Cleaving the oxidative repair protein Ape1 enhances cell death mediated by granzyme A. Nat Immunol 4(2):145–153
Fishel ML, He Y, Reed AM, Chin-Sinex H, Hutchins GD, Mendonca MS et al (2008) Knockdown of the DNA repair and redox signaling protein Ape1/Ref-1 blocks ovarian cancer cell and tumor growth. DNA Repair (Amst) 7(2):177–186. Epub 2007/11/03
Bapat A, Fishel ML, Kelley MR (2009) Going ape as an approach to cancer therapeutics. Antioxid Redox Signal 11(3):651–668. Epub 2008/08/22
Abbotts R, Madhusudan S (2010) Human AP endonuclease 1 (APE1): from mechanistic insights to druggable target in cancer. Cancer Treat Rev 36(5):425–435. Epub 2010/01/09
Phillps JH, Brown DM, Grossman L (1996) The efficiency of induction of mutations by hydroxylamine. J Mol Biol 21(3):405–419. Epub 1966/11/28
Goldszer F, Tindell GL, Walle UK, Walle T (1981) Chemical trapping of labile aldehyde intermediates in the metabolism of propranolol and oxprenolol. Res Commun Chem Pathol Pharmacol 34(2):193–205. Epub 1981/11/01
Beger RD, Bolton PH (1998) Structures of apurinic and apyrimidinic sites in duplex DNAs. J Biol Chem 273(25):15565–15573. Epub 1998/06/23
Talpaert-Borle M, Liuzzi M (1983) Reaction of apurinic/apyrimidinic sites with [14C] methoxyamine. A method for the quantitative assay of AP sites in DNA. Biochim Biophys Acta 740(4):410–416. Epub 1983/09/09
Liu L, Taverna P, Whitacre CM, Chatterjee S, Gerson SL (1999) Pharmacologic disruption of base excision repair sensitizes mismatch repair-deficient and -proficient colon cancer cells to methylating agents. Clin Cancer Res 5(10):2908–2917
Goellner EM, Grimme B, Brown AR, Lin YC, Wang XH, Sugrue KF et al (2011) Overcoming temozolomide resistance in glioblastoma via dual inhibition of NAD+ biosynthesis and base excision repair. Cancer Res 71(6):2308–2317. Epub 2011/03/17
Tang JB, Svilar D, Trivedi RN, Wang XH, Goellner EM, Moore B et al (2011) N-methylpurine DNA glycosylase and DNA polymerase beta modulate BER inhibitor potentiation of glioma cells to temozolomide. Neurooncology 13(5):471–486. Epub 2011/03/08
Taverna P, Liu L, Hwang HS, Hanson AJ, Kinsella TJ, Gerson SL (2001) Methoxyamine potentiates DNA single strand breaks and double strand breaks induced by temozolomide in colon cancer cells. Mutat Res 485(4):269–281
Taverna P, Hwang HS, Schupp JE, Radivoyevitch T, Session NN, Reddy G et al (2003) Inhibition of base excision repair potentiates iododeoxyuridine-induced cytotoxicity and radiosensitization. Cancer Res 63(4):838–846. Epub 2003/02/20
Liu L, Yan L, Donze JR, Gerson SL (2003) Blockage of abasic site repair enhances antitumor efficacy of 1,3-bis-(2-chloroethyl)-1-nitrosourea in colon tumor xenografts. Mol Cancer Ther 2(10):1061–1066. Epub 2003/10/28
She M, Pan I, Sun L, Yeung SC (2005) Enhancement of manumycin A-induced apoptosis by methoxyamine in myeloid leukemia cells. Leukemia 19(4):595–602. Epub 2005/03/04
Bulgar AD, Snell M, Donze JR, Kirkland EB, Li L, Yang S et al (2010) Targeting base excision repair suggests a new therapeutic strategy of fludarabine for the treatment of chronic lymphocytic leukemia. Leukemia 24(10):1795–1799. Epub 2010/09/03
Kinsella TJ (2009) Coordination of DNA mismatch repair and base excision repair processing of chemotherapy and radiation damage for targeting resistant cancers. Clin Cancer Res 15(6):1853–1859. Epub 2009/02/26
Vermeulen C, Verwijs-Janssen M, Cramers P, Begg AC, Vens C (2007) Role for DNA polymerase beta in response to ionizing radiation. DNA Repair (Amst) 6(2):202–212
Bulgar AD, Weeks LD, Miao Y, Yang S, Xu Y, Guo C et al (2012) Removal of uracil by uracil DNA glycosylase limits pemetrexed cytotoxicity: overriding the limit with methoxyamine to inhibit base excision repair. Cell Death Dis 3:e252. Epub 2012/01/13
Srinivasan A, Wang L, Cline CJ, Xie Z, Sobol RW, Xie XQ et al (2012) Identification and characterization of human apurinic/apyrimidinic endonuclease-1 inhibitors. Biochemistry 51(31):6246–6259. Epub 2012/07/14
Rai G, Vyjayanti VN, Dorjsuren D, Simeonov A, Jadhav A, Wilson DM 3rd et al (2012) Synthesis, biological evaluation, and structure-activity relationships of a novel class of apurinic/apyrimidinic endonuclease 1 inhibitors. J Med chem 55(7):3101–3112. Epub 2012/03/30
Bapat A, Glass LS, Luo M, Fishel ML, Long EC, Georgiadis MM et al (2010) Novel small-molecule inhibitor of apurinic/apyrimidinic endonuclease 1 blocks proliferation and reduces viability of glioblastoma cells. J Pharmacol Exp Ther 334(3):988–998. Epub 2010/05/28
Zawahir Z, Dayam R, Deng J, Pereira C, Neamati N (2009) Pharmacophore guided discovery of small-molecule human apurinic/apyrimidinic endonuclease 1 inhibitors. J Med Chem 52(1):20–32
Al-Safi RI, Odde S, Shabaik Y, Neamati N (2012) Small-molecule inhibitors of APE1 DNA repair function: an overview. Curr Mol Pharmacol 5(1):14–35. Epub 2011/11/30
Sultana R, McNeill DR, Abbotts R, Mohammed MZ, Zdzienicka MZ, Qutob H et al (2012) Synthetic lethal targeting of DNA double-strand break repair deficient cells by human apurinic/apyrimidinic endonuclease inhibitors. Int J Cancer 131(10):2433–2444. Epub 2012/03/02
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K et al (2005) DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434(7035):864–870. Epub 2005/04/15
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T et al (2005) Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434(7035):907–913. Epub 2005/04/15
Peasland A, Wang LZ, Rowling E, Kyle S, Chen T, Hopkins A et al (2011) Identification and evaluation of a potent novel ATR inhibitor, NU6027, in breast and ovarian cancer cell lines. Br J Cancer 105(3):372–381. Epub 2011/07/07
Toledo LI, Murga M, Zur R, Soria R, Rodriguez A, Martinez S et al (2011) A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations. Nat Struct Mol Biol 18(6):721–727. Epub 2011/05/10
Rainey MD, Charlton ME, Stanton RV, Kastan MB (2008) Transient inhibition of ATM kinase is sufficient to enhance cellular sensitivity to ionizing radiation. Cancer Res 68(18):7466–7474. Epub 2008/09/17
Vens C, Begg AC (2010) Targeting base excision repair as a sensitization strategy in radiotherapy. Semin Radiat Oncol 20(4):241–249. Epub 2010/09/14
O’Connor MJ, Martin NM, Smith GC (2007) Targeted cancer therapies based on the inhibition of DNA strand break repair. Oncogene 26(56):7816–7824. Epub 2007/12/11
Nutley BP, Smith NF, Hayes A, Kelland LR, Brunton L, Golding BT et al (2005) Preclinical pharmacokinetics and metabolism of a novel prototype DNA-PK inhibitor NU7026. Br J Cancer 93(9):1011–1018. Epub 2005/10/27
Zhao Y, Thomas HD, Batey MA, Cowell IG, Richardson CJ, Griffin RJ et al (2006) Preclinical evaluation of a potent novel DNA-dependent protein kinase inhibitor NU7441. Cancer Res 66(10):5354–5362. Epub 2006/05/19
Quanz M, Berthault N, Roulin C, Roy M, Herbette A, Agrario C et al (2009) Small-molecule drugs mimicking DNA damage: a new strategy for sensitizing tumors to radiotherapy. Clin Cancer Res 15(4):1308–1316. Epub 2009/02/05
Munck JM, Batey MA, Zhao Y, Jenkins H, Richardson CJ, Cano C et al (2012) Chemosensitization of cancer cells by KU-0060648, A Dual Inhibitor of DNA-PK and PI-3 K. Mol Cancer Ther. Epub 2012/05/12
Veuger SJ, Curtin NJ, Richardson CJ, Smith GC, Durkacz BW (2003) Radiosensitization and DNA repair inhibition by the combined use of novel inhibitors of DNA-dependent protein kinase and poly(ADP-ribose) polymerase-1. Cancer Res 63(18):6008–6015. Epub 2003/10/03
Azad A, Jackson S, Cullinane C, Natoli A, Neilsen PM, Callen DF et al (2011) Inhibition of DNA-dependent protein kinase induces accelerated senescence in irradiated human cancer cells. Mol Cancer Res 9(12):1696–1707. Epub 2011/10/20
Gurley KE, Kemp CJ (2001) Synthetic lethality between mutation in ATM and DNA-PK(cs) during murine embryogenesis. Curr Biol 11(3):191–194. Epub 2001/03/07
Taccioli GE, Amatucci AG, Beamish HJ, Gell D, Xiang XH, Torres Arzayus MI et al (1998) Targeted disruption of the catalytic subunit of the DNA-PK gene in mice confers severe combined immunodeficiency and radiosensitivity. Immunity 9(3):355–366. Epub 1998/10/13
Mukherjee B, McEllin B, Camacho CV, Tomimatsu N, Sirasanagandala S, Nannepaga S et al (2009) EGFRvIII and DNA double-strand break repair: a molecular mechanism for radioresistance in glioblastoma. Cancer Res 69(10):4252–4259. Epub 2009/05/14
Golding SE, Rosenberg E, Neill S, Dent P, Povirk LF, Valerie K (2007) Extracellular signal-related kinase positively regulates ataxia telangiectasia mutated, homologous recombination repair, and the DNA damage response. Cancer Res 67(3):1046–1053. Epub 2007/02/07
Kriegs M, Kasten-Pisula U, Rieckmann T, Holst K, Saker J, Dahm-Daphi J et al (2010) The epidermal growth factor receptor modulates DNA double-strand break repair by regulating non-homologous end-joining. DNA Repair (Amst) 9(8):889–897. Epub 2010/07/10
Golding SE, Rosenberg E, Valerie N, Hussaini I, Frigerio M, Cockcroft XF et al (2009) Improved ATM kinase inhibitor KU-60019 radiosensitizes glioma cells, compromises insulin, AKT and ERK prosurvival signaling, and inhibits migration and invasion. Mol Cancer Ther 8(10):2894–2902. Epub 2009/10/08
Toulany M, Kehlbach R, Florczak U, Sak A, Wang S, Chen J et al (2008) Targeting of AKT1 enhances radiation toxicity of human tumor cells by inhibiting DNA-PKcs-dependent DNA double-strand break repair. Mol Cancer Ther 7(7):1772–1781. Epub 2008/07/23
Zaidi SH, Huddart RA, Harrington KJ (2009) Novel targeted radiosensitisers in cancer treatment. Curr Drug Discov Technol 6(2):103–134. Epub 2009/06/13
Dittmann K, Mayer C, Fehrenbacher B, Schaller M, Raju U, Milas L et al (2005) Radiation-induced epidermal growth factor receptor nuclear import is linked to activation of DNA-dependent protein kinase. J Biol Chem 280(35):31182–31189. Epub 2005/07/08
Kirshner J, Jobling MF, Pajares MJ, Ravani SA, Glick AB, Lavin MJ et al (2006) Inhibition of transforming growth factor-beta1 signaling attenuates ataxia telangiectasia mutated activity in response to genotoxic stress. Cancer Res 66(22):10861–10869. Epub 2006/11/09
Sinnberg T, Lasithiotakis K, Niessner H, Schittek B, Flaherty KT, Kulms D et al (2009) Inhibition of PI3K-AKT-mTOR signaling sensitizes melanoma cells to cisplatin and temozolomide. J Invest Dermatol 129(6):1500–1515. Epub 2008/12/17
Chen L, Han L, Shi Z, Zhang K, Liu Y, Zheng Y et al (2012) LY294002 enhances cytotoxicity of temozolomide in glioma by down-regulation of the PI3K/Akt pathway. Mol Med Rep 5(2):575–579. Epub 2011/11/17
Pal J, Fulciniti M, Nanjappa P, Buon L, Tai YT, Tassone P et al (2012) Targeting PI3K and RAD51 in Barrett’s adenocarcinoma: impact on DNA damage checkpoints, expression profile and tumor growth. Cancer Genomics Proteomics 9(2):55–66. Epub 2012/03/09
Kimbung S, Biskup E, Johansson I, Aaltonen K, Ottosson-Wadlund A, Gruvberger-Saal S et al (2012) Co-targeting of the PI3K pathway improves the response of BRCA1 deficient breast cancer cells to PARP1 inhibition. Cancer Lett 319(2):232–241. Epub 2012/01/24
Huang TT, D’Andrea AD (2006) Regulation of DNA repair by ubiquitylation. Nat Rev Mol Cell Biol 7(5):323–334 Epub 2006/04/25
Vucic D, Dixit VM, Wertz IE (2011) Ubiquitylation in apoptosis: a post-translational modification at the edge of life and death. Nat Rev Mol Cell Biol 12(7):439–452. Epub 2011/06/24
Frezza M, Schmitt S, Dou QP (2011) Targeting the ubiquitin-proteasome pathway: an emerging concept in cancer therapy. Curr Top Med Chem 11(23):2888–2905. Epub 2011/08/10
Ruschak AM, Slassi M, Kay LE, Schimmer AD (2011) Novel proteasome inhibitors to overcome bortezomib resistance. J Natl Cancer Inst 103(13):1007–1017. Epub 2011/05/25
Galli M, Van Gool F, Rongvaux A, Andris F, Leo O (2010) The nicotinamide phosphoribosyltransferase: a molecular link between metabolism, inflammation, and cancer. Cancer Res 70(1):8–11. Epub 2009/12/24
Wang T, Zhang X, Bheda P, Revollo JR, Imai S, Wolberger C (2006) Structure of Nampt/PBEF/visfatin, a mammalian NAD+ biosynthetic enzyme. Nat Struct Mol Biol 13(7):661–662. Epub 2006/06/20
Berger F, Lau C, Dahlmann M, Ziegler M (2005) Subcellular compartmentation and differential catalytic properties of the three human nicotinamide mononucleotide adenylyltransferase isoforms. J Biol Chem 280(43):36334–36341
Watson M, Roulston A, Belec L, Billot X, Marcellus R, Bedard D et al (2009) The small molecule GMX1778 is a potent inhibitor of NAD+ biosynthesis: strategy for enhanced therapy in nicotinic acid phosphoribosyltransferase 1-deficient tumors. Mol Cell Biol 29(21):5872–5888. Epub 2009/08/26
Fleischer TC, Murphy BR, Flick JS, Terry-Lorenzo RT, Gao ZH, Davis T et al Chemical proteomics identifies Nampt as the target of CB30865, an orphan cytotoxic compound. Chem Biol 17(6):659–664. Epub 2010/07/09
Olesen UH, Christensen MK, Bjorkling F, Jaattela M, Jensen PB, Sehested M et al (2008) Anticancer agent CHS-828 inhibits cellular synthesis of NAD. Biochem Biophys Res Commun 367(4):799–804
Zhang RY, Qin Y, Lv XQ, Wang P, Xu TY, Zhang L et al (2011) A fluorometric assay for high-throughput screening targeting nicotinamide phosphoribosyltransferase. Anal Biochem 412(1):18–25. Epub 2011/01/08
Khan JA, Tao X, Tong L (2006) Molecular basis for the inhibition of human NMPRTase, a novel target for anticancer agents. Nat Struct Mol Biol 13(7):582–588
Hasmann M, Schemainda I (2003) FK866, a highly specific noncompetitive inhibitor of nicotinamide phosphoribosyltransferase, represents a novel mechanism for induction of tumor cell apoptosis. Cancer Res 63(21):7436–7442
Billington RA, Genazzani AA, Travelli C, Condorelli F (2008) NAD depletion by FK866 induces autophagy. Autophagy 4(3):385–387. Epub 2008/01/30
Pogrebniak A, Schemainda I, Azzam K, Pelka-Fleischer R, Nussler V, Hasmann M (2006) Chemopotentiating effects of a novel NAD biosynthesis inhibitor, FK866, in combination with antineoplastic agents. Eur J Med Res 11(8):313–321
Bi TQ, Che XM, Liao XH, Zhang DJ, Long HL, Li HJ et al (2011) Overexpression of nampt in gastric cancer and chemopotentiating effects of the nampt inhibitor FK866 in combination with fluorouracil. Oncol Rep 26(5):1251–1257. Epub 2011/07/12
Travelli C, Drago V, Maldi E, Kaludercic N, Galli U, Boldorini R et al (2011) Reciprocal potentiation of the antitumoral activities of FK866, an inhibitor of nicotinamide phosphoribosyltransferase, and etoposide or cisplatin in neuroblastoma cells. J Pharmacol Exp Ther 338(3):829–840. Epub 2011/06/21
Zoppoli G, Cea M, Soncini D, Fruscione F, Rudner J, Moran E et al (2010) Potent synergistic interaction between the Nampt inhibitor APO866 and the apoptosis activator TRAIL in human leukemia cells. Exp Hematol 38(11):979–988. Epub 2010/08/11
Olesen UH, Thougaard AV, Jensen PB, Sehested M (2010) A preclinical study on the rescue of normal tissue by nicotinic acid in high-dose treatment with APO866, a specific nicotinamide phosphoribosyltransferase inhibitor. Mol Cancer Ther 9(6):1609–1617
Goellner EM, Svilar D, Almeida KH, Sobol RW (2012) Targeting DNA polymerase β for therapeutic intervention. Curr Mol Pharmacol 5(1):68–87. Epub 2011/11/30
Martin LP, Hamilton TC, Schilder RJ (2008) Platinum resistance: the role of DNA repair pathways. Clin Cancer Res 14(5):1291–1295. Epub 2008/03/05
Orelli B, McClendon TB, Tsodikov OV, Ellenberger T, Niedernhofer LJ, Scharer OD (2010) The XPA-binding domain of ERCC1 is required for nucleotide excision repair but not other DNA repair pathways. J Biol Chem 285(6):3705–3712. Epub 2009/11/27
Su Y, Orelli B, Madireddy A, Niedernhofer LJ, Scharer OD (2012) Multiple DNA binding domains mediate the function of the ERCC1-XPF protein in nucleotide excision repair. J Biol Chem 287(26):21846–21855. Epub 2012/05/02
Tsodikov OV, Ivanov D, Orelli B, Staresincic L, Shoshani I, Oberman R et al (2007) Structural basis for the recruitment of ERCC1-XPF to nucleotide excision repair complexes by XPA. Embo J 26(22):4768–4776. Epub 2007/10/20
Sijbers AM, van der Spek PJ, Odijk H, van den Berg J, van Duin M, Westerveld A et al (1996) Mutational analysis of the human nucleotide excision repair gene ERCC1. Nucleic Acids Res 24(17):3370–3380. Epub 1996/09/01
Barakat KH, Torin Huzil J, Luchko T, Jordheim L, Dumontet C, Tuszynski J (2009) Characterization of an inhibitory dynamic pharmacophore for the ERCC1-XPA interaction using a combined molecular dynamics and virtual screening approach. J Mol Graph Model 28(2):113–130. Epub 2009/05/29
Lee-Kwon W, Park D, Bernier M (1998) Involvement of the ras/extracellular signal-regulated kinase signalling pathway in the regulation of ERCC-1 mRNA levels by insulin. Biochem J 331 (Pt 2):591–597. Epub 1998/06/11
Yadav A, Kumar B, Teknos TN, Kumar P (2011) Sorafenib enhances the antitumor effects of chemoradiation treatment by downregulating ERCC-1 and XRCC-1 DNA repair proteins. Mol Cancer Ther 10(7):1241–1251. Epub 2011/05/10
Burgers PM, Koonin EV, Bruford E, Blanco L, Burtis KC, Christman MF et al (2001) Eukaryotic DNA polymerases: proposal for a revised nomenclature. J Biol Chem 276(47):43487–43490
Lange SS, Takata K, Wood RD (2011) DNA polymerases and cancer. Nat Rev Cancer 11(2):96–110. Epub 2011/01/25
Bebenek K, Kunkel TA (2004) Functions of DNA polymerases. Adv Protein Chem 69:137–65. Epub 2004/12/14
Wilson SH, Beard WA, Shock DD, Batra VK, Cavanaugh NA, Prasad R et al (2010) Base excision repair and design of small molecule inhibitors of human DNA polymerase beta. Cell Mol Life Sci 67(21):3633–3647. Epub 2010/09/17
Barakat KH, Gajewski MM, Tuszynski JA (2012) DNA polymerase beta (pol beta) inhibitors: a comprehensive overview. Drug discovery today 17(15–16):913–920. Epub 2012/05/09
Higgins GS, Prevo R, Lee YF, Helleday T, Muschel RJ, Taylor S et al (2010) A small interfering RNA screen of genes involved in DNA repair identifies tumor-specific radiosensitization by POLQ knockdown. Cancer Res 70(7):2984–2993. Epub 2010/03/18
Higgins GS, Harris AL, Prevo R, Helleday T, McKenna WG, Buffa FM (2010) Overexpression of POLQ confers a poor prognosis in early breast cancer patients. Oncotarget 1(3):175–184. Epub 2010/08/12
Lemee F, Bergoglio V, Fernandez-Vidal A, Machado-Silva A, Pillaire MJ, Bieth A et al (2010) DNA polymerase theta up-regulation is associated with poor survival in breast cancer, perturbs DNA replication, and promotes genetic instability. Proc Natl Acad Sci USA 107(30):13390–13395. Epub 2010/07/14
Slupphaug G, Mol CD, Kavli B, Arvai AS, Krokan HE, Tainer JA (1996) A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature 384(6604):87–92. Epub 1996/11/07
Engelward BP, Allan JM, Dreslin AJ, Kelly JD, Wu MM, Gold B et al (1998) A chemical and genetic approach together define the biological consequences of 3-methyladenine lesions in the mammalian genome. J Biol Chem 273(9):5412–5418
Noren Hooten N, Kompaniez K, Barnes J, Lohani A, Evans MK (2011) Poly(ADP-ribose) polymerase 1 (PARP-1) binds to 8-oxoguanine-DNA glycosylase (OGG1). J Biol Chem 286(52):44679–44690. Epub 2011/11/08
Endres M, Biniszkiewicz D, Sobol RW, Harms C, Ahmadi M, Lipski A et al (2004) Increased postischemic brain injury in mice deficient in uracil-DNA glycosylase. J Clin Invest 113(12):1711–1721
Kruman II, Schwartz E, Kruman Y, Cutler RG, Zhu X, Greig NH et al (2004) Suppression of uracil-DNA glycosylase induces neuronal apoptosis. J Biol Chem 279(42):43952–43960
Imai K, Slupphaug G, Lee WI, Revy P, Nonoyama S, Catalan N et al (2003) Human uracil-DNA glycosylase deficiency associated with profoundly impaired immunoglobulin class-switch recombination. Nat Immunol 4(10):1023–1028. Epub 2003/09/06
Wang Z, Mosbaugh DW. Uracil-DNA glycosylase inhibitor gene of bacteriophage PBS2 encodes a binding protein specific for uracil-DNA glycosylase. J Biol Chem. 1989;264(2):1163-71. Epub 1989/01/15
Komander D, Clague MJ, Urbe S (2009) Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol 10(8):550–563. Epub 2009/07/25
Shabek N, Ciechanover A (2010) Degradation of ubiquitin: the fate of the cellular reaper. Cell Cycle 9(3):523–530. Epub 2010/01/29
Mattern MR, Wu J, Nicholson B (2012) Ubiquitin-based anticancer therapy: carpet bombing with proteasome inhibitors vs surgical strikes with E1, E2, E3, or DUB inhibitors. Biochim Biophys Acta 1823(11):2014–2021. Epub 2012/05/23
Opferman JT, Green DR (2010) DUB-le trouble for cell survival. Cancer Cell 17(2):117–119. Epub 2010/02/18
Altun M, Kramer HB, Willems LI, McDermott JL, Leach CA, Goldenberg SJ et al (2011) Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chem Biol 18(11):1401–1412. Epub 2011/11/29
Kapuria V, Peterson LF, Fang D, Bornmann WG, Talpaz M, Donato NJ (2010) Deubiquitinase inhibition by small-molecule WP1130 triggers aggresome formation and tumor cell apoptosis. Cancer Res 70(22):9265–9276. Epub 2010/11/04
Kramer HB, Nicholson B, Kessler BM, Altun M (2012) Detection of ubiquitin-proteasome enzymatic activities in cells: application of activity-based probes to inhibitor development. Biochim Biophys Acta 1823(11):2029–2037. Epub 2012/05/23
Seiberlich V, Goldbaum O, Zhukareva V, Richter-Landsberg C (2012) The small molecule inhibitor PR-619 of deubiquitinating enzymes affects the microtubule network and causes protein aggregate formation in neural cells: implications for neurodegenerative diseases. Biochim Biophys Acta 1823(11):2057–2068. Epub 2012/05/09
Chernikova SB, Game JC, Brown JM (2012) Inhibiting homologous recombination for cancer therapy. Cancer Biol Ther 13(2):61–68. Epub 2012/02/18
Huang F, Motlekar NA, Burgwin CM, Napper AD, Diamond SL, Mazin AV (2011) Identification of specific inhibitors of human RAD51 recombinase using high-throughput screening. ACS chemical biology 6(6):628–635. Epub 2011/03/25
Quiros S, Roos WP, Kaina B (2011) Rad51 and BRCA2–New molecular targets for sensitizing glioma cells to alkylating anticancer drugs. PLoS ONE 6(11):e27183. Epub 2011/11/11
Short SC, Giampieri S, Worku M, Alcaide-German M, Sioftanos G, Bourne S et al (2011) Rad51 inhibition is an effective means of targeting DNA repair in glioma models and CD133+ tumor-derived cells. Neurooncology 13(5):487–499. Epub 2011/03/03
Sprong D, Janssen HL, Vens C, Begg AC (2005) Resistance of hypoxic cells to ionizing radiation is influenced by homologous recombination status. Int J Radiat Oncol Biol Phys 64(2):562–572. Epub 2005/12/14
Neijenhuis S, Verwijs-Janssen M, van den Broek LJ, Begg AC, Vens C (2010) Targeted radiosensitization of cells expressing truncated DNA polymerase {beta}. Cancer Res 70(21):8706–8714. Epub 2010/10/28
Slade D, Dunstan MS, Barkauskaite E, Weston R, Lafite P, Dixon N et al (2011) The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase. Nature 477(7366):616–620. Epub 2011/09/06
Brochu G, Duchaine C, Thibeault L, Lagueux J, Shah GM, Poirier GG (1994) Mode of action of poly(ADP-ribose) glycohydrolase. Biochim Biophys Acta 1219(2):342–350
Affar EB, Germain M, Winstall E, Vodenicharov M, Shah RG, Salvesen GS et al (2001) Caspase-3-mediated processing of poly(ADP-ribose) glycohydrolase during apoptosis. J Biol Chem 276(4):2935–42. Epub 2000/10/29
Li Q, Li M, Wang YL, Fauzee NJ, Yang Y, Pan J et al (2012) RNA interference of PARG could inhibit the metastatic potency of colon carcinoma cells via PI3-kinase/Akt pathway. Cell Physiol Biochem Int J Exp Cell Physiol Biochem Pharmacol 29(3–4):361–72. Epub 2012/04/18
Koh DW, Lawler AM, Poitras MF, Sasaki M, Wattler S, Nehls MC et al (2004) Failure to degrade poly(ADP-ribose) causes increased sensitivity to cytotoxicity and early embryonic lethality. Proc Natl Acad Sci USA 101(51):17699–17704
Ying W, Swanson RA (2000) The poly(ADP-ribose) glycohydrolase inhibitor gallotannin blocks oxidative astrocyte death. Neuroreport 11(7):1385–8. Epub 2000/06/07
Sun Y, Zhang T, Wang B, Li H, Li P (2012) Tannic acid, an inhibitor of poly(ADP-ribose) glycohydrolase, sensitizes ovarian carcinoma cells to cisplatin. Anti-cancer drugs 23(9):979–990. Epub 2012/07/13
Formentini L, Arapistas P, Pittelli M, Jacomelli M, Pitozzi V, Menichetti S et al (2008) Mono-galloyl glucose derivatives are potent poly(ADP-ribose) glycohydrolase (PARG) inhibitors and partially reduce PARP-1-dependent cell death. Br J Pharmacol 155(8):1235–1249. Epub 2008/09/23
Cuzzocrea S, Mazzon E, Genovese T, Crisafulli C, Min WK, Di Paola R et al (2007) Role of poly(ADP-ribose) glycohydrolase in the development of inflammatory bowel disease in mice. Free Radic Biol Med 42(1):90–105
Cuzzocrea S, Genovese T, Mazzon E, Crisafulli C, Min W, Di Paola R et al (2006) Poly(ADP-ribose) glycohydrolase activity mediates post-traumatic inflammatory reaction after experimental spinal cord trauma. J Pharmacol Exp Ther 319(1):127–138
Cuzzocrea S, Di Paola R, Mazzon E, Cortes U, Genovese T, Muia C et al (2005) PARG activity mediates intestinal injury induced by splanchnic artery occlusion and reperfusion. Faseb J 19(6):558–566
Falsig J, Christiansen SH, Feuerhahn S, Burkle A, Oei SL, Keil C et al (2004) Poly(ADP-ribose) glycohydrolase as a target for neuroprotective intervention: assessment of currently available pharmacological tools. Eur J Pharmacol 497(1):7–16
Lu XC, Massuda E, Lin Q, Li W, Li JH, Zhang J (2003) Post-treatment with a novel PARG inhibitor reduces infarct in cerebral ischemia in the rat. Brain Res 978(1–2):99–103. Epub 2003/07/02
Steffen JD, Coyle DL, Damodaran K, Beroza P, Jacobson MK (2011) Discovery and structure-activity relationships of modified salicylanilides as cell permeable inhibitors of poly(ADP-ribose) glycohydrolase (PARG). J Med Chem 54(15):5403–5413. Epub 2011/06/23
Okita N, Ashizawa D, Ohta R, Abe H, Tanuma S (2010) Discovery of novel poly(ADP-ribose) glycohydrolase inhibitors by a quantitative assay system using dot-blot with anti-poly(ADP-ribose). Biochem Biophys Res Commun 392(4):485–489. Epub 2010/01/19
Kaye SB, Lubinski J, Matulonis U, Ang JE, Gourley C, Karlan BY et al (2012) Phase II, open-label, randomized, multicenter study comparing the efficacy and safety of olaparib, a poly (ADP-ribose) polymerase inhibitor, and pegylated liposomal doxorubicin in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer. J Clin Oncol 30(4):372–379. Epub 2011/12/29
Rajan A, Carter CA, Kelly RJ, Gutierrez M, Kummar S, Szabo E et al (2012) A phase I combination study of olaparib with cisplatin and gemcitabine in adults with solid tumors. Clin Cancer Res 18(8):2344–2351. Epub 2012/03/01
van Vuurden DG, Hulleman E, Meijer OL, Wedekind LE, Kool M, Witt H et al (2011) PARP inhibition sensitizes childhood high grade glioma, medulloblastoma and ependymoma to radiation. Oncotarget 2(12):984–996. Epub 2011/12/21
Clark CC, Weitzel JN, O’Connor TR (2012) Enhancement of synthetic lethality via combinations of ABT-888, a PARP inhibitor, and carboplatin in vitro and in vivo using BRCA1 and BRCA2 isogenic models. Mol Cancer Ther 11(9):1948–1958. Epub 2012/07/11
Kummar S, Chen A, Ji J, Zhang Y, Reid JM, Ames M et al (2011) Phase I study of PARP inhibitor ABT-888 in combination with topotecan in adults with refractory solid tumors and lymphomas. Cancer Res 71(17):5626–5634. Epub 2011/07/29
Palma JP, Wang YC, Rodriguez LE, Montgomery D, Ellis PA, Bukofzer G et al (2009) ABT-888 confers broad in vivo activity in combination with temozolomide in diverse tumors. Clin Cancer Res 15(23):7277–7290. Epub 2009/11/26
Donawho CK, Luo Y, Penning TD, Bauch JL, Bouska JJ, Bontcheva-Diaz VD et al (2007) ABT-888, an orally active poly(ADP-ribose) polymerase inhibitor that potentiates DNA-damaging agents in preclinical tumor models. Clin Cancer Res 13(9):2728–2737. Epub 2007/05/03
Kummar S, Ji J, Morgan R, Lenz HJ, Puhalla SL, Belani CP et al (2012) A phase I study of veliparib in combination with metronomic cyclophosphamide in adults with refractory solid tumors and lymphomas. Clin Cancer Res 18(6):1726–1734. Epub 2012/02/07
Barreto-Andrade JC, Efimova EV, Mauceri HJ, Beckett MA, Sutton HG, Darga TE et al (2011) Response of human prostate cancer cells and tumors to combining PARP inhibition with ionizing radiation. Mol Cancer Ther 10(7):1185–1193. Epub 2011/05/17
Efimova EV, Mauceri HJ, Golden DW, Labay E, Bindokas VP, Darga TE et al (2010) Poly(ADP-ribose) polymerase inhibitor induces accelerated senescence in irradiated breast cancer cells and tumors. Cancer Res 70(15):6277–6282. Epub 2010/07/09
Nowsheen S, Bonner JA, Yang ES (2011) The poly(ADP-Ribose) polymerase inhibitor ABT-888 reduces radiation-induced nuclear EGFR and augments head and neck tumor response to radiotherapy. Radiother Oncol 99(3):331–338. Epub 2011/07/02
Apisarnthanarax N, Wood GS, Stevens SR, Carlson S, Chan DV, Liu L et al (2012) Phase I clinical trial of O6-benzylguanine and topical carmustine in the treatment of cutaneous T-cell lymphoma, mycosis fungoides type. Arch Dermatol 148(5):613–620. Epub 2012/01/18
Warren KE, Gururangan S, Geyer JR, McLendon RE, Poussaint TY, Wallace D et al (2012) A phase II study of O6-benzylguanine and temozolomide in pediatric patients with recurrent or progressive high-grade gliomas and brainstem gliomas: a pediatric brain tumor consortium study. J Neurooncol 106(3):643–649. Epub 2011/10/05
Kefford RF, Thomas NP, Corrie PG, Palmer C, Abdi E, Kotasek D et al (2009) A phase I study of extended dosing with lomeguatrib with temozolomide in patients with advanced melanoma. Br J Cancer 100(8):1245–1249. Epub 2009/04/16
Watson AJ, Middleton MR, McGown G, Thorncroft M, Ranson M, Hersey P et al (2009) O(6)-methylguanine-DNA methyltransferase depletion and DNA damage in patients with melanoma treated with temozolomide alone or with lomeguatrib. Br J Cancer 100(8):1250–1256. Epub 2009/04/16
Ma Z, Yao G, Zhou B, Fan Y, Gao S, Feng X (2012) The Chk1 inhibitor AZD7762 sensitises p53 mutant breast cancer cells to radiation in vitro and in vivo. Mol Med Rep 6(4):897–903. Epub 2012/07/25
Montano R, Chung I, Garner KM, Parry D, Eastman A (2011) Preclinical development of the novel Chk1 inhibitor SCH900776 in combination with DNA-damaging agents and antimetabolites. Mol Cancer Ther 11(2):427–438. Epub 2011/12/29
Ferrao PT, Bukczynska EP, Johnstone RW, McArthur GA (2012) Efficacy of CHK inhibitors as single agents in MYC-driven lymphoma cells. Oncogene 31(13):1661–1672. Epub 2011/08/16
Hickson I, Zhao Y, Richardson CJ, Green SJ, Martin NM, Orr AI et al (2004) Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res 64(24):9152–9159. Epub 2004/12/18
Leahy JJ, Golding BT, Griffin RJ, Hardcastle IR, Richardson C, Rigoreau L et al (2004) Identification of a highly potent and selective DNA-dependent protein kinase (DNA-PK) inhibitor (NU7441) by screening of chromenone libraries. Bioorg Med Chem Lett 14(24):6083–6087
Devun F, Bousquet G, Biau J, Herbette A, Roulin C, Berger F et al (2012) Preclinical study of the DNA repair inhibitor Dbait in combination with chemotherapy in colorectal cancer. J gastroenterol 47(3):266–275. Epub 2011/11/10
Kim CH, Park SJ, Lee SH (2002) A targeted inhibition of DNA-dependent protein kinase sensitizes breast cancer cells following ionizing radiation. J Pharmacol Exp Ther 303(2):753–759. Epub 2002/10/22
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Vens, C., Sobol, R.W. (2013). Targeting DNA Repair Pathways for Cancer Therapy. In: Johnson, D. (eds) Cell Death Signaling in Cancer Biology and Treatment. Cell Death in Biology and Diseases. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4614-5847-0_6
Download citation
DOI: https://doi.org/10.1007/978-1-4614-5847-0_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4614-5846-3
Online ISBN: 978-1-4614-5847-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)