Abstract
Here, we focus on the implications of two cellular degradation pathways on prion replication and clearance. The first one is autophagy which can have a promoting and inhibiting role in prion infection. Lysosomal prion clearance can be enhanced in vitro and in vivo by drug-induced activation of autophagy. More recent work revealed that a certain level of autophagy is needed for establishing acute and persistent prion infection, implicating that autophagy might represent a functional equivalent for a disaggregase function. Such one was postulated for seed fragmentation in prion propagation, similar to sonication in PMCA or Hsp104 in yeast prion biology. The second pathway described here is the proteasomal one. We have challenged various cell lines by inducing ER stress or compromising proteasomal activity and analyzed the effects on PrP metabolism strictly in the secretory pathway. Both events led to enhanced detection of PrP aggregates and significant increase of PrPSc in prion-infected cells, which could be reversed by overexpression of proteins of the cellular quality control. These findings suggest a novel pathway which possibly provides additional substrate and template for prion formation when protein clearance by the proteasome is impaired and point to mechanisms which might play a role in prion de novo generation in sporadic prion diseases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Prion propagation
- Prion clearance
- Autophagy
- Lysosomal clearance
- Endosomal recycling
- Disaggregase
- Cellular quality control
- ER stress
- Proteasomal impairment
- Sporadic CJD
11.1 Autophagy in Health and Disease
Degradation and recycling of organelles or cytoplasmic proteins and protein aggregates is mediated by an intracellular bulk degradation process called macroautophagy (referred here as autophagy). The name autophagy originally denotes a cellular self-digestion (self-eating) program, in its simplest form as a single cell’s adaptation to starvation. During autophagy, portions of the cytosol are engulfed by a membrane sac resulting in a double-membrane vesicle, called autophagosome, which delivers cytoplasmic cargo to endosomes and lysosomes (Klionsky 2007; Mizushima 2009, 2011). After fusion with lysosomes, the protein and organelle contents of the autophagolysosome are degraded by acidic lysosomal hydrolases and recycled. Beyond its classical role in nutrient supply under starvation and turnover of organelles and proteins, autophagy greatly contributes to various physiological processes such as intracellular cleansing, differentiation, longevity, elimination of invading pathogens, antigen transport to the innate and adaptive immune system, or counteracting endoplasmic reticulum stress (Levine and Kroemer 2008; Mizushima et al. 2008; Meijer and Codogno 2004). Besides its role in physiology, autophagy is also directly implicated in pathophysiology and disease. Autophagy plays a role in cancer, in a number of infectious and inflammatory diseases, and in protein misfolding diseases (Levine and Kroemer 2008; Mizushima et al. 2008; Chu et al. 2009; Batlevi and La Spada 2011; Martinez-Vicente and Cuervo 2007). With respect to the importance of tight regulation of autophagy, perhaps the most fundamental point is that either too little or too much autophagy can be deleterious, a complex balance resulting in its dual role in survival and adaptation or cell death. However, in response to most forms of cellular stress, autophagy plays a protective role. Autophagy has long been defined as a form of non-apoptotic (type II) programmed cell death. However, a consensus is emerging that autophagy might be a cell death impostor which, in reality, functions primarily to promote cellular and organism health (Levine and Kroemer 2008).
Autophagy occurs at basal, constitutive levels in cells. In tissues where cells do not divide, such as neurons and myocytes, basal autophagy is of great relevance (Hara et al. 2006; Komatsu et al. 2006). Several studies suggest a crucial role of autophagy in neurodegenerative diseases, including Alzheimer’s disease (AD), Parkinson’s disease (PD), tauopathies, and polyglutamine expansion diseases like Huntington’s disease (HD) (Nixon 2005; Rubinsztein et al. 2005; Rubinsztein 2006; Ventruti and Cuervo 2007). A number of in vivo studies showed that conventional autophagy knockout mice die during embryogenesis or the neonatal period. Mice with neural-tissue-specific knockouts of these genes survive the postnatal starvation period. However, these mice develop progressive motor deficits and display abnormal reflexes, and ubiquitin-positive inclusion bodies accumulate in their neurons (Hara et al. 2006; Komatsu et al. 2006). Studies showed that the CNS displays only low levels of autophagosomes under normal conditions and even after starvation, but it was also demonstrated that constitutive turnover of cytosolic contents by autophagy is indispensable, even in the absence of expression of any disease-associated mutant proteins (Nixon 2005; Mizushima et al. 2004).
The requirement for autophagy is even more evident under disease conditions, and levels of autophagosomes can be dramatically increased in injured or degenerating neurons. Available data state that autophagy has a beneficial effect of protecting against neurodegeneration. One idea is that autophagy eliminates aggregated and aggregate-prone proteins (Cherra et al. 2010; Jeong et al. 2009; Korolchuk and Rubinsztein 2011; Krainc 2010). Thus, it is reasonable to assume that autophagy could be a therapeutic target for treatment of these neurodegenerative diseases (Rubinsztein 2006). Recent studies underlined that degradation of disease-related mutant proteins is highly dependent on autophagy. It was shown that the clearance of aggregate-prone proteins, such as mutant huntingtin fragments or mutant forms of α-synuclein, respectively, can be mediated by autophagy (Martinez-Vicente and Cuervo 2007; Winslow and Rubinsztein 2008; Chew et al. 2011; Vives-Bauza et al. 2010). Animal models of HD and of other proteinopathies revealed that treatment with rapamycin, a known inducer of autophagy via the mTOR pathway, accelerates the clearance of toxic proteins. Induction of autophagy, mediated by compounds like lithium and trehalose, also has been seen to accelerate the clearance of mutant huntingtin and α-synucleins (Sarkar et al. 2007; Sarkar and Rubinsztein 2008). The beneficial effect of upregulated autophagy has also been described for other diseases associated with aggregate-prone proteins, such as AD and ALS. The possible involvement of autophagy in prion pathogenesis was first described morphologically in form of autophagic vacuoles (Boellaard et al. 1991). These were detected in neurons of prion-infected mice and hamsters and of patients with CJD and other human prion disorders (Liberski et al. 2004, 2008; Sikorska et al. 2004). The appearance of multilamellar bodies and autophagic vacuoles was observed in prion-infected cultured neuronal cells (Schatzl et al. 1997).
11.1.1 Drug-Induced Autophagy Counteracts Prion Infection In Vitro and In Vivo
Our research program over the last years attempted to decipher the molecular and cellular mechanisms which underlie prion diseases (Gilch et al. 2007, 2008; Gilch and Schatzl 2003; Krammer et al. 2009a). Such molecular understanding was then used by us to define and characterize novel molecular targets against prion diseases. One example is our finding that the clearance of prions can be intensified by drug-induced increase of the autophagic flux, both in vitro and in vivo (Aguib et al. 2009; Heiseke et al. 2009, 2010; Ertmer et al. 2004, 2007; Yun et al. 2007). As autophagosomes fuse with the endosomal–lysosomal machinery for final degradation in autophagolysosomes, PrPSc present in endosomal–lysosomal compartments can be subject to changes in the activity of autophagy (Heiseke et al. 2010). PrPSc/prions produced within cells only indirectly have access to autophagy pathways. They are neither a direct substrate for autophagy, nor is autophagy in any way specific for prions/PrPSc and their clearance.
We showed that the c-abl inhibitor imatinib (Gleevec), a drug used to treat chronic myelogenous leukemia, activates the lysosomal degradation of PrPSc and, at the same time, induces autophagy (Ertmer et al. 2004, 2007). In prion-infected mice, imatinib treatment at an early phase of peripheral infection delayed neuroinvasion and onset of clinical disease (Yun et al. 2007). Since imatinib is not effectively crossing the blood–brain barrier, there was no beneficial effect when the process of neuroinvasion was already completed (Yun et al. 2007). Follow-up work with other chemical inducers of autophagy corroborated these findings. We showed recently that lithium and trehalose enhance the clearance of PrPSc by induction of autophagy (Aguib et al. 2009; Heiseke et al. 2009). To demonstrate that indeed induction of autophagy is the underlying mechanism, we inhibited autophagy by pharmacological interference or siRNA gene silencing of essential members of the autophagy machinery. Such co-treatment impaired or antagonized the capacity of compound-induced autophagy in reducing cellular levels of PrPSc. Besides compounds inducing autophagy in an mTor-independent manner (e.g., lithium, trehalose), we studied rapamycin, a drug widely used to activate autophagy by inhibiting mTor. Rapamycin also reduced PrPSc, showing that both autophagy-inducing pathways, mTor-dependent and mTor-independent, can be involved in the degradation of PrPSc (Heiseke et al. 2009, 2010).
To test whether autophagy-inducing compounds are candidates for therapeutic approaches against prion infection, we treated intraperitoneally prion-infected mice. Oral rapamycin treatment of prion-infected mice initiated in the last third of incubation time, mimicking a preclinical therapeutic situation, showed a significant prolongation of prion incubation times as compared to mock-treated control mice (Heiseke et al. 2009). Similar findings were obtained with lithium, although less uniform (Heiseke et al. 2009). Trehalose treatment did not prolong incubation times, but showed effects on PrPSc levels in spleen, and depending on when treatment was started, the peripheral accumulation of PrPSc was delayed (Aguib et al. 2009). As was the case with imatinib (Yun et al. 2007), this reflects that the process of neuroinvasion was decelerated. Taken together, these in vivo studies strongly indicate that autophagy-inducing compounds are beneficial in prion disease scenarios and ask for further studies, including also combination of drugs (Heiseke et al. 2010).
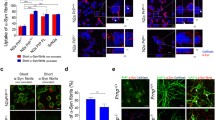
Before doing so, we studied another compound which has a promising bioavailability profile: the selective estrogen receptor modulator tamoxifen, a widely used anticancer drug (Heel et al. 1978). In fact, tamoxifen was the most potent enhancer of PrPSc clearance in our recent studies. We assessed its mode of action and found that tamoxifen treatment leads to robust clearance of prion infection after only a few days. Attenuation of the autophagic pathway (e.g., knockdown of Beclin-1 or Atg5) antagonized these effects. Time kinetics experiments showed that induction of autophagy is also of importance after prions are taken up by the cell, restricting the accumulation of intracellular aggregated prion protein. In vivo, tamoxifen treatment was able to reduce the PrPSc load in spleens, prolonged survival in infected animals, and led to reduced microgliosis in brain tissue of treated mice (Fig. 11.1).
Effect of tamoxifen in in vivo prion infection. (a) Prolonged survival times in tamoxifen-treated mice. Oral treatment with tamoxifen initiated at day 100 post intracerebral infection with prion strain 139A. Solid line depicts control mice (mean 170.6 ± 7.9 days) and broken tamoxifen-treated mice (mean 186.4 ± 13.6 days, *p > 0.01); n = 7. (b) Reduced PrPSc level in spleens. Immunoblot detection of PrPSc in spleens of mock- (lanes 1, 3) and tamoxifen-treated (lanes 2, 4) mice at terminal time points. (c) Reduced microgliosis in brain tissue of tamoxifen-treated mice. Immunohistochemical analysis of disease-associated microgliosis via detection of ionized calcium binding adapter molecule 1 (Iba1) in hippocampus (left row) and cortex (right row) of treated (lower row) and nontreated (upper row) mice at 125 dpi. Representative sections are shown in a magnification of ×400
Taken together, our data convincingly show that autophagy is a potent modifier of the cellular clearance of prions and that chemically induced autophagy shifts the delicate equilibrium between propagation and clearance of prions towards the latter.
11.1.2 A Second Role for Autophagy in Prion Infection: Basal Autophagy Is Required for Establishing Prion Infection and Might Provide Disaggregase Function
Our further studies focused on a novel biological function of autophagy recently found by us: Basal (i.e., normal, non-induced) autophagy is required for cellular prion propagation. We believe that autophagy compartments function as a biological “disaggregase” providing fragmentation activity which is postulated in protein aggregation/disaggregation (Borchsenius et al. 2006; Shorter 2011). PMCA technology uses physical disintegration to accomplish this task (Castilla et al. 2005). In yeast prion biology, Hsp104, for which no mammalian homologue is known, mainly fulfills this part (Shorter 2008, 2011; Chernoff et al. 1995; Lindquist et al. 1995). The molecular characterization of this finding might be of significance also for other diseases involving prion-like mechanisms. For no neurodegenerative disease, it is understood how aggregates build up in the cell and exit and enter neighboring cells, starting the cycle there again (Brundin et al. 2010; Kaganovich et al. 2008; Ren et al. 2009; Krammer et al. 2009b).
Our initial finding which brought us to look into this direction was that autophagy is transiently induced when cells start propagating PrPSc, implicating a more general role of autophagy in prion conversion. We infected neuronal cells with different known susceptibilities to primary prion infection and analyzed whether levels of autophagy were modulated. The recipient cells harbored a prion protein tagged with an epitope for mAb 3F4 (Maas et al. 2007). Therefore, only newly synthesized PrPSc was detected and discriminated from PrPSc in inocula. Inoculated cells were lysed at various time points postinfection and analyzed both for newly converted PrPSc and LC3-II levels. Upon induction of autophagy, posttranslationally processed LC3 (LC3-I) is converted into LC3-II. An increase in the level of LC3-II is commonly used as marker for autophagy induction, as the amount of LC3-II associated with autophagosome membranes correlates with the extent of autophagosome formation (Klionsky et al. 2008). In comparison to mock-brain infection, increased amounts of LC3-II were detected in prion-susceptible cell populations upon prion inoculation. This phenomenon was observed concomitant with the ability of cells to propagate PrPSc in detectable levels. When primary prion infection manifested in cells, the increased level of LC3-II went back to levels as observed in controls. Similar results were found for the medium susceptible clone, which started propagating PrPSc at a later time point. This phenomenon was lacking in prion-unsusceptible cells, indicating that autophagosome formation is transiently induced only in cells actively propagating PrPSc and is not the result of a cellular response to PrPSc in inocula.
We then wanted to further analyze whether basal levels of autophagy indeed play a role in primary prion infection. We inoculated wild-type mouse embryonic fibroblasts (MEFwt) and autophagy-deficient MEFs (MEFATG5−/−), originating from ATG5−/− transgenic mice, with prion-infected brain homogenate. Since these cells did not contain a 3F4-tagged PrP, we waited until day 20 postinfection to be sure that we do not detect PrPSc from inocula. Interestingly, MEFATG5−/− cells showed only very weak amounts of PrPSc at days 20 and 30 p.i., whereas MEFwt cells propagated PrPSc very efficiently. In addition, when we used siRNA-targeting Atg5 or beclin-1 genes, both genes necessary for execution of autophagy, around the time of primary prion infection of neuronal cells, we also observed a reduction in PrPSc propagation compared to mock-treated cells. This data indicated that the absence of autophagy very strongly decreased the cellular susceptibility to primary prion infection and that autophagy potentially also plays a role in maintenance of productive prion infection over time.
To rule out the possibility that the difference in prion susceptibility between Atg5−/− and +/+ cells was not based on ATG5 alone and might depend on cell clone issues, we decided to reintroduce the Atg5 gene into ATG5−/− cells. MEFATG5−/− cells were stably transduced with a lentivirus construct-encoding ATG5 to restore autophagy competence (cells termed MEFATG5). As done before, autophagy-competent and autophagy-deficient cells were inoculated with 22 L prion- or mock-infected brain homogenates in parallel experiments. Reintroduction of autophagy competence in MEFATG5 cells provoked a clearly increased susceptibility to primary prion infection as compared to autophagy-deficient counterparts. Interestingly, analysis of prion-infected cells at a later time point postinfection (55 days postinfection, dpi) revealed that the reduced PrPSc level in autophagy-deficient cells is not a transient phenomenon and that lack of autophagy may even result in abrogation of cellular prion infection. In contrast, autophagy-competent cells efficiently propagated PrPSc at 55 dpi, reflecting persistent prion infection. We also quantitatively addressed this phenomenon. We stepwise increased autophagy competence by transducing MEFATG5−/− cells with different dilutions of ATG5-encoding virus. In contrast to autophagy-deficient cells, gradually restored autophagy competence resulted in accordingly elevated levels in prion infection and PrPSc propagation, both after 20 and 30 dpi. These results indicate that restored autophagy competence renders cells more prone to PrPSc propagation, validating a pivotal role of functional active basal autophagy in primary and persistent prion infection.
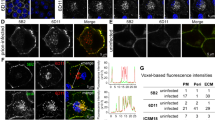
We hypothesize that autophagy plays a general role in the subcellular recycling of prions. The previous view that PrPSc is generated along the early endocytic pathway and is unidirectional transported to lysosomes for final degradation is not compatible anymore with recent findings (Beranger et al. 2002; Marijanovic et al. 2009; Gilch et al. 2009; Yamasaki et al. 2012). Without obtaining new prions steadily from the outside of the cell, such a unidirectional mechanism is rather incompatible with persistent prion propagation in terminally differentiated cells. It is likely that a fraction of PrPSc is retro-transported to the more upstream locale of prion conversion, thereby allowing continuous flow of prion generation (see Fig. 11.2). Experimental evidence for such a scenario was reported by our and the Lehmann, Zurzolo, and Horiuchi groups (Beranger et al. 2002; Marijanovic et al. 2009; Gilch et al. 2009; Yamasaki et al. 2012). We postulate that autophagic flux mechanisms play a role in this scenario; we even hypothesize that the level of autophagic flux represents a crossing point which decides whether PrPSc gets recycled or degraded. Increasing the autophagic flux counteracts the recycling pathway and thereby also affects prion conversion, taking away template for conversion and inducing its degradation (Fig. 11.2). We postulate that autophagic compartments provide disaggregase function and increase seeds as needed or are at least supportive for efficient prion propagation. When late endosomes containing PrPSc aggregates fuse with autophagosomes, the autophagic machinery and the cellular locale containing PrPSc/prions physically meet and get interconnected. How the endosomal recycling compartment (ESCRT) machinery is involved in completion of autophagy is presently subject of intensive research (Raiborg and Stenmark 2009; Rusten and Stenmark 2009; Rusten and Simonsen 2008). A general view is that the ESCRT machinery is required in fusion of autophagosomes with endosomes and lysosomes (Raiborg and Stenmark 2009; Rusten and Stenmark 2009). Two rab proteins have been found involved in this process (Rab11 and Rab7) (Rusten and Simonsen 2008; Fader and Colombo 2009). Interestingly, work from the Zurzolo laboratory has identified the ERC as a likely site of prion conversion (Marijanovic et al. 2009).
Model for how the level of autophagy impacts prion infection. Exogenously elevated levels of autophagy (e.g., by chemical compounds) affect prion clearance and thereby prion infection. When the autophagic flux increases more autophagosomes fuse with late endosomes (LL, in some cell types also known as multivesicular bodies, MVBs) which contain PrPSc/prions to form amphisomes (not shown). This results in increased fusion to lysosomes (Ly) and steadily sequesters the template for prion conversion. At the same time, the increase in prion clearance negatively affects the role of autophagy in prion propagation (left side). We hypothesize that PrPSc/prions recycle from early endosomes (EE) directly or indirectly (via LE/TGN) to the ERC (endosomal recycling complex) and from there back towards the cellular compartment of prion conversion. When this goes over LE, the autophagic machinery gets connected with compartments recycling PrPSc/prions and can provide disaggregation activity to them. Based on our new findings and data as published by us and others (Beranger et al. 2002; Marijanovic et al. 2009; Gilch et al. 2009; Yamasaki et al. 2011)
Is such a dual function of autophagy conceivable? It is known that basal autophagy and moderately enhanced autophagy help in cell survival but that both impairment of autophagy or strongly enhanced autophagy can lead to cell death (Martinez-Vicente and Cuervo 2007; Ventruti and Cuervo 2007; Ertmer et al. 2004; Cuervo et al. 2010; Wong and Cuervo 2010). We postulate that components of the autophagic flux might represent the biological equivalent for the postulated disaggregase activity in mammalian prion and prion-like biology. Whereas PMCA uses physical disintegration, for yeast prions this is done by Hsp104. Interestingly, Hsp104 also has a dual role, and depending on the level of activity, it is involved in aggregation and disaggregation (Shorter and Lindquist 2004). Having a system in hand which provides PrPSc originating from cells with normal and impaired autophagy, we can study now how this affects molecular, biophysical, and infectivity features of prions.
11.2 Effect of Proteasome Dysfunction and ER Stress on PrPSc Biogenesis
We recently found that proteasomal impairment and ER stress can have a direct impact on the level and quality of PrPc in the secretory pathway and its “fitness” for substrate in prion conversion. This finding is different from previous findings which focused on PrP moieties in the cytosol (Ma et al. 2002; Ma and Lindquist 2001, 2002; Kristiansen et al. 2005, 2007; Cohen and Taraboulos 2003). Our data suggest a novel pathway which contributes to “conventional” prion propagation. Such conversion favoring or disfavoring cellular conditions might also be of relevance for the pathogenesis of sporadic CJD, where initial conversion might take place without a bona fide PrPSc template. Improving protein quality in ER and post-ER compartments in trans in order to generate PrPc populations which have a more stable conformation and/or are less efficiently converted into PrPSc might provide translational potential (Nunziante et al. 2011).
Proteasomal dysfunction and ER stress enhance trafficking of prion protein aggregates through the secretory pathway and increase PrPSc.
In infectious forms of prion diseases, a direct interaction between PrPSc template and PrPc substrate underlies the conformational change of PrPc into PrPSc (Prusiner 1998). It is assumed that a preceding plasma membrane localization of PrPc is mandatory for conversion into PrPSc (Prusiner 2001; Caughey and Raymond 1991; Caughey et al. 1998; Borchelt et al. 1992). Much less is known in this respect about events occurring in ER and in early secretory compartments. The cellular mechanisms underlying sporadic prion diseases are mostly unknown and are difficult to assess in experimental systems. Various models propose the existence of a PrP isoform which is more prone to conversion into PrPSc (Billeter et al. 1997; Glockshuber 2001; Hornemann and Glockshuber 1998). The fundamental role of the ER environment and of the ER-associated degradation pathway (ERAD) in metabolism and turnover of wild-type and some mutant prion proteins has been highlighted in the past with regard to implications for prion diseases (Rogers et al. 1990; Yedidia et al. 2001; Lorenz et al. 2002; Drisaldi et al. 2003). Whereas work done by other groups mainly focused on aberrant PrP moieties in the cytosol or in aggresomes and its possible impact in execution of neurodegeneration (Ma et al. 2002; Ma and Lindquist 2002; Kristiansen et al. 2005, 2007; Yedidia et al. 2001), the aim of our study was to investigate how perturbations of ER homeostasis or proteasomal impairment affect PrPc metabolism in the secretory pathway and thereby directly PrPSc biogenesis.
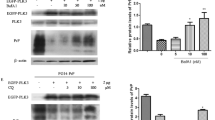
We found that induction of ER stress resulted in a general attenuation of PrPc level (Nunziante et al. 2011). In addition, we found aggregated PrP species that localized mainly in secretory compartments and at the cell surface. Inhibition of proteasomal function led to a significant increase of the total PrPc level and to accumulation of detergent soluble and insoluble PrPc isoforms. PrP species detected under these conditions were fully glycosylated, were properly processed through the secretory pathway, and localized at the outer leaflet of the plasma membrane. This was the case in cells with endogenous PrP expression, in primary neurons as well as in PrP-transfected cells. The majority of studies conducted on proteasomal degradation of PrP describe cytosolic accumulation of toxic PrP aggregates upon inhibition of this pathway. Although not extensively investigated for PrP metabolism before, it was assumed for other proteins that ER and quality control compartments are connected to the secretory pathway. In our hands, experimental manipulation of both pathways led to accumulation of insoluble PrP species in the secretory pathway, but the events underlying their formation seemed to be different, as were the effects on PrPc localization and expression.
Strikingly, inhibition of proteasomal activity amplified PrPSc levels in persistently prion-infected cells. The direct correlation between proteasome and PrPSc accumulation within cells represents a new aspect in prion metabolism. Previous studies reported formation of cytoplasmic PrPSc aggregates which associated with aggresomes and led to apoptotic death in prion-infected neurons, but only after mild inhibition of the proteasome (Kristiansen et al. 2005, 2007). In addition, purified PrPSc preparations were seen to inhibit the proteolytic activity of the proteasome (Kristiansen et al. 2007). These data support the view of a cytosolic localization for portions of PrPSc either by retro-translocation or by endosomal–lysosomal membrane destabilization (Laszlo et al. 1992). In our study, upon proteasomal inhibition, PrPc and detergent-insoluble aggregates were extensively transported to the cell surface, one of the putative sites for prion formation. Such PrP molecules might represent additional substrate binding to existing PrPSc seeds and leading to the increased formation of PrPSc as detected in our study.
We further underlined the fundamental role of the early secretory pathway in folding and transport of PrPc with respect to prion formation by overexpressing molecules known to promote cellular quality. Overexpression of EDEM-3 or ERGIC-53 significantly reduced PrP aggregates and PrPSc in infected cells. EDEM proteins are ER-resident lectins which recognize N-linked glycans on aberrantly folded proteins, accelerate their release from the calnexin/calreticulin cycle, and sort them for ERAD-degradation (Molinari et al. 2003; Oda et al. 2003; Ruddock and Molinari 2006). It is therefore plausible that by enhancing ERAD-degradation of PrP aggregates, EDEM-3 subtracts the substrate necessary for prion conversion. A similar explanation for reduction of PrPSc could apply to ERGIC-53, which selectively transports functionally folded proteins from ER to ERGIC vesicles and also operates in the quality control of glycoproteins (Appenzeller et al. 1999). ERGIC-53 might therefore promote proper folding of PrPc and selectively transport this cargo to the cell surface. This PrPc population would have a more stable conformation and/or be less efficiently converted into PrPSc.
Taken together, our data support the notion that ER and cellular quality control mechanisms tightly modulate PrP maturation and PrPSc formation. We show that proteasomal degradation and ERAD play a physiological role for endogenous PrPc in the secretory pathway. Impairments in this pathway as well as disturbances in ER homeostasis cause accumulation of PrP aggregates which are increasingly recycled through the secretory pathway, resulting in enhanced PrPSc replication. Of note, such conversion favoring or disfavoring cellular conditions might also be of relevance for the pathogenesis of sporadic CJD, where initial conversion might take place without a bona fide PrPSc template.
References
Aguib Y, Heiseke A, Gilch S, Riemer C, Baier M, Schatzl HM, Ertmer A (2009) Autophagy induction by trehalose counteracts cellular prion infection. Autophagy 5:361–369
Appenzeller C, Andersson H, Kappeler F, Hauri HP (1999) The lectin ERGIC-53 is a cargo transport receptor for glycoproteins. Nat Cell Biol 1:330–334
Batlevi Y, La Spada AR (2011) Mitochondrial autophagy in neural function, neurodegenerative disease, neuron cell death, and aging. Neurobiol Dis 43:46–51
Beranger F, Mange A, Goud B, Lehmann S (2002) Stimulation of PrP(C) retrograde transport toward the endoplasmic reticulum increases accumulation of PrP(Sc) in prion-infected cells. J Biol Chem 277:38972–38977
Billeter M, Riek R, Wider G, Hornemann S, Glockshuber R, Wuthrich K (1997) Prion protein NMR structure and species barrier for prion diseases. Proc Natl Acad Sci USA 94:7281–7285
Boellaard JW, Kao M, Schlote W, Diringer H (1991) Neuronal autophagy in experimental scrapie. Acta Neuropathol 82:225–228
Borchelt DR, Taraboulos A, Prusiner SB (1992) Evidence for synthesis of scrapie prion proteins in the endocytic pathway. J Biol Chem 267:16188–16199
Borchsenius AS, Muller S, Newnam GP, Inge-Vechtomov SG, Chernoff YO (2006) Prion variant maintained only at high levels of the Hsp104 disaggregase. Curr Genet 49:21–29
Brundin P, Melki R, Kopito R (2010) Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat Rev Mol Cell Biol 11:301–307
Castilla J, Saa P, Hetz C, Soto C (2005) In vitro generation of infectious scrapie prions. Cell 121:195–206
Caughey B, Raymond GJ (1991) The scrapie-associated form of PrP is made from a cell surface precursor that is both protease- and phospholipase-sensitive. J Biol Chem 266:18217–18223
Caughey B, Race RE, Ernst D, Buchmeier MJ, Chesebro B (1998) Prion protein biosynthesis in scrapie-infected and uninfected neuroblastoma cells. J Virol 63:175–181
Chernoff YO, Lindquist SL, Ono B, Inge-Vechtomov SG, Liebman SW (1995) Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [psi+]. Science 268:880–884
Cherra SJ III, Dagda RK, Chu CT (2010) Review: autophagy and neurodegeneration: survival at a cost? Neuropathol Appl Neurobiol 36:125–132
Chew KC, Ang ET, Tai YK, Tsang F, Lo SQ, Ong E, Ong WY, Shen HM, Lim KL, Dawson VL, Dawson TM, Soong TW (2011) Enhanced autophagy from chronic toxicity of iron and mutant A53T alpha-synuclein: implications for neuronal cell death in Parkinson disease. J Biol Chem 286:33380–33389
Chu CT, Plowey ED, Dagda RK, Hickey RW, Cherra SJ III, Clark RS (2009) Autophagy in neurite injury and neurodegeneration: in vitro and in vivo models. Methods Enzymol 453:217–249
Cohen E, Taraboulos A (2003) Scrapie-like prion protein accumulates in aggresomes of cyclosporin A-treated cells. EMBO J 22:404–417
Cuervo AM, Wong ES, Martinez-Vicente M (2010) Protein degradation, aggregation, and misfolding. Mov Disord 25(Suppl 1):S49–S54
Drisaldi B, Stewart RS, Adles C, Stewart LR, Quaglio E, Biasini E, Fioriti L, Chiesa R, Harris DA (2003) Mutant PrP is delayed in its exit from the endoplasmic reticulum, but neither wild-type nor mutant PrP undergoes retrotranslocation prior to proteasomal degradation. J Biol Chem 278:21732–21743
Ertmer A, Gilch S, Yun SW, Flechsig E, Klebl B, Stein-Gerlach M, Klein MA, Schatzl HM (2004) The tyrosine kinase inhibitor STI571 induces cellular clearance of PrPSc in prion-infected cells. J Biol Chem 279:41918–41927
Ertmer A, Huber V, Gilch S, Yoshimori T, Erfle V, Duyster J, Elsasser HP, Schatzl HM (2007) The anticancer drug imatinib induces cellular autophagy. Leukemia 21:936–942
Fader CM, Colombo MI (2009) Autophagy and multivesicular bodies: two closely related partners. Cell Death Differ 16:70–78
Gilch S, Schatzl HM (2003) Promising developments bringing prion diseases closer to therapy and prophylaxis. Trends Mol Med 9:367–369
Gilch S, Nunziante M, Ertmer A, Schatzl HM (2007) Strategies for eliminating PrP(c) as substrate for prion conversion and for enhancing PrP(Sc) degradation. Vet Microbiol 123:377–386
Gilch S, Krammer C, Schatzl HM (2008) Targeting prion proteins in neurodegenerative disease. Expert Opin Biol Ther 8:923–940
Gilch S, Bach C, Lutzny G, Vorberg I, Schatzl HM (2009) Inhibition of cholesterol recycling impairs cellular PrP(Sc) propagation. Cell Mol Life Sci 66:3979–3991
Glockshuber R (2001) Folding dynamics and energetics of recombinant prion proteins. Adv Protein Chem 57:83–105
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, Yokoyama M, Mishima K, Saito I, Okano H, Mizushima N (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441:885–889
Heel RC, Brogden RN, Speight TM, Avery GS (1978) Tamoxifen - review of its pharmacological properties and therapeutic use in treatment of breast-cancer. Drugs 16:1–24
Heiseke A, Aguib Y, Riemer C, Baier M, Schatzl HM (2009) Lithium induces clearance of protease resistant prion protein in prion-infected cells by induction of autophagy. J Neurochem 109:25–34
Heiseke A, Aguib Y, Schatzl HM (2010) Autophagy, prion infection and their mutual interactions. Curr Issues Mol Biol 12:87–98
Hornemann S, Glockshuber R (1998) A scrapie-like unfolding intermediate of the prion protein domain PrP(121–231) induced by acidic pH. Proc Natl Acad Sci USA 95:6010–6014
Jeong H, Then F, Melia TJ, Mazzulli JR, Cui L, Savas JN, Voisine C, Paganetti P, Tanese N, Hart AC, Yamamoto A, Krainc D (2009) Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 137:60–72
Kaganovich D, Kopito R, Frydman J (2008) Misfolded proteins partition between two distinct quality control compartments. Nature 454:1088–1095
Klionsky DJ (2007) Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 8:931–937
Klionsky DJ et al (2008) Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy 4:151–175
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, Ueno T, Koike M, Uchiyama Y, Kominami E, Tanaka K (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441:880–884
Korolchuk VI, Rubinsztein DC (2011) Regulation of autophagy by lysosomal positioning. Autophagy 7:927–928
Krainc D (2010) Clearance of mutant proteins as a therapeutic target in neurodegenerative diseases. Arch Neurol 67:388–392
Krammer C, Vorberg I, Schatzl HM, Gilch S (2009a) Therapy in prion diseases: from molecular and cellular biology to therapeutic targets. Infect Disord Drug Targets 9:3–14
Krammer C, Kryndushkin D, Suhre MH, Kremmer E, Hofmann A, Pfeifer A, Scheibel T, Wickner RB, Schatzl HM, Vorberg I (2009b) The yeast Sup35NM domain propagates as a prion in mammalian cells. Proc Natl Acad Sci USA 106:462–467
Kristiansen M, Messenger MJ, Klohn PC, Brandner S, Wadsworth JD, Collinge J, Tabrizi SJ (2005) Disease-related prion protein forms aggresomes in neuronal cells leading to caspase activation and apoptosis. J Biol Chem 280:38851–38861
Kristiansen M, Deriziotis P, Dimcheff DE, Jackson GS, Ovaa H, Naumann H, Clarke AR, van Leeuwen FW, Menendez-Benito V, Dantuma NP, Portis JL, Collinge J, Tabrizi SJ (2007) Disease-associated prion protein oligomers inhibit the 26S proteasome. Mol Cell 26:175–188
Laszlo L, Lowe J, Self T, Kenward N, Landon M, McBride T, Farquhar C, McConnell I, Brown J, Hope J (1992) Lysosomes as key organelles in the pathogenesis of prion encephalopathies. J Pathol 166:333–341
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132:27–42
Liberski PP, Sikorska B, Bratosiewicz-Wasik J, Gajdusek DC, Brown P (2004) Neuronal cell death in transmissible spongiform encephalopathies (prion diseases) revisited: from apoptosis to autophagy. Int J Biochem Cell Biol 36:2473–2490
Liberski PP, Brown DR, Sikorska B, Caughey B, Brown P (2008) Cell death and autophagy in prion diseases (transmissible spongiform encephalopathies). Folia Neuropathol 46:1–25
Lindquist S, Patino MM, Chernoff YO, Kowal AS, Singer MA, Liebman SW, Lee KH, Blake T (1995) The role of Hsp104 in stress tolerance and [PSI+] propagation in Saccharomyces cerevisiae. Cold Spring Harb Symp Quant Biol 60:451–460
Lorenz H, Windl O, Kretzschmar HA (2002) Cellular phenotyping of secretory and nuclear prion proteins associated with inherited prion diseases. J Biol Chem 277:8508–8516
Ma J, Lindquist S (2001) Wild-type PrP and a mutant associated with prion disease are subject to retrograde transport and proteasome degradation. Proc Natl Acad Sci USA 98:14955–14960
Ma J, Lindquist S (2002) Conversion of PrP to a self-perpetuating PrPSc-like conformation in the cytosol. Science 298:1785–1788
Ma J, Wollmann R, Lindquist S (2002) Neurotoxicity and neurodegeneration when PrP accumulates in the cytosol. Science 298:1781–1785
Maas E, Geissen M, Groschup MH, Rost R, Onodera T, Schatzl H, Vorberg IM (2007) Scrapie infection of prion protein-deficient cell line upon ectopic expression of mutant prion proteins. J Biol Chem 282:18702–18710
Marijanovic Z, Caputo A, Campana V, Zurzolo C (2009) Identification of an intracellular site of prion conversion. PLoS Pathog 5:e1000426
Martinez-Vicente M, Cuervo AM (2007) Autophagy and neurodegeneration: when the cleaning crew goes on strike. Lancet Neurol 6:352–361
Meijer AJ, Codogno P (2004) Regulation and role of autophagy in mammalian cells. Int J Biochem Cell Biol 36:2445–2462
Mizushima N (2009) Regulation of autophagosome formation in mammalian cells. Autophagy 5:898–899
Mizushima N (2011) Autophagy in protein and organelle turnover. Cold Spring Harb Symp Quant Biol 76:397–402
Mizushima N, Yamamoto A, Matsui M, Yoshimori T, Ohsumi Y (2004) In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell 15:1101–1111
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451:1069–1075
Molinari M, Calanca V, Galli C, Lucca P, Paganetti P (2003) Role of EDEM in the release of misfolded glycoproteins from the calnexin cycle. Science 299:1397–1400
Nixon RA (2005) Endosome function and dysfunction in Alzheimer’s disease and other neurodegenerative diseases. Neurobiol Aging 26:373–382
Nunziante M, Ackermann K, Dietrich K, Wolf H, Gadtke L, Gilch S, Vorberg I, Groschup M, Schatzl HM (2011) Proteasomal dysfunction and endoplasmic reticulum stress enhance trafficking of prion protein aggregates through the secretory pathway and increase accumulation of pathologic prion protein. J Biol Chem 286:33942–33953
Oda Y, Hosokawa N, Wada I, Nagata K (2003) EDEM as an acceptor of terminally misfolded glycoproteins released from calnexin. Science 299:1394–1397
Prusiner SB (1998) Prions. Proc Natl Acad Sci USA 95:13363–13383
Prusiner SB (2001) Shattuck lecture–neurodegenerative diseases and prions. N Engl J Med 344:1516–1526
Raiborg C, Stenmark H (2009) The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 458:445–452
Ren PH, Lauckner JE, Kachirskaia I, Heuser JE, Melki R, Kopito RR (2009) Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine aggregates. Nat Cell Biol 11:219–225
Rogers M, Taraboulos A, Scott M, Groth D, Prusiner SB (1990) Intracellular accumulation of the cellular prion protein after mutagenesis of its Asn-linked glycosylation sites. Glycobiology 1:101–109
Rubinsztein DC (2006) The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 443:780–786
Rubinsztein DC, DiFiglia M, Heintz N, Nixon RA, Qin ZH, Ravikumar B, Stefanis L, Tolkovsky A (2005) Autophagy and its possible roles in nervous system diseases, damage and repair. Autophagy 1:11–22
Ruddock LW, Molinari M (2006) N-glycan processing in ER quality control. J Cell Sci 119:4373–4380
Rusten TE, Simonsen A (2008) ESCRT functions in autophagy and associated disease. Cell Cycle 7:1166–1172
Rusten TE, Stenmark H (2009) How do ESCRT proteins control autophagy? J Cell Sci 122:2179–2183
Sarkar S, Rubinsztein DC (2008) Small molecule enhancers of autophagy for neurodegenerative diseases. Mol Biosyst 4:895–901
Sarkar S, Davies JE, Huang ZB, Tunnacliffe A, Rubinsztein DC (2007) Trehalose, a novel mTOR-independent autophagy enhancer, accelerates the clearance of mutant huntingtin and alpha-synuclein. J Biol Chem 282:5641–5652
Schatzl HM, Laszlo L, Holtzman DM, Tatzelt J, Dearmond SJ, Weiner RI, Mobley WC, Prusiner SB (1997) A hypothalamic neuronal cell line persistently infected with scrapie prions exhibits apoptosis. J Virol 71:8821–8831
Shorter J (2008) Hsp104: a weapon to combat diverse neurodegenerative disorders. Neurosignals 16:63–74
Shorter J (2011) The mammalian disaggregase machinery: hsp110 synergizes with hsp70 and hsp40 to catalyze protein disaggregation and reactivation in a cell-free system. PLoS One 6:e26319
Shorter J, Lindquist S (2004) Hsp104 catalyzes formation and elimination of self-replicating Sup35 prion conformers. Science 304:1793–1797
Sikorska B, Liberski PP, Giraud P, Kopp N, Brown P (2004) Autophagy is a part of ultrastructural synaptic pathology in Creutzfeldt–Jakob disease: a brain biopsy study. Int J Biochem Cell Biol 36:2563–2573
Ventruti A, Cuervo AM (2007) Autophagy and neurodegeneration. Curr Neurol Neurosci Rep 7:443–451
Vives-Bauza C, Zhou C, Huang Y, Cui M, de Vries RL, Kim J, May J, Tocilescu MA, Liu W, Ko HS, Magrane J, Moore DJ, Dawson VL, Grailhe R, Dawson TM, Li C, Tieu K, Przedborski S (2010) PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc Natl Acad Sci USA 107:378–383
Winslow AR, Rubinsztein DC (2008) Autophagy in neurodegeneration and development. Biochim Biophys Acta 1782:723–729
Wong E, Cuervo AM (2010) Autophagy gone awry in neurodegenerative diseases. Nat Neurosci 13:805–811
Yamasaki T, Suzuki A, Shimizu T, Watarai M, Hasebe R, Horiuchi M (2012) Characterization of intracellular localization of PrPSc in prion-infected cells using monoclonal antibody that recognizes the region consisting of amino acids 119–127 of mouse PrP. J Gen Virol 93(Pt 3):668–80
Yedidia Y, Horonchik L, Tzaban S, Yanai A, Taraboulos A (2001) Proteasomes and ubiquitin are involved in the turnover of the wild-type prion protein. EMBO J 20:5383–5391
Yun SW, Ertmer A, Flechsig E, Gilch S, Riederer P, Gerlach M, Schatzl HM, Klein MA (2007) The tyrosine kinase inhibitor imatinib mesylate delays prion neuroinvasion by inhibiting prion propagation in the periphery. J Neurovirol 13:328–337
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Schatzl, H.M. (2013). Cellular Mechanisms of Propagation and Clearance. In: Zou, WQ., Gambetti, P. (eds) Prions and Diseases. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-5305-5_11
Download citation
DOI: https://doi.org/10.1007/978-1-4614-5305-5_11
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-5304-8
Online ISBN: 978-1-4614-5305-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)