Abstract
While the human GW182 gene was discovered over 10 years ago, functional characterization of the Drosophila melanogaster GW182 othologue—Gawky (Gw, previously denoted as CG31992, CG11484, CG9905, or dGW182) has been relatively recent. (Rehwinkel et al. 2005; Schneider et al. 2006) However, the Drosophila model has contributed greatly to studying the role(s) of the GW182 family proteins in multiple pathways and in particular their role in RNA interference (RNAi). Of the commonly used metazoan models, Drosophila is unique in that there is only one Gw protein encoded by the Drosophila genome and this homologue retains a high level of sequence and/or organizational identity to vertebrate GW182 proteins (Fig. 8.1). Thus, the potential functional redundancy associated with the multiple GW182 family proteins encoded by the mammalian genome is less of a concern in Drosophila studies (Schneider et al. 2006; Eystathioy et al. 2002). The bulk of the currently published literature regarding Drosophila Gw can be divided into two main categories. Functional studies describing the Drosophila gw mutant phenotype and cell-biological/biochemical studies probing the vital role of Gw in the mechanics of Drosophila miRNA pathway.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
8.1 Introduction
While the human GW182 gene was discovered over 10 years Ago, functional characterization of the Drosophila melanogaster GW182 othologue—Gawky (GW, previously denoted as CG31992, CG11484, CG9905, or dGW182) has been relatively recent. (Rehwinkel et al. 2005; Schneider et al. 2006) However, the Drosophila model has contributed greatly to studying the role(s) of the GW182 family proteins in multiple pathways and in particular their role in RNA interference (RNAi). Of the commonly used metazoan models, Drosophila is unique in that there is only one Gw protein encoded by the Drosophila genome and this homologue retains a high level of sequence and/or organizational identity to vertebrate GW182 proteins (Fig. 8.1). Thus, the potential functional redundancy associated with the multiple GW182 family proteins encoded by the mammalian genome is less of a concern in Drosophila studies (Schneider et al. 2006; Eystathioy et al. 2002). The bulk of the currently published literature regarding Drosophila Gw can be divided into two main categories. Functional studies describing the Drosophila gw mutant phenotype and cell-biological/biochemical studies probing the vital role of Gw in the mechanics of Drosophila miRNA pathway.
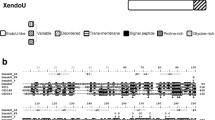
The structural organization of the Drosophila Gw protein. MI MII MIII—Motif I, highly conserved regions—Motif II and Motif III (Eulalio et al. 2009a). DUF conserved domain of unknown function; UBA ubiquitin associated domain; Q/QN Rich glutamine/glutamine and asparagine rich domain; RRM RNA recognition Motif; Ser serine rich domain
8.2 Drosophila Cells Make Extensive Use of Cytoplasmic mRNA Regulation
In eukaryotic cells, cytoplasmic mRNA regulation is thought to occur largely within ribonucleoprotein (RNP) complexes. These complexes contain both RNAs and proteins and are often aggregated into larger regulatory structures (Zhang et al. 2003; Muddashetty et al. 2002; Ohashi et al. 2000; Kobayashi et al. 1998). Many investigations of these regulatory mechanisms in Drosophila pre-date the discovery of the RNA interference (RNAi) pathway or the functional characterization of gw. One of the first RNP structures to be described is the Nuage/Polar granules located within Drosophila oocytes and embryos, where regulation of genes critical patterning and development makes extensive use of localized expression of mRNAs at the cellular poles (Hay et al. 1988; Wilsch-Brauninger et al. 1997).
Many other patterning events within the developing Drosophila oocyte and fertilized embryos are also extensively regulated by post-translational gene regulation. Drosophila screens to identify genes involved in developmental processes have identified several genes encoding multiple components of regulatory RNPs. Examples of these include: Staufen (STAU); Exupurentia (Exu); Ypsilon schachtel (YPS), a Y box binding protein One homologue and Oo18 RNA-binding protein (Orb), the Drosophila Cytoplasmic Poly (A) Element Binding protein homologue (St Johnston et al. 1991; Mansfield et al. 2002; Lin et al. 2006). Notably, in Drosophila, it seems that multiple mRNA regulatory events can be functionally linked. For example, there is appears to be a coupling of translational suppression and cytoplasmic mRNA localization and/or transport in Drosophila embryos. Me31B, a DEAD box helicase translational repressor and decapping activator, transiently localizes with RNP granules during transport, until they reach the posterior of the oocytes (Lin et al. 2006). Notably, many of these previously characterized RNA regulatory proteins have since been associated with GW or mammalian GW182 (Eulalio et al. 2007b; Ikeda et al. 2006; Quaresma et al. 2009; Huntzinger et al. 2010; Tritschler et al. 2010; Yao et al. 2011).
This functional linkage between multiple aspects of mRNA regulation and Recently, Dcp1, a key part of the decapping enzyme complex that is often found associated with GW182 family of proteins in mRNA processing (P-)bodies was also identified as a component of RNP granules that localize to the posterior of the oocyte (Lin et al. 2006). Degradation of some posteriorly localized transcripts does occur during early embryogenesis (Ephrussi et al. 1991; Kim-Ha et al. 1993). One possibility is that early recruitment of Dcp1 may facilitate the rapid assembly of the degradation machinery at a later time during embryo development (Lin et al. 2006). Much of the early mRNA deposited in the embryo maternally is co-ordinately degraded at approximately 120 min after eggs are deposited at the mid-zygotic (or mid-blastula) transition (reviewed in (Tadros and Lipshitz 2009)). At this stage of development, foci are seen within the embryo that have a typical P-body-like composition including Dcp2 and the 5′-3′ exonuclease Pacman (Pcm), a homologue of human Xrn1 (Lin et al. 2008). The fact that the various different regulatory RNPs active in Drosophila cells often share many of the same protein components between various regulatory structures supports that they may be linked functionally. This might mean that RNPs containing translationally repressed and localized mRNAs that are initially formed in the oocyte may later acquire additional components to degrade these mRNAs when they are no longer needed.
Extensive regulation of mRNA within cytoplasmic RNPs is not limited to Drosophila embryogenesis. Drosophila neurons also contain cytoplasmic RNPs that include factors involved in P-body mediated mRNA decay including Pcm, Dcp1, Ago2 the RNAi component, and Up-frameshift suppressor (Upf), a component of RNA nonsense mediated decay (NMD) pathway (Metzstein and Krasnow 2006). These neuronal RNPs also have been reported to share components with maternal mRNA regulatory RNPs including STAU, FRMP and Barentz or protein components normally localized to stress granules (G3BP and eIF2) (Barbee et al. 2006). Of particular note is the observation that a number of RNPs contained different subsets of these components. Additionally, the composition of RNPs in neuronal cells appears to be influenced directly by the relative level of particular protein components. Over-expression of Stau or a GFP fusion of dFMR1 resulted in an increase in the degree of co-localization of these two proteins in cytoplasmic RNPs. This concurrent increase in particle size and decrease in particle number suggests that the increase in co-localization may be the result of fusion of different types of RNA granules. Fusion of these granules further supports a model where there is a functional relationship between them. Thus, while many functionally diverse mRNA regulatory bodies have been discovered independently in various Drosophila cell types, they share significant similarities, both in composition and function. Thus, there is a distinct possibility that our current differentiation of cytoplasmic RNA regulatory bodies in Drosophila could be largely artificial or that there is significant cross-talk between different aspects of mRNA regulation. However, given that most of these proteins were identified in functional screens affecting specific aspects of Drosophila development, it is clear these cytoplasmic RNPs have a direct role in regulating many different aspects of cellular function balancing competing cytoplasmic events: mRNA translation and sequestering/degrading mRNAs in RNP complexes. Elucidating the role of Drosophila Gw in some or all of these various aspects of mRNA regulation during initial cellular differentiation and later homeostasis has only just begun.
8.3 The Drosophila Genome Project Predicted Only one Gene Similar to GW182
The majority of the Drosophila genome was first sequenced in 2000 (Adams et al. 2000; Stapleton et al. 2002; Drysdale 2003). Drosophila sequences similar to human GW182 were first identified as two different genes on chromosome 4 given the sequential identifiers (Celera Genomics) CG11484, CG9905 (Adams et al. 2000). Later efforts focused on both functional annotation of the genome and conformation of the mRNAs expressed from each identified gene refined this prediction to a single gene id CG31992, which was subsequently named gawky (gw) (Schneider et al. 2006). Follow up projects that mass sequenced multiple cDNAs indicated that the gw locus produces 8 transcripts via alternative splicing: gw-RA, gw-RB, gw-RC, gw-RD, gw-RE, gw-RF, gw-RG and gw-RH (Fig. 8.2). However, all of these transcripts differ only in their 5′ untranlsated region (UTR) and the open reading frame of each of these alternative splicing forms is identical, encoding a protein with a molecular weight 143 kD. The modENCODE project has confirmed that Drosophila gw expression is seen in all development stages. The relative expression levels of gw are higher during early embryogenesis and at the beginning stages of pupariation, implying that during these two stages, cells may have elevated requirements for Gw (Celniker et al. 2009). Finally, the Drosophila genome project has further sequenced gw homologues from multiple related species and have found that there is significant conservation of the gw locus among the Drosophilids (Gilbert 2007).
The gw gene produces eight mRNA isoforms differing only at the 5′ untranslated region (UTR). The 5′ UTR (blue) is composed of several different exons that are selectively spliced and expressed from three different start sites. However, the coding region of the Gw protein (red ) and the 3′ UTR (green) are the same in each gw mRNA isoform. The numbered boxes indicate the positions of RT-PCR primers that can be used to identify mRNAs transcribed from each of the three alternative start-sites and, based upon the length of the resulting product, each of the mRNAs
8.4 Functional Identification of a the Gawky (gw) Mutation
Traditionally, gene discovery in Drosophila focuses on the identification of gene mutations affecting specific cellular or developmental activities (St Johnston 2002). Despite an extensive history of screening of the Drosophila genome for mutations affecting embryo development, the identification of gw as a critical gene required for early embryonic development was quite recent (Schneider et al. 2006). The likely reason for this is that the gw gene is located on the right arm of chromosome 4 at sequence location 4:670575..682391, cytological map location 102D2-102D3. Unfortunately, the large scale screens for mutations that are so effective in isolating critical Drosophila genes on other chromosomes largely ignore the few genes on chromosome IV. Drosophila has two sex chromosomes and 3 autosomes. Chromosome IV has two major regions: the centromeric domain is a-heterochromatic and consists primarily of about ∼3–4 Mbp of short, satellite repeats. This region forms part of the highly condensed chromocenter seen in polytene chromosome spreads. The remaining ∼1.2 Mbp constitutes cytogenetic regions 101E to 102F (Locke and McDermid 1993).
One aspect of chromosome IV genes that needs to be considered is that this autosome may be regulated by an expression-regulation system similar to some dosage-compensation systems that regulate sex chromosomes. The Painting-of-Fourth (POF) protein seems to act in concert with Heterochromatic protein 1 (HP1) in a feedback mediated regulatory system to “fine-tune” the expression of genes on chromosome IV (Stenberg et al. 2009; Riddle et al. 2009; Johansson et al. 2007a, b; Tzeng et al. 2007; Larsson et al. 2001, 2004). The POF protein is encoded by a gene that it is itself on chromosome IV. Notably, flies hemizygous for chromosome IV can survive with few ill effects. However, if the pof gene is mutated, loss of one copy of chromosome IV is lethal (Stenberg et al. 2009). The DNA encompassing the gw gene locus was found to be bound and potentially regulated by POF (Johansson et al. 2007b). Notably, in pof mutant larvae, gw mRNA expression is reduced by one half. Such a fine-tuning mechanism which can compensate for the loss of a whole copy of chromosome IV might explain the variability of defects in within homozygous gw mutant larvae (Schneider et al. 2006).
The Drosophila chromosome IV mapping project has made a concerted effort to expand the relatively small group of mapped and characterized mutations within genes along the fourth chromosome (Sousa-Neves et al. 2005). The gw mutation was identified via screening for mutations in the region predicted to encode a potential GW182 homologue by the Drosophila genome (Schneider et al. 2006). One particular mutation exhibited a striking phenotype, which caused early embryo lethality due to progressive loss of intact embryonic nuclei due to what appeared to be lack of coordination of the early nuclear divisions. This mutation was termed “gawky (gw)” based on the uncoordinated nuclear division phenotype and in anticipation that it was a mutation in the Drosophila GW182 homologue (Schneider et al. 2006).
Using a novel approach exploiting site-directed terminal deficiencies (Sousa-Neves et al. 2005) the gawky recessive zygotic lethal mutation was mapped to a single previously uncharacterized gene, the same locus predicted by the Drosophila genome project to be the single Drosophila GW182 homologue (Adams et al. 2000). Subsequently, this gw mutation was confirmed to be the gw gene, via PCR-sequencing and western blot analysis (Schneider et al. 2006). A particular quirk regarding Drosophila genetic nomenclature, dating back to the original isolation of the white mutation by Morgan (Morgan 1910) is that gene names are traditionally derived from the mutant phenotype (Wilkins 2001). Therefore, anticipation that the uncoordinated gw mutation identified in the mutation screen would be the GW182 homologue was fortunate as it preserved the nomenclature pattern of “GW” while avoiding the inherent logical lapse of referring to the 143 kDa Drosophila Gw protein with the name GW182. While some groups still refer to “Drosophila GW182,” this name is confusing and is not supported by the Flybase consortium which represents the official register of Drosophila nomenclature (Ashburner and Drysdale 1994; Gelbart et al. 1997; Misra et al. 2002). The name Gawky (Gw) also avoids the situation present in other model organisms where GW182 homologues have been given unrelated names (e.g., C. elegans Ain1), while at the same time respecting the long standing tradition of Drosophila gene nomenclature.
8.5 The Phenotype of Drosophila gw 1 Mutation
Drosophila embryos (and many other insect eggs) are syncytial during the earliest stages of development. Notably, cellularization of the rapidly dividing cortical nuclei is not complete until after the 14th nuclear division. After fertilization, the zygotic nucleus undergoes several rounds of mitosis within the center of the egg. In Drosophila, this continues seven more times until 256 nuclei are present within a single syncytial embryo. Most of these nuclei then migrate to the periphery of the embryo. During nuclear division cycle nine several nuclei at the posterior pole become surrounded by invaginating apical membrane to generate the pole cells. These pole cells ultimately give rise to the adult gametes. Notably, this process requires extensive post-transcriptional gene regulatory events (Jin and Xie 2006; Mahowald 2001). The majority of the remaining nuclei arrive at the embryo cortex following nuclear cycle 10. These cortical nuclei then undergo four more mitotic division cycles. Also of note is that the earliest nuclear cycles (1–8) are relatively less sensitive to regulation by cyclins (Edgar and Lehner 1996) and seem to move rapidly from S to M phase, with a complete mitotic cycle occurring approximately every 10 min. Later nuclear cycles occur more slowly, and seem to have greater requirements for the cyclin-based mitotic regulatory machinery. During these later nuclear divisions of the cortical nuclei within the syncytial embryo (syncytial blastoderm), mitosis occur less rapidly.
Beginning at nuclear cycle 13, the apical cell membrane surrounding the embryo begins to invaginate between the nuclei, a process that eventually partitions each somatic nucleus into a single cell—commonly referred to as cellular blastoderm (Foe and Alberts 1983; Turner and Mahowald 1977). Thus, after the first 4 h of development, Drosophila embryos are composed of a cellular blastoderm of approximately 6,000 cells surrounding a central yolk which then undergoes gastrulation to form the cellular layers of the embryo. Notably, in the developing Drosophila embryo, the first 14 nuclear division cycles are precisely synchronized (Edgar and O’Farrell 1989).
The gw 1 mutant lacking the RNA recognition motif (RRM) is the result of a nonsense mutation of the tryptophan codon at position 967 to stop (Schneider et al. 2006). The gw 1 mutant embryos die soon after the nuclear cycle 10 around 2 h after egg deposition (AED) (Schneider et al. 2006). The homozygous mutant shows a disorganized internal structure accompanying abnormal nuclei and cytoskeleton network, consequently failing complete cellularization (Fig. 8.3). High-resolution confocal images of homozygous mutant embryos showed enlarged nuclei accompanied by disposition of centrosomes and severely disorganized microtubule network. These disruptions were confirmed by transmission electron micrography (Schneider et al. 2006).
Initial identification of a gw mutation causing embryonic lethality. A sample of 22 h old embryos produced by ci D /gw parents (ci D is a dominant mutation used to mark the gw + chromosome). Embryos A1-A2 have a wild-type cuticle pattern, A3 is characteristic of a homozygous ci mutation while A4 is characteristic of a homozygous gw mutant. This vacuole was seen consistently in approximately one quarter of the embryos
A difficulty of working with Drosophila genes on chromosome IV is the paucity of visible genetic markers that allow unambiguous sorting of wild-type vs. homozygous mutant animals. Genotyping of homozygous gw 1 mutant embryos required a tedious restriction fragment length polymorphism analysis (Schneider et al. 2006). Similarly, the location of the gw gene made it impossible to use many of the Drosophila methodologies to create embryos that do not have a significant maternal protein contribution (Perrimon 1998). To circumvent these difficulties, a complete loss of Gw phenotype was induced by injection of affinity-purified polyclonal anti-Gw-antibody into the wildtype developing embryos. Blocking Gw function by antibody injection had a rapid effect on embryo nuclear division with the primary phenotypes being mitotic arrest with sister chromatids unable to separate (Schneider et al. 2006). In anti-Gw injected embryos the cytoskeleton network was no longer anchored at the embryo cortex This phenotype shares a great similarity with the gw 1 mutant and the phenotype caused by injection of Ago2 antibody (Schneider et al. 2006).
8.6 Several Drosophila Screens Have Implicated gw in Multiple Processes
Drosophila, as a genetic model system, is used extensively for unbiased screening to discover genes involved in particular processes (St Johnston 2002). Its development, although analogous to mammals, is less complex, requiring only one or two members of the known gene families with defined roles in embryonic differentiation. Interestingly, there seems to be functional conservation between members of the mammalian GW182 gene and Drosophila gw. This would indicate that the relative simplicity of Drosophila compared to mammalian genomes largely represents a lack of redundancy, rather than functional differences in the requirement for a particular gene (Ball and Cherry 2001; Venter et al. 2001). Those working with the Drosophila model system have devised multiple methods to screen the Drosophila genome for genes involved in specific processes (St Johnston 2002; Mathey-Prevot and Perrimon 2006; Reiter and Bier 2002). Accordingly, several screens for a wide variety of biological processes have identified Gw. These include a whole-genome microarray assay of genes involved in the response of females to mating. Gw was one of 23 genes that was reduced at least 1.5-fold in virgin females after they were exposed to courtship by males (Lawniczak and Begun 2004).
Drosophila S2 cells are particularly amenable to large-scale dsRNA knockdown screens. Boutros et al. (2004) showed that knocking down gw (then named CG9905) caused a significant reduction in S2 growth and viability (Boutros et al. 2004). The gw gene was identified as one of 488 genes in a dsRNA based knockout screen for genes involved in cell-cycle progression (Bjorklund et al. 2006). This screen was unique in that it employed flow cytometry to identify specific changes in DNA replication associated with the knockdown phenotypes. Consequently, it identified a large number of loci not found in other screens for cell size and cell cycle progression. One of the most interesting conclusions of this screen was that functional clustering of identified genes tentatively placed gw into a category of p38β/MAPK associated regulators of G2 phase. It is particularly interesting that these recent screens have identified a potential role for gw in widely divergent functional processes suggesting either that mRNA regulation is also important or that Drosophila Gw has roles in addition to mRNA regulation.
8.7 The Organization of the Drosophila Gw Protein is Similar to Mammalian GW182
The Drosophila genome project predicted that all splice isoforms of the gw gene encoded a 143 kDa protein with a high ratio of glycine and tryptophan as GW/WG repeats throughout its sequences (Adams et al. 2000; Stapleton et al. 2002; Drysdale 2003). This gene encoded a protein with a predicted sequence that is 17.8–20% identical and 24–28.3% similar to the human GW182 protein family (Eystathioy et al. 2002; Schneider et al. 2006). The percentage of glycine (G) and tryptophan (W) in Drosophila gw is 12.43% and 2.53 %, respectively with 15 pairs of GW/WG repeats, 12 of which are located within the N-terminal of the protein broadly defined as the GW-rich region (Schneider et al. 2006). This region is followed by a ubiquitin-associated-like domain (UBA) (539–604) and a Q-rich/QN-rich domain (635–861) rich in glutamine (Q) and asparagines (N), whose percentages are 16.81% and 14.61% in this region, respectively. Three additional pairs of GW/WG are interspersed within the following sequences (861–1116) before the RRM (domain—1116–1198). Within the C-terminal region, there is a domain that is rich in serine (S) accounting for 27.62% of the total amino acids (Schneider et al. 2006). Similarly, multiple alignment of Drosophila Gw with other GW182 family proteins identified an additional three highly conservative regions termed: Motif I (1–35), II (312–355), III/Domain of unknown function (DUF 937–1003) (Fig. 8.1) (Behm-Ansmant et al. 2006; Zekri et al. 2009). Finally, there is functional evidence for the region of Gw encompassing amino acids 205–490 exerts the minimal repressive function in its N-terminal in miRNA pathway. Thus, this region has been termed the N-terminal effector domain (NED) (Chekulaeva et al. 2010).
Many of the notable amino acid motifs found within Drosophila Gw, including the GW-rich region, Q-rich domain and RRM are also found within all three human GW182 family members (Eystathioy et al. 2002; Schneider et al. 2006). A homologous region to the UBA domain can only be found between GW and TNRC6C, but iterative PSI-BLAST sequence comparison suggests that all mammalian GW182 family proteins may have this UBA domain (Behm-Ansmant et al. 2006). Therefore, strict comparison to mammalian GW182 would suggest that, TNRC6C is most homologous to GW. However, the basic domain structure of GW is conserved with the entire mammalian GW182 protein family. This is particularly notable as reported GW182 orthologues in another widely used model system Caenorhabditis elegans, are considerably more highly divergent in their overall protein organization. For example, neither C. elegans AIN-1 nor AIN-2 has a well conserved RRM binding domain (Ding et al. 2005). There is some divergence between the human GW182 family and Drosophila GW. A conserved region for binding Ago1 termed as the Ago-hook (Till et al. 2007) reported in human TNRC6B is not present within Drosophila GW. Also, human GW182 family members do not have the concentrated Ser-rich domain within the C-terminal domain. Despite these differences, there is evidence for functional conservation. Notably, when human GW182, TNRC6B and TNRC6C are expressed in Drosophila Schneider2 cells, they form cytoplasmic foci that also recruit Drosophila GW (Schneider et al. 2006). However, a functional conservation for the activities of human GW182 family proteins in S2 cells has not been shown directly.
8.8 Drosophila GW Bodies
Some of the initial biochemical characterizations of the role of GW in the miRNA silencing pathway were reported as early as in 2005 using S2 cells (Rehwinkel et al. 2005). Note that this study referred to GW as Drosophila GW182 (dGW182) as the characterization of the mutant phenotype had yet to be published. In cells of most organisms, GW182 family proteins form cytoplasmic foci (Ding et al. 2005; Eystathioy et al. 2002). Fluorescent-tagged GW was seen co localizing with cytoplasmic bodies, Ago2 (Behm-Ansmant et al. 2006), Me31B (Behm-Ansmant et al. 2006). These were subsequently supported by observations showing that Drosophila Pacman (PCM), the othologue of human being 5′-3′ exonuclease XRN1 also co-localizes with GW in S2 cells. The best proof that Drosophila GW localizes to nonmembrane-bound punctate cytoplasmic bodies shown by transmission electron microscopy and confocal microscopy (Fig. 8.4) (Schneider et al. 2006).
Drosophila Gw bodies. (top) Antibody staining against Drosophila Gw (red) and Ago1 (green) in developing embryos. Significant, but not complete, co-localization is seen between thse two proteins. (bottom) Transmission electron microscopy of a section of the cytoplasm of a Drosophila embryo. Immunogold staining using an anti-Gw antibody Gw bodies detects Gw-bodies of various sizes
The functional localization of GW182 families appears to be a highly conserved process as all 3 human GW182 family proteins also were targeted to GW containing bodies when these human proteins are expressed in Drosophila S2 cells (Schneider et al. 2006). This implies that GW is part of Drosophila mRNA processing bodies as it is consistent with the result of others showing that that GW182 co-localizes with XRN1 in human HEp-2 cells (Eystathioy et al. 2003). Similarly, both mammalian and Drosophila GW bodies dissociate after RNase A treatment indicating that RNA is a significant component of structures in both cell types (Schneider et al. 2006). Many groups are still expanding the list of known GW-body components using Drosophila S2 cells. Recently, an immunoprecipitation assay showed that GW interacts with the decapping activator HPat (Jager and Dorner 2010). Accumulated evidence confirmed that the yeast HPat homologue, Pat1p, is an essential component of P-bodies and required for translational repression and decapping (Eulalio et al. 2007b). Knocking-down HPat in Drosophila cells caused the levels of miRNA-targeted mRNAs level to be slightly elevated (Eulalio et al. 2007a).
8.9 The Role of Drosophila GW in Cytoplasmic mRNA Regulation
Much of the recent research on Drosophila GW has concentrated on elucidating the specifics of its role in miRNA repression and decay. Depleting Ago1, GW and DCP1:DCP2 does not affect NMD and this observation differentiates Ago1 and GW from NMD pathway components UPF1 and SMG7 (Rehwinkel et al. 2005). Using a specific luciferase reporter that measures activity of specific miRNA silencing, Ago1 and GW were confirmed to be primary effectors of the Drosophila miRNA pathway, while Ago2 was revealed to have relatively poor miRNA repression ability (Rehwinkel et al. 2005). This is particularly interesting in light of the fact that some punctate GW co-localized with Ago2 in S2 cells in several studies (Rehwinkel et al. 2005; Schneider et al. 2006). This implies that in Drosophila, the miRNA pathway can function independently of siRNA pathway. Both Dcp1 and Dcp2assist Ago1-GW miRNA repression activities, as the depletion of these decapping factors increased the release of repression by another twofold (Rehwinkel et al. 2005).
The GW protein itself appears to have silencing function independent of some or all of the other members of the canonical miRNA silencing pathway. This was shown by fusing GW to a phage λN-peptide which binds with high affinity to a phage λ BoxB RNA hairpin. By incorporating repeats of these hairpins into a 3′ UTR (F-Luc-5BoxB) downstream of a luciferase reporter, it was shown that GW could independently promote the target degradation without the presence of either Ago1 or miRNA (Behm-Ansmant et al. 2006). Moreover, artificial targeting of GW to mRNAs increases their degradation rate. However, in these same experiments, co-deletion of deadenylation complex components CAF1, NOT1, or the decapping complex component Dcp1:Dcp2restored the cellular levels of the reporter mRNA (Behm-Ansmant et al. 2006). This would suggest that GW is able to trigger mRNA degradation by recruiting deadenylation and decapping complexes from the cytoplasmic pool independently of Ago1 (Iwasaki et al. 2009; Eulalio et al. 2007b). This suggests that GW would functions downstream of Ago1 during miRNA repression in Drosophila cells. This would agree with studies in human cells where GW182 is co-localized with proteins of the 5′ mRNA decapping and deadenylase complex usually associated with P-bodies (Eystathioy et al. 2002, 2003). However, other studies using different reporters that would interact with a 3′ histone H4 stem-loop structure instead of linked to poly-A tail show that GW also represses mRNA independently of adenylation. Therefore, recruitment of the adenylation complex may be a necessary step ONLY for the degradation of the intact RNAs with poly-A tails. Notably this poly-A tail independent RNA degradation seems to require both GW and Ago1 (Eulalio et al. 2009b).
The mechanism by which GW participates in miRNA-mediated degradation remains unclear. GW is released from the target mRNP only when the deadenylase complex is absent, suggesting GW dissociates from the mRNA target after it is deadenylated (Zekri et al. 2009). The C-terminal region is necessary for the release of GW from the target mRNP. GW without C-terminal is not released from a complex with the Ago1 and miRNA targets. Other functional studies have shown that the middle region conserved sequences MII, together with Motif III and C-terminal region of GW bind to PABP1 (Fig. 8.1). This binding is required for the degradation of and interfere with miRNA target interacting with eIF4G (Zekri et al. 2009). The binding is required for the degradation of target RNA possibly through promoting recruitment of the deadenylase complex. However, what remains to be determined is which subset of the total cellular pool of PABP1 binds to GW. It could be the free PABP1from the cytoplasm pool or as part of a complex that circularizes miRNA-targeted mRNAs.
The biochemical interaction between GW and Ago1 has been probed extensively in Drosophila. The Phe594 (F594V) and Phe629 (F629V) amino acids of Ago1 are crucial in miRNA silencing but not important for cap binding (Eulalio et al. 2008). However, mutating both sites may cause a conformational change and lose the ability to bind either miRNA or GW directly or indirectly. This study also showed that binding of GW to endogenous miRNAs was not impaired after reducing Ago1 function, indicating that GW is not involved in miRNA being loaded onto RNA induced silencing complex (RISC) and acts downstream of the assembly (Eulalio et al. 2008). Notably, overexpression of Ago1 seems to alter the GW-Ago1 complex into an inactive state independently of miRNA binding, resulting in a release of the miRNA repression in S2 cells. Therefore, interaction between Ago1 and GW is necessary for Ago1/miRNA-mediated repression (Eulalio et al. 2008). The GW/Ago1 interaction seems to be a regulated process as Ago1 cannot dissociate from GW as well as the decapping and deadenylase complex when ATP is depleted (Iwasaki et al. 2009). This is particularly notable as Ago1-RISC binding to RNA target requires ATP. Finally, Ago1 seems to require the presence of GW for targeting to cytoplasmic P-bodies (Eulalio et al. 2009a). This would suggest that GW has at least two roles in mRNA repression, one independent of the Ago1/miRNA pathway and the other assisting Ago1 to assist in the miRNA repression function possibly through targeting the GW/Ago2 RNP complex to processing bodies where some or all of the associated mRNAs are degraded. Moreover, GW was also reported not to be related to miRNA repression mediated by Ago2 blocking mRNA’s cap structure (Iwasaki et al. 2009). Thus, an unambiguous role for GW in this process is still to be determined.
8.10 Structure/Function Studies of the Role of GW in the Drosophila miRNA Pathway
It has been reported that at least three independent domains within GW protein have potential roles during miRNA repression. Fragments of GW containing amino acids 1–605, 605–830 and 940–1215 decrease the rate of mRNA translation similar to full length GW (Chekulaeva et al. 2009). Later studies mapped a minimal region of GW required for miRNA repression more specifically to amino acids 205–490, and the Ago1 binding domain resides within amino acids 1–204. This domain has been proven to be required for miRNA-mediated repression and degradation and this process is independent of poly-A tail (Chekulaeva et al. 2010).
The role of the N-terminal GW-repeat rich region of GW is still not entirely clear. It has been reported that when 12 GW/WG repeats within GW were mutated to AA pairs, the interaction between GW and Ago1 were severely disrupted (Chekulaeva et al. 2010). The GW 1–539 fragment is sufficient to coimmunoprecipitate Ago1 (Behm-Ansmant et al. 2006). However, only GW/WG repeats in Motif I (M I—Fig. 8.1) are required for GW interaction with Ago1. The GW/WG repeats in the middle the GW protein appear not be essential for binding Ago1 and/or miRNA (Eulalio et al. 2009a). This middle region comprising 3 GW/WG repeats as well as the C-terminal regions of GW are thought to be more important for miRNA based gene silencing (Eulalio et al. 2009a). Finally, it has been reported that the Ago1 binding domain and Q-rich domains, but not UBA-like region, are required for the localization of GW in cytoplasmic foci P-bodies (Eulalio et al. 2009a).
Structural studies of the Drosophila GW RRM domain indicates that it is an RRM fold, with an additional C-terminal α-helix lying on the β-sheet surface shielding the spot used to bind RNA in canonical RRM domain (Eulalio et al. 2009c). The absence of two aromatic amino acids in RNP1 and RNP2 domains would seem to indicate a low affinity for binding RNA. Rather, this domain was suggested to bind other proteins via its hydrophobic cleft (Eulalio et al. 2009c). This region is not essential for the interaction of GW and Ago1, or for P-body localization and is not required for repression function or poly-A independent deadenylation, but assists in a target-specific manner (Eulalio et al. 2009c; Iwasaki et al. 2009).
8.11 A Link Between Drosophila GW-Bodies and Multivesicular Bodies
Many in the field of mRNA regulation have considered GW-bodies and P-bodies as identical structures because GW-containing punctate structures often share many of the same proteins components with P-bodies. This confusion was enhanced by the lack of clear evidence differentiating the biological roles of P-bodies from other GW-containing bodies. However, studies of exosomes (small microvesicles that are released from late endosomal compartments of cells but unrelated to the RNA degradation machinery) in human monocytes first suggested that our concept of a cellular GW-body may need to be considered independently from P-bodies (Gibbings et al. 2009). In these mammalian cells, GW182, Ago2, miRNA and miRNA-repressible mRNA are concentrated with multivesicular bodies(MVB) and endosomes, suggesting that they are the accumulation sites of miRNA-loaded RISC. However while Ago2, which is the core protein in human miRNA-RISC, may be recruited into this subset of GW182 exosome associated structures, these same structures appear to be devoid of the functional P-body marker Dcp1. This would suggest that the Endosomal Sorting Complex Required for Transport (ESCRT) may partition GW182 into the exosomes-lysosomes degradation pathway (Gibbings et al. 2009).
Notably, a role in RNAi for ESCRT sorting of GW has also been confirmed in Drosophila. In a mutagenesis screen devised to identify genes that increase siRNA-mediated RNA silencing discovered that mutation of the locus CG4966 can cause stronger RNAi effect. CG4966 encodes a human Hermansky-Pudlak Syndrome 4 (HPS4) orthologue controlling the turnover of MVBs. Interestingly, RNAi based mRNA silencing is severely impaired when the MVB formation is blocked by mutating Drosophila ESCRT genes hrs and vps25 (Lee et al. 2009). Gibbings et al. found that knocking-down the ESCRT genes vps36, hrs and alix in human monocytes mildly compromised miRNA repression but did not change the miRNA accumulation (Gibbings et al. 2009). In human cells, mutations in HPS4 significantly increase the number of GW-bodies and the quantity of miRNAs being loaded onto the Ago1-RISC, whereas the mutations in MVB formation proteins HRS and TSG101 result in fewer GW-bodies (Gibbings et al. 2009). In Drosophila S2 cells, GW and ME31B are found juxtaposed to the cytosolic phase of MVBs and/or lysosomes. It is also found that mutation in Drosophila homologue HPS4 enhances both siRNA and miRNA-mediated silencing, which would seem to support the findings of Gibbings et al. in mammalian cells. Both groups do agree on a hypothesized mechanistic model that recruiting GW into MVBs is a necessary step for miRNA being loaded on to RISC so as to be a rate-limiting step for miRNA silencing. However, the critical details of this process still need to be addressed.
8.12 Summary and Future Directions
The clear conservation of Drosophila GW to the mammalian GW182 protein family, in terms of both sequence and function has made it a valuable system to model the requirements for these proteins in both cellular functions like miRNA based repression. However, Drosophila studies have also been key to advancing knowledge regarding the function of GW in cellular and developmental processes. An advantage to Drosophila studies is that our findings regarding GW can be fit into an extensive knowledge of the role of mRNA regulation in the cell. Further modeling of the developmental role of GW in early embryonic development as well as later tissue formation and cellular homeostasis should be particularly interesting.
References
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF, George RA, Lewis SE, Richards S, Ashburner M, Henderson SN, Sutton GG, Wortman JR, Yandell MD, Zhang Q, Chen LX, Brandon RC, Rogers YH, Blazej RG, Champe M, Pfeiffer BD, Wan KH, Doyle C, Baxter EG, Helt G, Nelson CR, Gabor Miklos GL, Abril JF, Agbayani A, An HJ, Andrews-Pfannkoch C, Baldwin D, Ballew RM, Basu A, Baxendale J, Bayraktaroglu L, Beasley EM, Beeson KY, Benos PV, Berman BP, Bhandari D, Bolshakov S, Borkova D, Botchan MR, Bouck J, Brokstein P, Brottier P, Burtis KC, Busam DA, Butler H, Cadieu E, Center A, Chandra I, Cherry JM, Cawley S, Dahlke C, Davenport LB, Davies P, de Pablos B, Delcher A, Deng Z, Mays AD, Dew I, Dietz SM, Dodson K, Doup LE, Downes M, Dugan-Rocha S, Dunkov BC, Dunn P, Durbin KJ, Evangelista CC, Ferraz C, Ferriera S, Fleischmann W, Fosler C, Gabrielian AE, Garg NS, Gelbart WM, Glasser K, Glodek A, Gong F, Gorrell JH, Gu Z, Guan P, Harris M, Harris NL, Harvey D, Heiman TJ, Hernandez JR, Houck J, Hostin D, Houston KA, Howland TJ, Wei MH, Ibegwam C, Jalali M, Kalush F, Karpen GH, Ke Z, Kennison JA, Ketchum KA, Kimmel BE, Kodira CD, Kraft C, Kravitz S, Kulp D, Lai Z, Lasko P, Lei Y, Levitsky AA, Li J, Li Z, Liang Y, Lin X, Liu X, Mattei B, McIntosh TC, McLeod MP, McPherson D, Merkulov G, Milshina NV, Mobarry C, Morris J, Moshrefi A, Mount SM, Moy M, Murphy B, Murphy L, Muzny DM, Nelson DL, Nelson DR, Nelson KA, Nixon K, Nusskern DR, Pacleb JM, Palazzolo M, Pittman GS, Pan S, Pollard J, Puri V, Reese MG, Reinert K, Remington K, Saunders RD, Scheeler F, Shen H, Shue BC, Siden-Kiamos I, Simpson M, Skupski MP, Smith T, Spier E, Spradling AC, Stapleton M, Strong R, Sun E, Svirskas R, Tector C, Turner R, Venter E, Wang AH, Wang X, Wang ZY, Wassarman DA, Weinstock GM, Weissenbach J, Williams SM, Woodage T, Worley KC, Wu D, Yang S, Yao QA, Ye J, Yeh RF, Zaveri JS, Zhan M, Zhang G, Zhao Q, Zheng L, Zheng XH, Zhong FN, Zhong W, Zhou X, Zhu S, Zhu X, Smith HO, Gibbs RA, Myers EW, Rubin GM, Venter JC (2000) The genome sequence of Drosophila melanogaster. Science 287:2185–2195
Ashburner M, Drysdale R (1994) FlyBase—the Drosophila genetic database. Development 120:2077–2079
Ball CA, Cherry JM (2001) Genome comparisons highlight similarity and diversity within the eukaryotic kingdoms. Curr Opin Chem Biol 5:86–89
Barbee SA, Estes PS, Cziko AM, Hillebrand J, Luedeman RA, Coller JM, Johnson N, Howlett IC, Geng C, Ueda R, Brand AH, Newbury SF, Wilhelm JE, Levine RB, Nakamura A, Parker R, Ramaswami M (2006) Staufen- and FMRP-containing neuronal RNPs are structurally and functionally related to somatic P bodies. Neuron 52:997–1009
Behm-Ansmant I, Rehwinkel J, Doerks T, Stark A, Bork P, Izaurralde E (2006) mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:Dcp2decapping complexes. Genes Dev 20:1885–1898
Bjorklund M, Taipale M, Varjosalo M, Saharinen J, Lahdenpera J, Taipale J (2006) Identification of pathways regulating cell size and cell-cycle progression by RNAi. Nature 439:1009–1013
Boutros M, Kiger AA, Armknecht S, Kerr K, Hild M, Koch B, Haas SA, Paro R, Perrimon N (2004) Genome-wide RNAi analysis of growth and viability in Drosophila cells. Science 303:832–835
Celniker SE, Dillon LA, Gerstein MB, Gunsalus KC, Henikoff S, Karpen GH, Kellis M, Lai EC, Lieb JD, MacAlpine DM, Micklem G, Piano F, Snyder M, Stein L, White KP, Waterston RH (2009) Unlocking the secrets of the genome. Nature 459:927–930
Chekulaeva M, Filipowicz W, Parker R (2009) Multiple independent domains of dGW182 function in miRNA-mediated repression in Drosophila. RNA 15:794–803
Chekulaeva M, Parker R, Filipowicz W (2010) The GW/WG repeats of Drosophila GW182 function as effector motifs for miRNA-mediated repression. Nucleic Acids Res 38(19):6673–6683
Ding L, Spencer A, Morita K, Han M (2005) The developmental timing regulator AIN-1 interacts with miRISCs and may target the argonaute protein ALG-1 to cytoplasmic P bodies in C. elegans. Mol Cell 19:437–447
Drysdale R (2003) The Drosophila melanogaster genome sequencing and annotation projects: a status report. Brief Funct Genomic Proteomic 2:128–134
Edgar BA, Lehner CF (1996) Developmental control of cell cycle regulators: a fly’s perspective. Science 274:1646–1652
Edgar BA, O’Farrell PH (1989) Genetic control of cell division patterns in the Drosophila embryo. Cell 57:177–187
Ephrussi A, Dickinson LK, Lehmann R (1991) oskar organizes the germ plasm and directs localization of the posterior determinant nanos. Cell 66:37–50
Eulalio A, Behm-Ansmant I, Schweizer D, Izaurralde E (2007a) P-body formation is a consequence, not the cause, of RNA-mediated gene silencing. Mol Cell Biol 27:3970–3981
Eulalio A, Rehwinkel J, Stricker M, Huntzinger E, Yang SF, Doerks T, Dorner S, Bork P, Boutros M, Izaurralde E (2007b) Target-specific requirements for enhancers of decapping in miRNA-mediated gene silencing. Genes Dev 21:2558–2570
Eulalio A, Huntzinger E, Izaurralde E (2008) GW182 interaction with Argonaute is essential for miRNA-mediated translational repression and mRNA decay. Nat Struct Mol Biol 15:346–353
Eulalio A, Helms S, Fritzsch C, Fauser M, Izaurralde E (2009a) A C-terminal silencing domain in GW182 is essential for miRNA function. RNA 15:1067–1077
Eulalio A, Huntzinger E, Nishihara T, Rehwinkel J, Fauser M, Izaurralde E (2009b) Deadenylation is a widespread effect of miRNA regulation. RNA 15:21–32
Eulalio A, Tritschler F, Buttner R, Weichenrieder O, Izaurralde E, Truffault V (2009c) The RRM domain in GW182 proteins contributes to miRNA-mediated gene silencing. Nucleic Acids Res 37:2974–2983
Eystathioy T, Chan EK, Tenenbaum SA, Keene JD, Griffith K, Fritzler MJ (2002) A phosphorylated cytoplasmic autoantigen, GW182, associates with a unique population of human mRNAs within novel cytoplasmic speckles. Mol Biol Cell 13:1338–1351
Eystathioy T, Jakymiw A, Chan EK, Seraphin B, Cougot N, Fritzler MJ (2003) The GW182 protein colocalizes with mRNA degradation associated proteins hDcp1 and hLSm4 in cytoplasmic GW bodies. RNA 9:1171–1173
Foe VE, Alberts BM (1983) Studies of nuclear and cytoplasmic behaviour during the five mitotic cycles that precede gastrulation in Drosophila embryogenesis. J Cell Sci 61:31–70
Gelbart WM, Crosby M, Matthews B, Rindone WP, Chillemi J, Russo Twombly S, Emmert D, Ashburner M, Drysdale RA, Whitfield E, Millburn GH, de Grey A, Kaufman T, Matthews K, Gilbert D, Strelets V, Tolstoshev C (1997) FlyBase: a Drosophila database. The FlyBase consortium. Nucleic Acids Res 25:63–66
Gibbings DJ, Ciaudo C, Erhardt M, Voinnet O (2009) Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat Cell Biol 11:1143–1149
Gilbert DG (2007) DroSpeGe: rapid access database for new Drosophila species genomes. Nucleic Acids Res 35:D480–D485
Hay B, Ackerman L, Barbel S, Jan LY, Jan YN (1988) Identification of a component of Drosophila polar granules. Development 103:625–640
Huntzinger E, Braun JE, Heimstadt S, Zekri L, Izaurralde E (2010) Two PABPC1-binding sites in GW182 proteins promote miRNA-mediated gene silencing. EMBO J 29:4146–4160
Ikeda K, Satoh M, Pauley KM, Fritzler MJ, Reeves WH, Chan EK (2006) Detection of the argonaute protein Ago2 and microRNAs in the RNA induced silencing complex (RISC) using a monoclonal antibody. J Immunol Methods 317:38–44
Iwasaki S, Kawamata T, Tomari Y (2009) Drosophila argonaute1 and argonaute2 employ distinct mechanisms for translational repression. Mol Cell 34:58–67
Jager E, Dorner S (2010) The decapping activator HPat a novel factor co-purifying with GW182 from Drosophila cells. RNA Biol 7:381–385
Jin Z, Xie T (2006) Germline specification: small things have a big role. Curr Biol 16:R966–R967
Johansson AM, Stenberg P, Bernhardsson C, Larsson J (2007a) Painting of fourth and chromosome-wide regulation of the 4th chromosome in Drosophila melanogaster. EMBO J 26:2307–2316
Johansson AM, Stenberg P, Pettersson F, Larsson J (2007b) POF and HP1 bind expressed exons, suggesting a balancing mechanism for gene regulation. PLoS Genet 3:e209
Kim-Ha J, Webster PJ, Smith JL, Macdonald PM (1993) Multiple RNA regulatory elements mediate distinct steps in localization of oskar mRNA. Development 119:169–178
Kobayashi S, Takashima A, Anzai K (1998) The dendritic translocation of translin protein in the form of BC1 RNA protein particles in developing rat hippocampal neurons in primary culture. Biochem Biophys Res Commun 253:448–453
Larsson J, Chen JD, Rasheva V, Rasmuson-Lestander A, Pirrotta V (2001) Painting of fourth, a chromosome-specific protein in Drosophila. Proc Natl Acad Sci U S A 98:6273–6278
Larsson J, Svensson MJ, Stenberg P, Makitalo M (2004) Painting of fourth in genus Drosophila suggests autosome-specific gene regulation. Proc Natl Acad Sci U S A 101:9728–9733
Lawniczak MK, Begun DJ (2004) A genome-wide analysis of courting and mating responses in Drosophila melanogaster females. Genome 47:900–910
Lee YS, Pressman S, Andress AP, Kim K, White JL, Cassidy JJ, Li X, Lubell K, Lim do H, Cho IS, Nakahara K, Preall JB, Preall P, Sontheimer EJ, Carthew RW (2009) Silencing by small RNAs is linked to endosomal trafficking. Nat Cell Biol 11:1150–1156
Lin MD, Fan SJ, Hsu WS, Chou TB (2006) Drosophila decapping protein 1, dDcp1, is a component of the oskar mRNP complex and directs its posterior localization in the oocyte. Dev Cell 10:601–613
Lin MD, Jiao X, Grima D, Newbury SF, Kiledjian M, Chou TB (2008) Drosophila processing bodies in oogenesis. Dev Biol 322:276–288
Locke J, McDermid HE (1993) Analysis of Drosophila chromosome 4 using pulsed field gel electrophoresis. Chromosoma 102:718–723
Mahowald AP (2001) Assembly of the Drosophila germ plasm. Int Rev Cytol 203:187–213
Mansfield JH, Wilhelm JE, Hazelrigg T (2002) Ypsilon Schachtel, a Drosophila Y-box protein, acts antAgonistically to Orb in the oskar mRNA localization and translation pathway. Development 129:197–209
Mathey-Prevot B, Perrimon N (2006) Drosophila genome-wide RNAi screens: are they delivering the promise? Cold Spring Harb Symp Quant Biol 71:141–148
Metzstein MM, Krasnow MA (2006) Functions of the nonsense-mediated mRNA decay pathway in Drosophila development. PLoS Genet 2:e180
Misra S, Crosby MA, Mungall CJ, Matthews BB, Campbell KS, Hradecky P, Huang Y, Kaminker JS, Millburn GH, Prochnik SE, Smith CD, Tupy JL, Whitfied EJ, Bayraktaroglu L, Berman BP, Bettencourt BR, Celniker SE, de Grey AD, Drysdale RA, Harris NL, Richter J, Russo S, Schroeder AJ, Shu SQ, Stapleton M, Yamada C, Ashburner M, Gelbart WM, Rubin GM, Lewis SE (2002) Annotation of the Drosophila melanogaster euchromatic genome: a systematic review. Genome Biol 3:RESEARCH0083
Morgan TH (1910) Sex limited inheritance in Drosophila. Science 32:120–122
Muddashetty R, Khanam T, Kondrashov A, Bundman M, Iacoangeli A, Kremerskothen J, Duning K, Barnekow A, Huttenhofer A, Tiedge H, Brosius J (2002) Poly(A)-binding protein is associated with neuronal BC1 and BC200 ribonucleoprotein particles. J Mol Biol 321:433–445
Ohashi S, Kobayashi S, Omori A, Ohara S, Omae A, Muramatsu T, Li Y, Anzai K (2000) The single-stranded DNA- and RNA-binding proteins pur alpha and pur beta link BC1 RNA to microtubules through binding to the dendrite-targeting RNA motifs. J Neurochem 75:1781–1790
Perrimon N (1998) Creating mosaics in Drosophila. Int J Dev Biol 42:243–247
Quaresma AJ, Bressan GC, Gava LM, Lanza DC, Ramos CH, Kobarg J (2009) Human hnRNP Q re-localizes to cytoplasmic granules upon PMA, thapsigargin, arsenite and heat-shock treatments. Exp Cell Res 315:968–980
Rehwinkel J, Behm-Ansmant I, Gatfield D, Izaurralde E (2005) A crucial role for GW182 and the DCP1:Dcp2decapping complex in miRNA-mediated gene silencing. RNA 11:1640–1647
Reiter LT, Bier E (2002) Using Drosophila melanogaster to uncover human disease gene function and potential drug target proteins. Expert Opin Ther Targets 6:387–399
Riddle NC, Shaffer CD, Elgin SC (2009) A lot about a little dot—lessons learned from Drosophila melanogaster chromosome 4. Biochem Cell Biol 87:229–241
Schneider MD, Najand N, Chaker S, Pare JM, Haskins J, Hughes SC, Hobman TC, Locke J, Simmonds AJ (2006) Gawky is a component of cytoplasmic mRNA processing bodies required for early Drosophila development. J Cell Biol 174:349–358
Sousa-Neves R, Lukacsovich T, Mizutani CM, Locke J, Podemski L, Marsh JL (2005) High-resolution mapping of the Drosophila fourth chromosome using site-directed terminal deficiencies. Genetics 170:127–138
St Johnston D (2002) The art and design of genetic screens: Drosophila melanogaster. Nat Rev Genet 3:176
St Johnston D, Beuchle D, Nusslein-Volhard C (1991) staufen, a gene required to localize maternal RNAs in the Drosophila egg. Cell 66:51–63
Stapleton M, Carlson J, Brokstein P, Yu C, Champe M, George R, Guarin H, Kronmiller B, Pacleb J, Park S, Wan K, Rubin GM, Celniker SE (2002) A Drosophila full-length cDNA resource. Genome Biol 3:RESEARCH0080
Stenberg P, Lundberg LE, Johansson AM, Ryden P, Svensson MJ, Larsson J (2009) Buffering of segmental and chromosomal aneuploidies in Drosophila melanogaster. PLoS Genet 5:e1000465
Tadros W, Lipshitz HD (2009) The maternal-to-zygotic transition: a play in two acts. Development 136:3033–3042
Till S, Lejeune E, Thermann R, Bortfeld M, Hothorn M, Enderle D, Heinrich C, Hentze MW, Ladurner AG (2007) A conserved motif in Argonaute-interacting proteins mediates functional interactions through the Argonaute PIWI domain. Nat Struct Mol Biol 14:897–903
Tritschler F, Huntzinger E, Izaurralde E (2010) Role of GW182 proteins and PABPC1 in the miRNA pathway: a sense of deja vu. Nat Rev Mol Cell Biol 11:379–384
Turner FR, Mahowald AP (1977) Scanning electron microscopy of Drosophila melanogaster embryogenesis. II. Gastrulation and segmentation. Dev Biol 57:403–416
Tzeng TY, Lee CH, Chan LW, Shen CK (2007) Epigenetic regulation of the Drosophila chromosome 4 by the histone H3K9 methyltransferase dSETDB1. Proc Natl Acad Sci U S A 104:12691–12696
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, Gocayne JD, Amanatides P, Ballew RM, Huson DH, Wortman JR, Zhang Q, Kodira CD, Zheng XH, Chen L, Skupski M, Subramanian G, Thomas PD, Zhang J, Gabor Miklos GL, Nelson C, Broder S, Clark AG, Nadeau J, McKusick VA, Zinder N, Levine AJ, Roberts RJ, Simon M, Slayman C, Hunkapiller M, Bolanos R, Delcher A, Dew I, Fasulo D, Flanigan M, Florea L, Halpern A, Hannenhalli S, Kravitz S, Levy S, Mobarry C, Reinert K, Remington K, Abu-Threideh J, Beasley E, Biddick K, Bonazzi V, Brandon R, Cargill M, Chandramouliswaran I, Charlab R, Chaturvedi K, Deng Z, Di Francesco V, Dunn P, Eilbeck K, Evangelista C, Gabrielian AE, Gan W, Ge W, Gong F, Gu Z, Guan P, Heiman TJ, Higgins ME, Ji RR, Ke Z, Ketchum KA, Lai Z, Lei Y, Li Z, Li J, Liang Y, Lin X, Lu F, Merkulov GV, Milshina N, Moore HM, Naik AK, Narayan VA, Neelam B, Nusskern D, Rusch DB, Salzberg S, Shao W, Shue B, Sun J, Wang Z, Wang A, Wang X, Wang J, Wei M, Wides R, Xiao C, Yan C, Yao A, Ye J, Zhan M, Zhang W, Zhang H, Zhao Q, Zheng L, Zhong F, Zhong W, Zhu S, Zhao S, Gilbert D, Baumhueter S, Spier G, Carter C, Cravchik A, Woodage T, Ali F, An H, Awe A, Baldwin D, Baden H, Barnstead M, Barrow I, Beeson K, Busam D, Carver A, Center A, Cheng ML, Curry L, Danaher S, Davenport L, Desilets R, Dietz S, Dodson K, Doup L, Ferriera S, Garg N, Gluecksmann A, Hart B, Haynes J, Haynes C, Heiner C, Hladun S, Hostin D, Houck J, Howland T, Ibegwam C, Johnson J, Kalush F, Kline L, Koduru S, Love A, Mann F, May D, McCawley S, McIntosh T, McMullen I, Moy M, Moy L, Murphy B, Nelson K, Pfannkoch C, Pratts E, Puri V, Qureshi H, Reardon M, Rodriguez R, Rogers YH, Romblad D, Ruhfel B, Scott R, Sitter C, Smallwood M, Stewart E, Strong R, Suh E, Thomas R, Tint NN, Tse S, Vech C, Wang G, Wetter J, Williams S, Williams M, Windsor S, Winn-Deen E, Wolfe K, Zaveri J, Zaveri K, Abril JF, Guigo R, Campbell MJ, Sjolander KV, Karlak B, Kejariwal A, Mi H, Lazareva B, Hatton T, Narechania A, Diemer K, Muruganujan A, Guo N, Sato S, Bafna V, Istrail S, Lippert R, Schwartz R, Walenz B, Yooseph S, Allen D, Basu A, Baxendale J, Blick L, Caminha M, Carnes-Stine J, Caulk P, Chiang YH, Coyne M, Dahlke C, Mays A, Dombroski M, Donnelly M, Ely D, Esparham S, Fosler C, Gire H, Glanowski S, Glasser K, Glodek A, Gorokhov M, Graham K, Gropman B, Harris M, Heil J, Henderson S, Hoover J, Jennings D, Jordan C, Jordan J, Kasha J, Kagan L, Kraft C, Levitsky A, Lewis M, Liu X, Lopez J, Ma D, Majoros W, McDaniel J, Murphy S, Newman M, Nguyen T, Nguyen N, Nodell M, Pan S, Peck J, Peterson M, Rowe W, Sanders R, Scott J, Simpson M, Smith T, Sprague A, Stockwell T, Turner R, Venter E, Wang M, Wen M, Wu D, Wu M, Xia A, Zandieh A, Zhu X (2001) The sequence of the human genome. Science 291:1304–1351
Wilkins AS (2001) Gene names: the approaching end of a century-long dilemma. Bioessays 23:377–378
Wilsch-Brauninger M, Schwarz H, Nusslein-Volhard C (1997) A sponge-like structure involved in the association and transport of maternal products during Drosophila oogenesis. J Cell Biol 139:817–829
Yao B, Li S, Jung HM, Lian SL, Abadal GX, Han F, Fritzler MJ, Chan EK (2011) Divergent GW182 functional domains in the regulation of translational silencing. Nucleic Acids Res 39:2534–2547
Zekri L, Huntzinger E, Heimstadt S, Izaurralde E (2009) The silencing domain of GW182 interacts with PABPC1 to promote translational repression and degradation of microRNA targets and is required for target release. Mol Cell Biol 29:6220–6231
Zhang HL, Pan F, Hong D, Shenoy SM, Singer RH, Bassell GJ (2003) Active transport of the survival motor neuron protein and the role of exon-7 in cytoplasmic localization. J Neurosci 23:6627–6637
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Li, J., Hobman, T.C., Simmonds, A.J. (2013). Gawky (GW) is the Drosophila melanogaster GW182 Homologue. In: Chan, E., Fritzler, M. (eds) Ten Years of Progress in GW/P Body Research. Advances in Experimental Medicine and Biology, vol 768. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-5107-5_8
Download citation
DOI: https://doi.org/10.1007/978-1-4614-5107-5_8
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-5106-8
Online ISBN: 978-1-4614-5107-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)