Abstract
Alleviating the crop loss due to biotic stress is the primary aim of plant biologists to achieve sustainable evergreen revolution in order to feed rapidly growing population. In nature, continuous evolution of plants while interacting with pathogens has generated a complex immune system that consists of preformed barriers and induced responses. The induced responses are further subdivided based upon the recognition of microbe-associated molecular patterns and effectors produced by pathogens; however, overlap exists between the downstream signaling pathways. In last decade, great deal of information about molecular aspects of plant–pathogen interactions has been generated which can be utilized for improving crops through genetic manipulation. Plant breeding has helped in the isolation of species-specific resistance components (R genes) from many plants. The molecular breeding techniques have also helped in pyramiding several components to a single variety, especially QTLs responsible for plant resistance, high yield, and nutritional quality. The identification of nonhost components in model plants and incorporation of genetically modified crops in our cropping system have raised hopes that nonhost resistance can be utilized for generating broad-spectrum pathogen tolerance breaking the barriers of species level resistance. This chapter describes the recent molecular aspects of plant–pathogen interactions focusing on the nonhost resistance components. Additionally, strategies like specific regulation of induced defense responses, manipulation of susceptibility factors, and host-induced gene silencing (HIGS) are discussed. The development of GM crops using such strategies will help in generating higher yields against pathogen infestations.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
In the face of an expanding world population, we need more quantity of food, especially when the cultivated land resources are shrinking. It is estimated that to feed the world population by 2050 our food requirement will be 70% more, which means an increase in crop production at the rate of 44 million metric tons per year is required (Tester and Langridge 2010). The increasing food and energy demand calls for intensive crop production but it is also visualized that in intensive cropping systems the growth of plant pathogens is rapid and new virulence mechanisms appear in pathogen population, and minor pathogens become a major production constrain. Therefore, the incorporation of resistance is a major focus of many breeding programs. However, certain limitations like lack of resistance against many diseases in the primary gene pool, difficulty in transfer of resistance in desired host due to crossability barriers, rapid evolution of virulent pathogens, existence of high pathogenic variability amongst the pathogens, etc. override the advantages of traditional breeding. It is the consensus of plant breeders, geneticists, and other biologists that biotechnological approaches can play an important role in alleviating some of these problems.
Research on host–pathogen interaction in crop plants mainly has been focused on production of toxic substances. Recent advances in molecular biology, however, have offered efficient and precise tools for better understanding of plant–pathogen interactions. In the first half of this chapter, recent developments towards understanding of molecular aspect of plant immunity, mostly against the bacteria and fungi, have been described, although many of these pathways play an important role against other pathogens also. This part is further divided into sections and subsections to provide clearly outlined apprehension of the topic. In the second half of the chapter various methods to achieve resistance against pathogens in crop plants have been discussed.
2 Plant–Pathogen Interactions
Plants are rich source of sugar, minerals, and water that attracts various organisms with heterotrophic lifestyle. The pathogenic organisms use host plant to serve basic aim of life, i.e., grow and reproduce. Plant diseases are comparatively less than the number and variety of potential pathogens in the surrounding environment of plants; this is due to the fact that they have developed a highly complex and multilayered immune system while coevolving with pathogens. The outcome of a plant–pathogen interaction can be either an incompatible (disease resistance/tolerance) or a compatible (pathogen infection and disease) interaction, which is governed by the genetic makeup of both the plant and pathogen. The ability of a pathogen to infect plants depends upon the repertoire of its effectors to suppress or evade plant immune responses and modulate host cellular metabolism for its own benefit. The plant resistance against a potential pathogen depends upon the capacity of the plant to recognize this pathogen as nonself and induce immune response to restrict its growth.
In an ecosystem, pathogens pass their life on host plants in different modes. Many pathogens have evolved to infect only a single plant species (narrow host range) while a few of these pathogens can implicate more plant species for their survival (broad host range). Based on their lifestyle on host, pathogens are classified as biotrophs, hemibiotrophs, and necrotrophs. The biotrophs are entirely dependent upon living host and keep their host alive throughout their life cycle, the hemibiotrophs keep host alive for some period and then kill them, and the necrotrophs feed on host plants by killing them. The evolution of such lifestyles in filamentous pathogens was correlated with gain/loss of genes by comparative analysis (Dodds 2010).
Molecular plant pathologists have broadly classified plant disease resistance operating in natural habitats into two categories: the host resistance and the nonhost resistance (Heath 2000). The nonhost resistance dominates in nature as every plant withstands the injurious effect of most of the potential pathogens while host resistance against a particular pathogen species is shown by the some genotypes of an otherwise host species. To define, the nonhost resistance is the ability of an entire plant species to resist infection by all isolates of a pathogen species. Many reports suggest that the defense signaling against host and nonhost pathogens is similar and many components of these resistance mechanisms are common but the final outcome of their interaction with pathogen or pathogen effectors differs (Thordal-Christensen 2003). It is opined that the components of host resistance were isolated earlier through forward genetics in many known plant–pathogen pairs; hence data towards host resistance seem to be unfair. Therefore, with the advancements in biotechnology biologists are prompted to use the components of nonhost resistance due to its effectiveness and durability.
3 Multilayered Plant Immune System
A simple way to define plant immune system is to define the obstacles that a pathogen must overcome to invade host tissue, proliferate, and cause disease (Thordal-Christensen 2003). Bacterial pathogens get access to host tissue through stomata or wounds. The filamentous pathogens make their entry in host through stomata and may even directly penetrate the cuticle layer. Plants try to restrict pathogens by preformed and induced defenses. The induced defense responses are controlled by PTI (pathogen-associated molecular patterns-triggered immunity) and ETI (effector-triggered immunity). Only pathogens that can evade/suppress/manipulate these defensive layers can cause disease.
3.1 Preformed Structural and Chemical Barriers
The cuticle covers the epidermal cell wall and functions as the first barrier for pathogens. It is composed of polysaccharides, cutin, and waxes, whose composition changes within each species and according to environmental conditions (Shepherd and Wagner 2007). After landing of pathogen on plant surface, the cuticle plays an important part in the plant–pathogen communications. Generally cuticle is considered as a barrier for the entry of pathogens but now it is clear that many pathogens like Uromyces appendiculatus, Fusarium solani f. sp. pisi, Ustilago maydis, Magnoporthe oryzae, Colletotrichum gloeosporioides, Puccinia graminis f. sp. tritici, etc. require signals from the host plant surface to differentiate and penetrate the host (Mendoza et al. 2009; Reina-Pinto and Yephremov 2009; Liu et al. 2011). Thus, the cuticle’s role is important towards resistance against nonadapted pathogens. The phytopathogenic fungi secrete cutinase to liberate cutin that serves as a signal for differentiation in M. grisea and Erysiphe graminis but not in Botrytis cinerea (Bessire et al. 2007). In case of necrotrophic fungi like B. cinerea, Alternaria brassicicola, and Fusarium graminearum secreted lipases are important for pathogenicity. The Blumeria graminis release lipolytic activity containing protein, Lip1, to release cues from the wheat plant surface for promoting pathogen development and infection (Feng et al. 2009). The Arabidopsis CUTE plants with cell wall targeted fungal cutinase, lipase, and mutants with altered cuticle showed higher resistance to B. cinerea but not to other necrotrophs like Plectosphaerella cucumerina, A. brassicicola and Sclerotinia sclerotiorum (Bessire et al. 2007; Chassot et al. 2008). The increased resistance against B. cinerea in these plants was correlated with the induction of few genes and higher fungitoxic activity. Clearly a single mechanism cannot be charted out for the role of cuticle against diverse pathogens but the studies on various cuticle-related mutants have shown that its composition affects the final outcome of plant–pathogen interaction (Table 16.1). The adhesion level of cuticle with cell wall also modulates the defense responses of plants. The glandular trichomes also release antimicrobial substances that can inhibit pathogen growth.
After alteration of cuticle, the pathogens adopt a course of action to break the host plant cell wall by mechanical force and degrading enzymes such as polygalacturonases, xylanases, cellulases, and proteinases. Changes in the host cell wall components like less O-acetylation of cell wall polysaccharides in Arabidopsis thaliana’s Reduced Wall Acetylation (RWA2) mutant plants displayed increased tolerance towards Botrytis cinerea, but mutation had no effect on infection by powdery mildew (Golovinomyces cichoracearum) suggesting differential mechanisms of fungal infection and plant resistance against these pathogens (Manabe et al. 2011). In another case, the atmyb46 mutants have high level of cell wall-associated peroxidases that are involved in phenolic cross-linking at cell wall and ROS scavenging leading to enhanced resistance against B. cinerea (Ramirez et al. 2011). Many other cell wall-associated genes had been reported to influence resistance and susceptibility to pathogens (Hückelhoven 2007; Cantu et al. 2008). The molecules released by cell wall breakdown of the host (i.e., endogenous elicitors) and the pathogen can induce plant defense response, which has been discussed under induced defenses.
The apoplastic space is the site where many pathogen and plant-derived molecules counteract each other. Molecules having antimicrobial activity are secreted in the apoplastic space constitutively by plant or they can be induced after pathogen perception. Many enzyme inhibitors block the activity of pathogen-released enzymes and the plant-derived lipid transfer proteins (LTPs) have inhibitory effects on fungal growth (Molina and Garcia-Olmedo 1997; Patkar and Chattoo 2006). The sad mutants of Avena strigosa can be infected by the nonhost fungal pathogens Gaeumannomyces graminis var. tritici and Fusarium culmorum due to the lack of avenacins, a type of saponin with antifungal activity (Papadopoulou et al. 1999).
3.2 Pathogen-Associated Molecular Pattern-Triggered Immunity
When pathogens are able to breach the constitutive defensive layers then they are recognized as nonself by plant cell membrane receptors, which recognize conserved microbial components (flagellin and chitin in bacteria and fungi respectively) or motifs present in them and molecules released by pathogen. These molecules, termed as PAMPs/MAMPs (microbe-associated molecular patterns), are mostly conserved within a class of microbes and are essential for microbial survival and fitness (Bent and Mackey 2007). They are non-race-specific inducers of plant defense so are often mentioned as exogenous elicitors in contrast to the endogenous elicitors, called damage-associated molecular patterns (DAMPs), released from the host plant by virtue of pathogen attack (Lotze et al. 2007). Some of the known pathogen-associated molecular pattern (PAMPs)/DAMPs are listed in Table 16.2. The importance of PAMP-triggered immunity (PTI) in plant defense is manifested from the fact that the adapted pathogens have evolved effectors to suppress it or they have evolved mechanisms to mask the recognition of PAMPs/DAMPs but in nonhost resistance growth of a nonadapted pathogen is effectively restricted by PTI. The PTI in plants is very similar to that of animals.
The typical responses initiated in plant cell after PAMP/DAMP perception are generation of ion fluxes across plasma membrane, enhanced Ca2+ concentration in cytosol, protein phosphorylation, GTPases activation, rapid increase in reactive oxygen species (ROS), generation of nitric oxide (NO) and ethylene (ET), and many more associated changes (Garcia-Brugger et al. 2006; Boller and Felix 2009). These changes lead to the activation of calcium-dependent protein kinases (CDPKs), calmodulins, and mitogen-activated protein kinases (MAPKs) through cascade of events that ultimately activates the transcription of numerous defense-related genes (Boudsocq et al. 2010). Scientists generally use alkalization of the growth medium, MAPK activation, hydrogen peroxide (H2O2) generation, callose deposition, and expression of early induced genes as markers for the flagellin, chitin, and other PAMP-activated responses (Asai et al. 2002; Denoux et al. 2008). In terms of the quality, responses elicited by various PAMPs from virus, bacteria, oomycetes, fungi, and other pathogens are same but quantitatively they may differ. The cumulative effect of these responses can often lead to hypersensitive response (HR) that is characterized by localized cell death at the site of attack to limit the pathogen spread (Heath 1998; Bolwell 1999).
Many PAMPs have been defined at molecular level based on the activation of PTI responses but their corresponding plant receptors working as sentinels at plasma membrane are not so well defined (Zipfel 2009). The first PAMP receptor cloned from plants was for flagellin (flg22). It is FLAGELLIN-SENSING 2 (FLS2) that encodes for a leucine-rich repeat receptor-like kinase (LRR-RK) (Gomez-Gomez and Boller 2000). The orthologs of FLS2 are present in other higher plants also suggesting that flagellin-mediated signaling is present in both monocot and dicot branches (Takai et al. 2008). Unlike flg22 responsiveness seen in many higher plants, the Brassicaceae family is only responsive to the N-terminus (elf18/26) of a highly conserved and abundant bacterial protein Elongation factor Tu (a GTPase). Its receptor in Arabidopsis, EFR, is also an LRR-RK (Kunze et al. 2004). Such is also the case with the recognition of Ax21 by some specific rice cultivars. It is thus apparent that each plant does not recognize every PAMP and not every pathogen displays all PAMPs (Zipfel and Robatzek 2010).
The nonhost interaction of Arabidopsis thaliana with Blumeria graminis f. sp. hordei (Bgh) has emerged as an excellent system to study the role of early induced genes as the infection is localized at the epidermal cells. Analysis of mutant plants for the various genes like PENETRATION (PEN1- a syntaxin, PEN2-a glycosyl hydolase, and PEN3-an ABC transporter) have suggested their role in plant immunity towards nonadapted pathogens (Ellis 2006).
3.3 Effector-Triggered Immunity
To suppress the PTI and to modulate host metabolism for their own benefit, pathogens secrete a variety of effector molecules inside the host cell (Hok et al. 2010). Bacteria mainly use type III secretion system while filamentous pathogens utilize host machinery to deliver effectors into the plant cell (Göhre and Robatzek 2008; Chibucos et al. 2009). These effectors can be proteases, toxins, transcriptional activators, etc. suggesting that diverse pathogens have evolved various strategies to subvert plant responses (de Jonge et al. 2011; Gheysen and Mitchum 2011; Hogenhout and Bos 2011; Stassen and Van den Ackerveken 2011). In a recent study, related to the interaction of pathogenic effectors with their target plant proteins, it was concluded that two diverse pathogens have evolved their effectors to target a selected set of plant proteins besides other individual targets. These common plant protein targets, in general, form large interaction networks in plants suggesting that pathogens target those proteins inside a host plant that are important for a signaling or interaction hub (Mukhtar et al. 2011). In response to effectors, plants have evolved an array of R (resistance) genes that recognizes these effectors directly or indirectly to rapidly induce a strong defense response. Many of the R proteins are associated with multi-protein immune complexes (Friedman and Baker 2007). Models have been proposed and experimentally verified to explain the evolution of R genes and the recognition of pathogen effectors by R proteins. Relevant among them are gene-for-gene, guard model, and decoy model (van der Hoom and Kamoun 2008).
Most of the known R proteins are multidomain NB-LRR (Nucleotide binding site and leucine-rich repeat) type but other types of R proteins are also known like protein kinase (Rpg1), LRR-receptor-like kinase (Xa21), LRR-TM (Cf’s), etc. and in some genes the promoter polymorphisms also genetically suggest it as R gene (Liu et al. 2007; Bogdanove et al. 2010; Chen et al. 2010). The NB-LRR proteins can be further subdivided based on N-terminal homology to TIR (Toll and Interleukin-1 Receptor; RPS4, SSI4, L6, etc.), CC (coiled-coil; RPM1, RCY1, Mi-1, etc.) or LZ (leucine-zipper; RPS5), and non-motif groups. The C-terminal LRR region binds to the decoy or the effector (direct Avr-R interaction) while the N-terminal is involved in transducing signals to the downstream components to initiate defense signaling. It is suggested that the intra-domain interaction or interaction with associated proteins keeps NB-LRR proteins under resting condition and with the perception of effectors or their activity the signaling is initiated (Caplan et al. 2008; Collier and Moffett 2009; Lukasik and Takken 2009). The signaling downstream to R-proteins is very complex, as some group of R-genes requires NDR1 (non-race-specific disease resistance 1) or EDS1 (enhanced disease susceptibility 1) or some are independent of these two parallel pathways. Further complexity appears in the requirement of RAR1 and SGT1 proteins (Thatcher et al. 2005; Shirasu 2008).
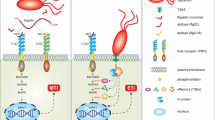
In model plant Arabidopsis and other crop plants various components of preformed and induced (PAMP and effector recognition-based) immunity have be isolated and from these analyses emerges a complex picture of plant immune responses (Fig. 16.1) (Thatcher et al. 2005; Chisholm et al. 2006; Knepper and Day, 2010; Nishimura and Dangl 2010; Zhang and Zhou 2010; Chen and Ronald 2011). The signaling initiated by ETI and PTI shares many common points (Thomma et al. 2011) but the final outcome of defense response, i.e., plant immunity is brought by the cumulative effects of all these components some of which may also be involved in primary and secondary metabolism. A common feature associated with resistance against biotrophic pathogen is the development of hypersensitive response (HR) and systemic acquired resistance (SAR) along with some associated processes (Durrant and Dong 2004; Vlot et al. 2008). Plant hormones like salicylic acid (SA), jasmonic acid (JA), ethylene (ET), auxin, etc. also play an important role along with a myriad of small molecules and proteins in this complex plant response. Role of these components in plant immunity has been extensively reviewed (Lorenzo and Solano 2005; Roberts-Seilaniantz et al. 2007; Spoel and Dong 2008; Bari and Jones 2009; Pieterse et al. 2009; Ton et al. 2009).
4 Strategies to Develop Biotic Stress-Tolerant Crops
Since a number of crop species are cultivated under adverse stress conditions, Varshney et al. (2011) emphasized that the scientists should take up multiple approaches to develop biotic and abiotic stress-tolerant crops with adequate nutritional food value. This will be useful in meeting the food and biofuel security with the growing population and changing environment. As discussed earlier, the plant breeding has played a significant role in crop improvement; still we need to do more. In this context the impact of agrobiotechnology is both productive and benign. We can utilize the most cutting edge works associated with genetic mapping, molecular markers, and biotechnology to accelerate the crop development process. Methods through which crops with enhanced immunity can be generated are discussed in the following sections.
4.1 Molecular Plant Breeding
The plant breeding was the basis of the green revolution that led to increase in wheat and rice production in the twentieth century. The merger of biotechnology with conventional plant breeding techniques along with increase in our knowledge about basic plant biology has led to evolution of molecular plant breeding. Many reviews have discussed the molecular techniques and essential requirements for efficient use of molecular plant breeding in future crops (Jauhar 2006; Wenzel 2006; Moose and Mumm 2008; Hospital 2009; Torres 2010). A number of molecular markers based on simple sequence repeats (SSRs), single nucleotide polymorphism (SNPs), insert-deletions, and candidate gene markers are being developed in several crop species that will assist in genetic analysis and breeding programs (Feuillet et al. 2010). In recent years the next-generation sequencing (NGS) technologies have positively influenced the breeding programs (Varshney et al. 2005, 2010). A greater impact of NGS is noted on the comparative genomic studies which is expected to facilitate breeding programs.
The breeding for disease resistance is the greatest challenge because there is great variability both in plants and pathogens. Although our knowledge about disease resistance mechanisms has increased but still its application for developing resistant varieties is not an easy task because only the genes responsible for species level resistance (host resistance) can be transferred to elite varieties through breeding. Against many pathogens the plant resistance is a complex trait governed by QTLs having major or minor roles; with the advancement of molecular breeding technologies it will be possible to transfer many of the QTLs in elite varieties (Poland et al. 2009).
In breeding programs the field trials need to be well designed as various others environmental factors can also influence the final outcome of plant–pathogen interactions. It is visualized that next decade will be dominated by the high yielding and stress-tolerant varieties developed through traditional and molecular breeding due to the sociopolitical reasons associated with genetically modified (GM) crops.
4.2 Induction of Plant Immunity
Although breeding strategies are useful in enhancing species level resistance, they are time-consuming and have some drawbacks like linkage drag and nonavailability of effective resistant germplasms. The crop production can improve if we espouse environment friendly chemicals that enhance plant immunity, use nonpathogenic microbes as biocontrol agents that induce SAR, and raise transgenic plants with greater potential to recognize the pathogens and execute defense responses (Mourgues et al. 1998; Dita et al. 2006; Collinge et al. 2010; Gust et al. 2010; Shoresh et al. 2010; Wulff et al. 2011).
The initial transgenic crops were developed to overcome pathogen infestations and herbicide tolerance for industrial (ethanol, oil, textile, sugar) use of crops like corn, cotton, sugarcane, soybean, etc (Marshall 2010). When this trend shifted to crops for food consumption then various biosafety and ethical issues were raised, which were also raised for industrial crops but to a lower level. These issues were successfully overcome by the use of marker free transgenic, field trials, and well-designed experiments on animal models, so GM crops are making greater impact on the economy and accepted by people now (Carpenter 2010). Several genes are regularly being tried to get biotic stress-tolerant plants. Transgenic approaches to control herbivore pests are mainly expression of recombinant protease inhibitors and Bacillus thuringiensis endotoxins along with some alternate strategies (Bravo and Soberon 2008; Gatehouse 2008; Schlüter et al. 2010; Sanahuja et al. 2011). Some of the recent publications in this regard are mentioned in Table 16.3. The cis-engineering has provided promoters that precisely express the useful genes in an organ-specific and pathogen-inducible manner depending upon mode of pathogen infection (Venter 2006).
4.3 Manipulation of Susceptibility Factors
It is now very clear that for pathogenesis, plant pathogens manipulate host metabolism and suppress plant defense. In some cases plant proteins behave as susceptibility factors, i.e., plant proteins help in pathogen growth and reproduction leading to disease establishment. The role of a gene in susceptibility can be either because of its own function as negative regulator of plant defense or plant effectors may target its protein product for their own growth, although the gene may have role in plant growth and development in normal conditions (Eckardt 2002; De Almeida et al. 2005; Pavan et al. 2010). The elimination or modification of such plant factors from crop plants can also be a method to achieve resistance against pathogens, although modifications of gene should not have obvious negative consequences on plant health and yield. Many recessive genes that act as negative regulators provide resistance by activating the cell death (cpr, lsd, cim, acd, and mlo) or by unknown mechanisms independent of salicylic acid, jasmonic acid, and ethylene signaling pathways (pmr6).
In one of the best examples of a susceptibility gene, barley’s Mlo (Mildew Resistance Locus o) gene is required for successful colonization by the ascomycete B. graminis f. sp. hordei (Humphry et al. 2006). Nonfunctional mutant alleles of this gene provide durable resistance in many elite varieties of barley after their introgression into them. Its role in powdery mildew pathogenesis has also been found in Arabidopsis, tomato, and pea plants (Consonni et al. 2006; Bai et al. 2008; Humphry et al. 2011). The gene seems to function as a suppressor of nonhost defense response components/signaling as resistance in mlo mutant plants and nonhost resistance share analogous features (Humphry et al. 2006). The pmr6 mutants showed enhanced recessive resistance to Erysiphe orontii and E. cichoracearum but these mutant plants were susceptible to P. parasitica (Vogel and Somerville 2000; Vogel et al. 2002). The pmr6 gene encodes for a pectate lyase-like protein with extended C-terminal, the mutations in this gene show pleiotropic effects on plant growth, and the cell wall of these plants have high pectin content. The eukaryotic translation initiation factor subunits (mostly elF4E and elF4G) act as susceptibility factors for viral infections mainly potyviruses (Robaglia and Caranta 2006; Piron et al. 2010). In Arabidopsis a pathogen-inducible patatin-like lipid acyl hydrolase (PLP2) facilitates fungal and bacterial colonization (La Camera et al. 2005) and in rice loss of a proline-rich protein (Pi21) confers durable disease resistance (Fukuoka et al. 2009). The transcription-activator-like (TAL) effector proteins of bacteria target many susceptible factors and in resistant plants they are recognized by many R-genes (Lewis et al. 2009; Bogdanove et al. 2010). A group of ‘SWEET’ sugar efflux transporters are induced by several pathogens and it was shown that TAL effectors in case of Xanthomonas spp. regulate their induction for pathogen growth (Chen et al. 2010).
The availability of genome editing in plants and further technology improvements will help scientists to manipulate the pathogen-induced expression or the whole susceptibility gene from plant. Thus, this powerful method can also increase the hope for improved GM crops with durable disease resistance (Weinthal et al. 2010).
4.4 Host-Induced Gene Silencing in Pathogens
The sequencing projects of various pathogens especially filamentous pathogens have revealed that their effectors are rapidly evolving as compared to other genes and their genomes are rich in transposons (Dodds 2010). This suggests that in near future more virulent strains of a pathogen will emerge like the highly virulent strain of Puccinia graminis f. sp. tritici Ug99 and events of host jumps may also be seen. In the long run, breeding and induced defense-based approaches will work only against pathogens that will evolve slowly but approaches that target the basic cellular and pathogenicity mechanisms of pathogens would provide long-lasting resistance. The RNA interference (RNAi; RNA-guided regulation of RNA transcripts) based approach would make an ideal choice against rapidly evolving pathogens, as it is known to provide resistance against viral infection in natural environment (Baulcombe 2004). Transgenic plants with RNAi constructs targeting specific genes of pathogens have shown resistance against viruses, parasitic nematodes, herbivorous insects, and parasitic weeds in many plants (Huang et al. 2006; Frizzi and Huang 2010; Niu et al. 2010; Wani et al. 2010). In an unsuccessful attempt, the Plasmodiophora brassicae gene was also checked for downregulation on transgenic Arabidopsis thaliana plants as this phytomyxea pathogen remains in intimate contact with host cell (Bulman 2006).
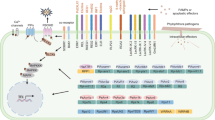
Considering the situation that ∼70% of all major crop diseases are caused by fungal pathogens (Agrios 2005), this RNAi technology against fungi would greatly help to increase crop yield. Two prerequisites for successful silencing of fungal genes on transgenic plants would be the transfer of silencing-RNAs from host plant cell to the fungi and a functional RNAi machinery of the pathogenic fungi. Many independent groups have reported the silencing of genes using RNAi constructs in fungi suggesting that the RNAi machinery works in many fungi. The uptake of dsRNA from outside the fungal cells and subsequent silencing of the targeted fungal gene transcripts were claimed in two US patents (Van De Craen et al. 2006; Roberts et al. 2008). Tinoco et al. (2010) reported silencing of the gus transcripts in transgenic Fusarium verticillioides when it was inoculated on transgenic tobacco plants expressing RNAi construct against gus gene. Nowara et al. (2010) also showed that dsRNA or siRNA molecules were exchanged between cereal hosts and the obligate biotrophic fungal pathogen Blumeria graminis and they called this technique of downregulating pathogen genes as host-induced gene silencing (HIGS). Using transient expression, virus-induced gene silencing (VIGS), and transgenic plants with RNAi constructs it was proved that HIGS could be an effective tool to study the role of fungal genes in pathogenesis and it has the potential of disease control against biotrophic fungal pathogens (Fig. 16.2). Using VIGS the genes that are expressed in haustorial cells were silenced efficiently in Puccinia striiformis f. sp. tritici rather than the genes that are constitutively expressed in whole pathogen, probably pointing towards the fact that tissue which remains in intimate contact with host will receive more silencing-RNAs (Yin et al. 2011). More experiments with other systems are needed to standardize this technology before engineering at mass level and also the questions regarding the silencing of genes in hemibiotrophs and necrotrophs need to be answered. The usefulness of fungal inducible promoters to drive the RNAi constructs should help but the most important thing is to check for RNAi constructs off-targets and avoid it inside the plant cell. Overall the HIGS technology holds promise for generating fungal-tolerant crops leading to higher grain yield and it is believed that in future a common terminology of HIGS will be followed to make scientific literature retrieval easy regarding this type of silencing.
Host-induced gene silencing (HIGS). Genes essential for pathogen growth on host plants can be downregulated by RNAi approach to limit the pathogen growth. (i) High-throughput approaches like virus-induced gene silencing (VIGS) can be used to identify the genes involved in pathogen growth and reproduction on host plants and (ii) the pathogen inducible promoter (PIP) can be used to generate transgenic plants having RNAi constructs against the gene/or genes of a pathogen
5 Conclusions and Future Prospects
We have come a long way in crop improvement from traditional elite variety selection to the development molecular breeding and transgenic crops. But our demand of food supply still needs rapid progress with growing population and nemesis of adverse environmental conditions. Also the increase in demand for biofuels will add more pressure on arable land. In this decade a great deal of information has been achieved about molecular aspects of plant–pathogen interactions. The technological advancements have certainly played a major role in this regard. Now, every aspect of plant–pathogen interaction is studied and sequencing of many crop plants and their pathogens will help in pyramiding various genes through marker-assisted selection especially against notorious pests and necrotrophic fungi where resistance is governed by many QTLs. Contrary to the biosafety-related opinions raised regarding GM crops, molecular plant biologists are optimistic about the need to incorporate GM crops in our crop improvement chain as it can be applied to all the crops outside the limits of species. Already more that 20% of arable land is under the GM crops in countries like USA, Brazil, and Argentina, which dictates the success story of GM crops.
We still need to study and effectively use the nonhost resistance components for high yielding disease-tolerant crops. In case of GM crops effective regulatory mechanisms and safeguards need to be installed to avoid any biosafety-related problem in future and the fields should be monitored regularly for the evolution of new pathogens against resistant crops. The need for translational of basic research to the field crops is more from public sector as investments are more in this sector. The areas where still we can improve for production of stress tolerance crops need to be evaluated and programs need to be implemented especially in developing countries.
References
Agrios GN (2005) Plant pathology, 5th edn. Academic, San Diego
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL (2002) MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415:977–983
Allen A, Islamovic E, Kaur J, Gold S, Shah D, Smith TJ (2011) Transgenic maize plants expressing the Totivirus antifungal protein, KP4, are highly resistant to corn smut. Plant Biotechnol J 9:857–864
Almasia NI, Bazzini AA, Hopp HE, Rovere CV (2008) Overexpression of snakin-1 gene enhances resistance to Rhizoctonia solani and Erwinia carotovora in transgenic potato plants. Mol Plant Pathol 9:329–338
Anuradha S, Divya K, Jami SK, Kirti PB (2008) Transgenic tobacco and peanut plants expressing a mustard defensin show resistance to fungal pathogens. Plant Cell Rep 27:1777–1786
Bai Y, Pavan S, Zheng Z, Zappel NF, Reinstädler A, Lotti C, De Giovanni C, Ricciardi L, Lindhout P, Visser R, Theres K, Panstruga R (2008) Naturally occurring broad-spectrum powdery mildew resistance in a Central American tomato accession is caused by loss of Mlo function. Mol Plant Microbe Interact 21:30–39
Ban H, Chai X, Lin Y, Zhou Y, Peng D, Zhou Y, Zou Y, Yu Z, Sun M (2009) Transgenic Amorphophallus konjac expressing synthesized acyl-homoserine lactonase (aiiA) gene exhibit enhanced resistance to soft rot disease. Plant Cell Rep 28:1847–1855
Bari R, Jones JDG (2009) Role of plant hormones in plant defense responses. Plant Mol Biol 69:473–488
Baulcombe DC (2004) RNA silencing in plants. Nature 431:356–363
Bent AF, Mackey D (2007) Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu Rev Phytopathol 45:399–436
Bessire M, Chassot C, Jacquat AC, Humphry M, Borel S, MacDonald-Comber Petétot J, Métraux JP, Nawrath C (2007) A permeable cuticle in Arabidopsis leads to a strong resistance to Botrytis cinerea. EMBO J 26:2158–2168
Bharathi Y, Kumar SV, Pasalu IC, Balachandran SM, Reddy VD, Rao KV (2011) Pyramided rice lines harbouring Allium sativum (asal) and Galanthus nivalis (gna) lectin genes impart enhanced resistance against major sap-sucking pests. J Biotechnol 152:63-71
Bogdanove AJ, Schornack S, Lahaye T (2010) TAL effectors: finding plant genes for disease and defense. Curr Opin Plant Biol 13:394–401
Boller T (1995) Chemoperception of microbial signals in plant cells. Annu Rev Plant Physiol Plant Mol Biol 46:189–214
Boller T, Felix G (2009) A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu Rev Plant Biol 60:379–406
Bolwell GP (1999) Role of active oxygen species and NO in plant defense responses. Curr Opin Plant Biol 2:287–294
Boudsocq M, Willmann MR, McCormack M, Lee H, Shan L, He P, Bush J, Cheng SH, Sheen J (2010) Differential innate immunity signaling via Ca2+ sensor protein kinases. Nature 464:418–422
Bravo A, Soberon M (2008) How to cope with insect resistance to Bt toxins? Trends Biotechnol 26:573–579
Brunner F, Rosahl S, Lee J, Rudd JJ, Geiler C, Kauppinen S, Rasmussen G, Scheel D, Nurnberger T (2002) Pep1-13, a plant defense-inducing pathogen associated pattern from Phytophthora transglutaminases. EMBO J 21:6681–6688
Bulman SR (2006) Testing the effect of in planta RNA silencing on Plasmodiophora brassicae infection. Ph.D thesis. Lincoln University, Chester County
Cantu D, Vicente AR, Labavitch JM, Bennett AB, Powell ALT (2008) Strangers in the matrix: plant cell walls and pathogen susceptibility. Trends Plant Sci 13:610–617
Cao A, Xing L, Wang X, Yang X, Wang W, Sun Y, Qian C, Ni J, Chen Y, Liu D, Wang X, Chen P (2011) Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc Natl Acad Sci USA 108:7727–7732
Caplan J, Padmanabhan M, Dinesh-Kumar SP (2008) Plant NB-LRR immune receptors: from recognition to transcriptional reprogramming. Cell Host Microbe 3:126–135
Carrillo L, Martinez M, Ramessar K, Cambra I, Castanera P, Ortego F, Diaz I (2011) Expression of a barley cystatin gene in maize enhances resistance against phytophagous mites by altering their cysteine-proteases. Plant Cell Rep 30:101–112
Carpenter JE (2010) Peer-reviewed surveys indicate positive impact of commercialized GM crops. Nat Biotechnol 28:319–321
Chassot C, Nawrath C, Metraux JP (2007) Cuticular defects lead to full immunity to a major plant pathogen. Plant J 49:972–980
Chassot C, Nawrath C, Metraux JP (2008) The cuticle: not only a barrier for plant defense. Plant Signal Behav 3:142–144
Chen L, Zhang ZY, Liang HX, Liu HX, Du LP, Xu H, Xin Z (2008) Overexpression of TiERF1 enhances resistance to sharp eyespot in transgenic wheat. J Exp Bot 59:4195–4204
Chen R, Li H, Zhang L, Zhang J, Xiao J, Ye Z (2007) CaMi, a root-knot nematode resistance gene from hot pepper (Capsium annuum L.) confers nematode resistance in tomato. Plant Cell Rep 26:895–905
Chen X, Ronald PC (2011) Innate immunity in rice. Trends Plant Sci 16:451–459
Chen Y, Tian JC, Shen ZC, Peng YF, Hu C, Guo YY, Ye GY (2010b) Transgenic rice plants expressing a fused protein of Cry1Ab/Vip3H has resistance to rice stem borers under laboratory and field conditions. J Econ Entomol 103:1444–1453
Chen L-Q, Hou B-H, Lalonde S, Takanaga H, Hartung ML, Qu X-Q, Guo W-J, Kim J-G et al (2010) Sugar transporters for intercellular exchange and nutrition of pathogens. Nature 468:527–532
Chibucos MC, Tseng TT, Setubal JC (2009) Describing commonalities in microbial effector delivery using the Gene Ontology. Trends Microbiol 17:312–319
Chisholm ST, Goaker G, Day B, Staskawicz BJ (2006) Host-Microbe interactions: shaping the evolution of the plant immune system. Cell 124:803–814
Choi MS, Kim YH, Park HM, Seo BY, Jung JK, Kim ST, Kim MC, Shin DB, Yun HT, Choi IS, Kim CK, Lee JY (2009) Expression of BrD1, a plant defensin from Brassica rapa, confers resistance against brown plant hopper (Nilaparvata lugens) in transgenic rices. Mol Cells 28:131–137
Collier SM, Moffett P (2009) NB-LRRs work a “bait and switch” on pathogens. Trends Plant Sci 14:521–529
Collinge DB, Jorgensen HJL, Lund OS, Lyngkjaer MF (2010) Engineering pathogen resistance in crop plants: current trends and future prospects. Annu Rev Plant Pathol 48:269–291
Consonni C, Humphry ME, Hartmann HA, Livaja M, Durner J, Westphal L, Vogel J, Lipka V, Kemmerling B, Schulze-Lefert P, Somerville SC, Panstruga R (2006) Conserved requirement for a plant host cell protein in powdery mildew pathogenesis. Nat Genet 38:716–720
Cox KD, Layne DR, Scorza R, Schnabel G (2006) Gastrodia anti-fungal protein from the orchid Gastrodia elata confers disease resistance to root pathogens in transgenic tobacco. Planta 224:1373–1383
Curvers K, Seifi H, Mouille G, de Rycke R, Asselbergh B, Van Hecke A, Vanderschaeghe D, Höfte H, Callewaert N, Van Breusegem F, Höfte M (2010) Abscisic acid deficiency causes changes in cuticle permeability and composition that influence tomato resistance to Botrytis cinerea. Plant Physiol 154:847–860
De Almeida EJ, Favery B, Engler G, Abad P (2005) Loss of susceptibility as an alternative for nematode resistance. Curr Opin Biotechnol 16:112–117
De Jonge R, Bolton MD, Thomma BPHJ (2011) How filamentous pathogen co-opt plants: the ins and outs of fungal effector. Curr Opin Plant Biol 14:400–406
Denoux C, Galletti R, Mammarella N, Gopalan S, Werck D, De Lorenzo G, Ferrari S, Ausubel FM, Dewdney J (2008) Activation of defense response pathways by OGs and flg22 elicitors in Arabidopsis seedlings. Mol Plant 1:423–445
Dita MA, Rispail N, Prats E, Rubiales D, Singh KB (2006) Biotechnological approaches to overcome biotic and abiotic stress constraints in legumes. Euphytica 147:1–24
Dodds PN (2010) Genome evolution in plant pathogens. Science 330:1486–1487
Dong N, Liu X, Lu Y, Du L, Xu H, Liu H, Xin Z, Zhang Z (2010) Overexpression of TaPIEP1, a pathogen-induced ERF gene of wheat, confers host-enhanced resistance to fungal pathogen Bipolaris sorokiniana. Funct Integr Genomics 10:215–226
Dong X, Ji R, Guo X, Foster SJ, Chen H, Dong C, Liu Y, Hu Q, Liu S (2008) Expressing a gene encoding wheat oxalate oxidase enhances resistance to Sclerotinia sclerotiorum in oilseed rape (Brassica napus). Planta 228:331–340
Dubouzet JG, Maeda S, Sugano S, Ohtake M, Hayashi N, Ichikawa T, Kondou Y, Kuroda H, Horii Y, Matsui M, Oda K, Hirochika H, Takatsuji H, Mori M (2011) Screening for resistance against Pseudomonas syringae in rice-FOX Arabidopsis lines identified a putative receptor-like cytoplasmic kinase gene that confers resistance to major bacterial and fungal pathogens in Arabidopsis and rice. Plant Biotechnol J 9:466–485
Dunse KM, Stevens JA, Lay FT, Gaspar YM, Heath RL, Anderson MA (2010) Coexpression of potato type I and II proteinase inhibitors gives cotton plants protection against insect damage in the field. Proc Natl Acad Sci USA 107:15011–15015
Durrant WE, Dong X (2004) Systemic acquired resistance. Annu Rev Phytopathol 42:185–209
Eckardt NA (2002) Plant disease susceptibility genes? Plant Cell 14:1983–1986
Ellis J (2006) Insights into nonhost disease resistance: can they assist disease control in agariculture? Plant Cell 18:523–528
Erbs G, Newman MA (2012) The role of lipopolysaccharide and peptidoglycan, two glycosylated bacterial microbe-associated molecular patterns (MAMPs), in plant innate immunity. Mol Plant Pathol 13:95–104
Felix G, Boller T (2003) Molecular sensing of bacteria in plants. The highly conserved RNA-binding motif RNP-1 of bacterial cold shock proteins is recognized as an elicitor signal in tobacco. J Biol Chem 278:6201–6208
Feng J, Wang F, Liu G, Greenshields D, Shen W, Kaminskyj S, Hughes GR, Peng Y, Selvaraj G, Zou J, Wei Y (2009) Analysis of Blumeria graminis-secreted lipase reveals the importance of host epicuticular wax components for fungal adhesion and development. Mol Plant Microbe Interact 22:1601–1610
Feuillet C, Leach JE, Rogers J, Schnable PS, Eversole K (2010) Crop genome sequencing: lessons and rationales. Trends Plant Sci 16:77–88
Fliegmann J, Mithofer A, Wanner G, Ebel J (2004) An ancient enzyme domain hidden in the putative β-glucan elicitor receptor of soybean may play an active part in the perception of pathogen-associated molecular patterns during broad host resistance. J Biol Chem 279:1132–1140
Fradin EF, Abd-El-Haliem A, Masini L, van den Berg GCM, Joosten MHAG, Thomma BPHJ (2011) Interfamily transfer of tomato Ve1 mediates Verticillium resistance in Arabidopsis. Plant Physiol 156:2255–2265
Friedman AR, Baker BJ (2007) The evolution of resistance genes in multi-protein plant resistance systems. Curr Opin Genet Dev 17:493–499
Frizzi A, Huang S (2010) Tapping the silencing pathways for plant biotechnology. Plant Biotechnol J 8:655–677
Fukuoka S, Saka N, Koga H, Ono K, Shimizu T, Ebana K, Hayashi N, Takahashi A, Hirochika H, Okuno K, Yano M (2009) Loss of function of a proline-containing protein confers durable disease resistance in rice. Science 325:998–1001
Garcia-Brugger A, Lamotte O, Vandelle E, Bourque S, Lecourieux D, Poinssot B, Wendehenne D, Pugin A (2006) Early signaling events induced by elicitors of plant defenses. Mol Plant Microbe Interact 19:711–724
Gatehouse JA (2008) Biotechnological prospects for engineering insect-resistant plants. Plant Physiol 146:881–887
Gaulin E, Drame N, Lafitte C, Torto-Alalibo T, Martinez Y, Torregrosa CA, Khatib M, Mazarguil H, Villalba-Mateos F, Kamoun S, Mazars C, Dumas B, Bottin A, Esquerre-Tugaye MT, Rickauer M (2006) Cellulose binding domains of a Phytophthora cell wall protein are novel pathogen-associated molecular patterns. Plant Cell 18:1766–1777
Gheysen G, Mitchum MG (2011) How nematodes manipulate plant development pathways for infection? Curr Opin Plant Biol 14:415–421
Göhre V, Robatzek S (2008) Breaking the barriers: microbial effector molecules subvert plant immunity. Annu Rev Phytopathol 46:189–215
Gomez-Gomez L, Boller T (2000) FLS2: An LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Granado J, Felix G, Boller T (1995) Perception of fungal sterols in plant (subnanomolar concentrations of ergosterol elicit extracellular alkalization in tomato cells). Plant Physiol 107:485–490
Gust AA, Brunner F, Nürnberger T (2010) Biotechnological concepts for improving plant innate immunity. Curr Opin Biotechnol 21:204–210
Heath MC (1998) Apoptosis, programmed cell death and the hypersensitive response. Eur J Plant Pathol 104:117–124
Heath MC (2000) Nonhost resistance and nonspecific plant defenses. Curr Opin Plant Biol 3:315–319
Hogenhout SA, Bos JIB (2011) Effector proteins that modulate plant-insect interaction. Curr Opin Plant Biol 14:422–428
Hok S, Attard A, Keller H (2010) Getting the most from the host: how pathogens force plants to cooperate in disease. Mol Plant Microbe Interact 23:1253–1259
Hospital F (2009) Challenges for effective marker-assisted selection in plants. Genetica 136:303–310
Huang G, Allen R, Davis EL, Baum TJ, Hussey RS (2006) Engineering broad root-knot resistance in transgenic plants by RNAi silencing of a conserved and essential root-knot nematode parasitism gene. Proc Natl Acad Sci USA 103:14302–14306
Hückelhoven R (2007) Cell wall-associated mechanisms of disease resistance and susceptibility. Annu Rev Phytopathol 45:101–127
Humphry M, Consonni C, Panstruga R (2006) mlo-based powdery mildew immunity: silver bullet or simply non-host resistance? Mol Plant Pathol 7:605–610
Humphry M, Reinstädler A, Ivanov S, Bisseling T, Panstruga R (2011) Durable broad-spectrum powdery mildew resistance in pea er1 plants is conferred by natural loss-of-function mutations in PsMLO1. Mol Plant Pathol 12:866–878
Hwang IS, Hwang BK (2011) The pepper mannose-binding lectin gene CaMBL1 is required to regulate cell death and defense responses to microbial pathogens. Plant Physiol 155:447–463
Imamura T, Yasuda M, Kusano H, Nakashita H, Ohno Y, Kamakura T, Taguchi S, Shimada H (2010) Acquired resistance to the rice blast in transgenic rice accumulating the antimicrobial peptide thanatin. Transgenic Res 19:415–442
Iqbal MM, Nazir F, Ali S, Asif MA, Zafar Y, Iqbal J, Ali GM (2012) Over expression of rice chitinase gene in transgenic peanut (Arachis hypogaea L.) improves resistance against leaf spot. Mol Biotechnol 50:129–136
Janni M, Sella L, Favaron F, Blechl AE, Lorenzo GD, D’Ovidio R (2008) The expression of a bean PGIP in transgenic wheat confers increased resistance to the fungal pathogen Bipolaris sorokiniana. Mol Plant Microbe Interact 21:171–177
Jayaraj J, Punja ZK (2007) Combined expression of chitinase and lipid transfer protein genes in transgenic carrot plants enhances resistance to foliar fungal pathogens. Plant Cell Rep 26:1539–1546
Jha S, Chattoo BB (2010) Expression of a plant defensin in rice confers resistance to fungal phytopathogens. Transgenic Res 19:373-384
Jauhar P (2006) Modern biotechnology as an integral supplement to conventional plant breeding: the prospects and challenges. Crop Sci 46:1841–1859
Kim JG, Jeon E, Oh J, Moon JS, Hwang I (2004) Mutational analysis of Xanthomonas hairpin HpaG identifies a key functional region that elicits the hypersensitive response in nonhost plants. J Bacteriol 186:6239–6247
Klarzynski O, Descamps V, Plesse B, Yvin JC, Kloareq B, Fritiq B (2003) Sulfated fucan oligosaccharides elicit defense responses in tobacco and local systemic resistance against tobacco mosaic virus. Mol Plant Microbe Interact 16:115–122
Kern MF, Maraschin SF, Endt DV, Schrank A, Vainstein MH, Pasquali G (2010) Expression of a chitinase gene from Metarhizium anisopliae in tobacco plants confers resistance against Rhizoctonia solani. Appl Biochem Biotechnol 160:1933-1946
Knecht K, Seyffarth M, Desel C, Thurau T, Sherameti I, Lou B, Oelmuller R, Cai D (2010) Expression of BvGLP-1 encoding a germin-like protein from sugar beet in Arabidopsis thaliana leads to resistance against phytopathogenic fungi. Mol Plant Microbe Interact 23:446–457
Knepper C, Day B (2010) From perception to activation: the molecular-genetic and biochemical landscape of disease resistance signaling in plants. In: The Arabidopsis book. American Society of Plant Biologists, Rockville
Koga J, Yamauchi T, Shimura M, Ogawa N, Oshima K, Umemura K, Kikuchi M, Ogasawara N (1998) Cerebrosides A and C, sphingolipid elicitors of hypersensitive cell death and phytoalexin accumulation in rice plants. J Biol Chem 273:31985–31991
Kunze G, Zipfel C, Robatzek S, Niehaus K, Boller T, Felix G (2004) The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16:3496–3507
Lacombe S, Rougon-Cardoso A, Sherwood E, Peeters N, Dahlbeck D, van Esse HP, Smoker M, Rallapalli G, Thomma BP, Staskawicz B, Jones JD, Zipfel C (2010) Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat Biotechnol 28:365–369
Lee SC, Hwang IS, Choi HW, Hwang BK (2008) Involvement of the pepper antimicrobial protein CaAMP1 gene in broad spectrum disease resistance. Plant Physiol 148:1004–1020
Li XQ, Wei JZ, Tan A, Aroian RV (2007b) Resistance to root-knot nematode in tomato roots expressing a nematicidal Bacillus thuringiensis crystal protein. Plant Biotechnol J 5:455–464
Li Z, Zhou M, Zhang Z, Ren L, Du L, Zhang B, Xu H, Xin Z (2011) Expression of a radish defensin in transgenic wheat confers increased resistance to Fusarium graminearum and Rhizoctonia cerealis. Funct Integr Genomics 11:63–70
L’Haridon F, Besson-Bard A, Binda M, Serrano M, Abou-Mansour E, Balet F, Schoonbeek HJ, Hess S, Mir R, Leon J, Lamotte O, Metraux JP (2011) A permeable cuticle is associated with the release of reactive oxygen species and induction of innate immunity. PLoS Pathog 7:e1002148
La Camera S, Geoffroy P, Samaha H, Ndiaye A, Rahim G, Legrand M, Heitz T (2005) A pathogen-inducible patatin-like lipid acyl hydrolase facilitates fungal and bacterial host colonization in Arabidopsis. Plant J 44:810–825
Laquitaine L, Gomes E, Francois J, Marchive C, Pascal S, Hamdi S, Atanassova R, Delrot S, Coutos-Thevenot P (2006) Molecular basis of ergosterol-induced protection of grape against Botrytis cinerea: induction of type I LTP promoter activity, WRKY, and stilbene synthase gene expression. Mol Plant Microbe Interact 19:1103–1112
Lee J, Klessig DF, Nurnberger T (2001) A Harpin binding site in tobacco plasma membranes mediates activation of the pathogenesis-related gene HIN1 independent of extracellular calcium but dependent on mitogen-activated protein kinase activity. Plant Cell 13:1079–1093
Lewis JD, Guttman DS, Desveaux D (2009) The targeting of plant cellular systems by injected type III effector proteins. Semin Cell Dev Biol 20:1055–1063
Li Y, Beisson F, Koo AJK, Molina I, Pollard M, Ohlrogge J (2007) Identification of acyltransferases required for cutin biosynthesis and production of cutin with suberin-like monomers. Proc Natl Acad Sci USA 104:18339–18344
Liu J, Liu X, Dai L, Wang G (2007) Recent progress in elucidating the structure, function and evolution of disease resistance genes in plants. J Genet Genomics 34:765–776
Liu W, Zhou X, Li G, Li L, Kong L, Wang C, Zhang H, Xu JR (2011) Multiple plant surface signals are sensed by different mechanisms in the rice blast fungus for appressorium formation. PLoS Pathog 7:e1001261
Lochman J, Mikes V (2006) Ergosterol treatment leads to the expression of a specific set of defense-related genes in tobacco. Plant Mol Biol 62:43–51
Lorenzo O, Solano R (2005) Molecular players regulating the jasmonate signaling network. Curr Opin Plant Biol 8:532–540
Lotze MT, Zeh HJ, Rubartelli A, Sparvero LJ, Amoscato AA, Washburn NR, DeVera ME, Liang X, Tör M, Billiar T (2007) The grateful dead: damage associated molecular pattern molecules and reduction/oxidation regulate immunity. Immunol Rev 220:60–81
Lukasik E, Takken FLW (2009) STANDing strong, resistance proteins instigators of plant defense. Curr Opin Plant Biol 12:427–436
Maheswaran G, Pridmore L, Franz P, Anderson MA (2007) A proteinase inhibitor from Nicotiana alata inhibits the normal development of light-brown apple moth, Epiphyas postvittana in transgenic apple plants. Plant Cell Rep 26:773–782
Manabe Y, Nafisi M, Verhertbruggen Y, Orfila C, Gille S, Rautengarten C, Cherk C, Marcus SE, Somerville S, Pauly M, Knox JP, Sakuragi Y, Scheller HV (2011) Loss-of-function mutation of REDUCED WALL ACETYLATION2 in Arabidopsis leads to reduced cell wall acetylation and increased resistance to Botrytis cinerea. Plant Physiol 155:1068–1078
Marshall A (2010) 2nd-generation GM traits progress. Nat Biotech 28:306
Mendoza AM, Berndt P, Djamei A, Linne U, Marahiel M, Vranes M, Kämper J, Kashmann R (2009) Physical-chemical plant-derived signals induce differentiation in Ustilago maydis. Mol Microbiol 71:895–911
Miao W, Wang X, Li M, Song C, Wang Y, Hu D, Wang J (2010) Genetic transformation of cotton with a harpin-encoding gene hpaXoo confers an enhanced defense response against different pathogens through a priming mechanism. BMC Plant Biol 10:67
Molina A, Garcia-Olmedo F (1997) Enhanced tolerance to bacterial pathogens caused by the transgenic expression of barley lipid transfer protein LTP2. Plant J 12:669–675
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation of 21st century crop improvement. Plant Physiol 147:969–977
Mourgues F, Brisset M-N, Chevreau E (1998) Strategies to improve plant resistance to bacterial diseases through genetic engineering. Trends Biotechnol 6:203–210
Mukhtar MS, Carvunis A-R, Dreze M, Epple P, Steinbrenner J, Moore J, Tasan M et al (2011) Independently evolved virulence effectors converge onto hubs in a plant immune system network. Science 333:596–601
Nishimura MT, Dangl JL (2010) Arabidopsis and the plant immune system. Plant J 61:1053–1066
Niu JH, Jian H, Xu JM, Guo YD, Liu Q (2010) RNAi technology extends its reach: engineering plant resistance against harmful eukaryotes. Afr J Biotechnol 9:7573–7582
Nowara D, Gay A, Lacomme C, Shaw J, Ridout C, Douchkov D, Hensel G, Kumlehn J, Schweizer P (2010) HIGS: Host-induced gene silencing in the obligate biotrophic fungal pathogen Blumeria graminis. Plant Cell 22:3130–3141
Ntui VO, Thirukkumaran G, Azadi P, Khan RS, Nakamura I, Mii M (2010) Stable integration and expression of wasabi defensin gene in “Egusi” melon (Colocynthis citrullus L.) confers resistance to Fusarium wilt and Alternaria leaf spot. Plant Cell Rep 29:943–954
Oh SK, Baek KH, Seong ES, Joung YH, Choi GJ, Park JM, Cho HS, Kim EA, Lee S, Choi D (2010) CaMsrB2, Pepper methionine sulfoxide reductase B2, is a novel defense regulator against oxidative stress and pathogen attack. Plant physiol 154:245–261
Osman H, Vauthrin S, Mikes V, Milat ML, Panabieres F, Marais A, Brunie S, Maume B, Ponchet M, Blein JP (2001) Mediation of elicitin activity on tobacco is assumed by elicitin-sterol complexes. Mol Biol Cell 12:2825–2834
Papadopoulou K, Melton RE, Leggett M, Daniels MJ, Osbourn AE (1999) Compromised disease resistance in saponin-deficient plants. Proc Natl Acad Sci USA 96:12923–12928
Parkhi V, Kumar V, Campbell LM, Bell AA, Shah J, Rathore KS (2010) Resistance against various fungal pathogens and reniform nematode in transgenic cotton plants expressing Arabidopsis NPR1. Transgenic Res 19:959–975
Patkar RN, Chattoo BB (2006) Transgenic indica rice expressing ns-LTP-like protein shows enhanced resistance to both fungal and bacterial pathogens. Mol Breed 17:159–171
Pavan S, Jacobsen E, Visser RGF, Bai Y (2010) Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol Breed 25:1–12
Pieterse CMJ, Leon-Reyes A, Van der Ent S, Van Wees SCM (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5:308–316
Piron F, Nicolai M, Minoïa S, Piednoir E, Moretti A, Salgues A, Zamir D, Caranta C, Bendahmane A (2010) An induced mutation in tomato elF4E leads to immunity to two potyviruses. PLoS One 5:e11313
Poland JA, Balint-Kurti PJ, Wisser RJ, Pratt RC, Nelson RJ (2009) Shades of gray: the world of quantitative disease resistance. Trends Plant Sci 14(1):21–29
Portieles R, Ayra C, Gonzalez E, Gallo A, Rodriguez R, Chacon O, Lopez Y, Rodriguez M, Castillo J, Pujol M, Enriquez G, Borroto C, Trujillo L, Thomma BP, Hidalgo OB (2010) NmDef02, a novel antimicrobial gene isolated from Nicotiana megalosiphon confers high-level pathogen resistance under greenhouse and field conditions. Plant Biotechnol J 8:678–690
Quilis J, Meynard D, Vila L, Aviles FX, Guiderdoni E, Segundo BS (2007) A potato carboxypeptidase inhibitor gene provides pathogen resistance in transgenic rice. Plant Biotechnol J 5:537–553
Rahnamaeian M, Langen G, Imani J, Khalifa W, Altincicek B, von Wettstein D, Kogel KH, Vilcinskas A (2009) Insect peptide metchnikowin confers on barley a selective capacity for resistance to fungal ascomycetes pathogens. J Exp Bot 60:4105–4114
Ramirez V, Agorio A, Coego A, Andrade JG, Hernandez MJ, Balaguer B, Ouwerkerk PBF, Zarra I, Vera P (2011) MYB46 modulates disease susceptibility to Botrytis cinerea in Arabidopsis. Plant Physiol 155:1920–1935
Reina-Pinto JJ, Yephremov A (2009) Surface lipids and plant defenses. Plant Physiol Biochem 47:540–549
Robaglia C, Caranta C (2006) Translation initiation factors: a weak link in plant RNA virus infection. Trends Plant Sci 11:40–45
Roberts JK, Pitkin JW, Adams TH. 2008; USA patent publication no. 2008/0022423.
Roberts-Seilaniantz A, Navarro L, Bari R, Jones JDG (2007) Pathological hormone imbalances. Curr Opin Plant Biol 10:372–379
Ron M, Avni A (2004) The receptor for the fungal elicitor ethylene-inducing xylanase is a member of a resistance-like gene family in tomato. Plant Cell 16:1604–1615
Saladie M, Matas AJ, Isaacson T, Jenks MA, Goodwin SM, Niklas KJ, Xiaolin R, Labavitch JM, Shackel KA, Fernie AR, Lytovchenko A, O’Neill MA, Watkins CB, Rose JKC (2007) The reevaluation of the key factors that influence tomato fruit softening and integrity. Plant Physiol 144:1012–1028
Sanahuja G, Banakar R, Twyman RM, Capell T, Christou P (2011) Bacillus thuringiensis: a century of research, development and commercial applications. Plant Biotechnol 9:283–300
Schlüter U, Benchabane M, Munger A, Kiggundu A, Vorster J, Goulet M-C, Cloutier C, Michaud D (2010) Recombinant protease inhibitors for herbivore pest control: a multitrophic perspective. J Exp Bot 61:4169–4183
Sengupta S, Chakraborti D, Mondal HA, Das S (2010) Selectable antibiotic resistance marker gene-free transgenic rice harbouring the garlic leaf lectin gene exhibits resistance to sap-sucking plant hoppers. Plant Cell Rep 29:261–271
Shah AD, Ahmed M, Mukhtar Z, Khan SA, Habib I, Malik ZA, Mansoor S, Saeed NA (2011) Spider toxin (Hvt) gene cloned under phloem specific RSs1 and RolC promoters provides resistance against American bollworm (Heliothis armigera). Biotechnol Lett 33:1457–1463
Shah JM, Raghupathy V, Veluthambi K (2009) Enhanced sheath blight resistance in transgenic rice expressing an endochitinase gene from Trichoderma virens. Biotechnol Lett 31:239–244
Shao M, Wang J, Dean RA, Lin Y, Gao X, Hu S (2008) Expression of a harpin-encoding gene in rice confers durable nonspecific resistance to Magnaporthe grisea. Plant Biotechnol J 6:73–81
Shepherd RW, Wagner GJ (2007) Phylloplane proteins: emerging defenses at the aerial frontline? Trends Plant Sci 12:51–56
Shirasu K (2008) The HSP90-SGT1 chaperone complex for NLR immune sensors. Annu Rev Plant Biol 60:139–164
Shin S, Mackintosh CA, Lewis J, Heinen SJ, Radmer L, Macky RD, Baldridge GD, Zeyen RJ, Muehlbauer GJ (2008) Transgenic wheat expressing a barley class II chitinase gene has enhanced resistance against Fusarium graminearum. J Exp Bot 59:2371–2378
Shoresh M, Harman GE, Mastouri F (2010) Induced systemic resistance and plant responses to fungal biocontrol agents. Annu Rev Phytopathol 48:21–43
Shukurov R, Voblikova V, Nikonorova AK, Komakhin RA, Komakhina V, Egorov T, Grishin E, Babakov A (2012) Transformation of tobacco and Arabidopsis plants with Stellaria media genes encoding novel hevein-like peptides increases their resistance to fungal pathogens. Transgenic Res 21:313–325
Solleti SK, Bakshi S, Purkayastha J, Panda SK, Sahoo L (2008) Transgenic cowpea (Vigna unguiculata) seeds expressing a bean alpha-amylase inhibitor 1 confer resistance to storage pests, bruchid beetles. Plant Cell Rep 27:1841–1850
Spoel SH, Dong X (2008) Making sense of hormone crosstalk during plant immune responses. Cell Host Microbe 3:348–351
Srinivasan T, Kumar KRR, Kirti PB (2009) Constitutive expression of a trypsin protease inhibitor confers multiple stress tolerance in transgenic tobacco. Plant Cell Physiol 50:541–553
Stassen JHM, Van den Ackerveken G (2011) How do oomycete effector interfere with plant life? Curr Opin Plant Biol 14:407–414
Sujatha M, Lakshminarayana M, Tarakeswari M, Singh PK, Tuli R (2009) Expression of the cry1EC gene in castor (Ricinus communis L.) confers field resistance to tobacco caterpillar (Spodoptera litura Fabr) and castor semilooper (Achoea janata L.). Plant Cell Rep 28:935–946
Takai R, Isogai A, Seiji S, Che FS (2008) Analysis of flagellin perception mediated by flg22 receptor OsFLS2 in rice. Mol Plant Microbe Interact 12:1635–1642
Takakura YY, Oka NN, Suzuki JJ, Tsukamoto HH, Ishida YY (2012) Intercellular production of tamavidin 1, a biotin-binding protein from Tamogitake mushroom, confers resistance to the blast fungus Magnaporthe oryzae in transgenic rice. Mol Biotechnol 51:9–17
Tang D, Simonich MT, Innes RW (2007) Mutations in LACS2, a long-chain acyl-coenzyme A synthetase, enhance susceptibility to avirulent Pseudomonas syringae but confer resistance to Botrytis cinerea in Arabidopsis. Plant Physiol 144:1093–1103
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
Thatcher LF, Anderson JP, Singh KB (2005) Plant defense responses: what have we learnt from Arabidopsis? Funct Plant Biol 32:1–19
Thomma BPHJ, Nürnberger T, Joosten MHAJ (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23:4–15
Thordal-Christensen H (2003) Fresh insights into processes of nonhost resistance. Curr Opin Plant Biol 6:351–357
Tinoco MLP, Dias BBA, Dall’Astta RC, Pamphile JA, Aragäo FJL (2010) In vivo trans-specific gene silencing in fungal cells by in planta expression of a double-stranded RNA. BMC Biol 8:27
Ton J, Flors V, Mauch-Mani B (2009) The multifaceted role of ABA in disease resistance. Trends Plant Sci 14:310–317
Torres AM (2010) Application of molecular markers for breeding disease resistant varieties in crop plants. In: Jain SM, Brar DS (eds.) Molecular techniques in crop improvement. Springer Science, Dordrecht, pp. 185–205
Tripathi L, Mwaka H, Tripathi JN, Tushemereirwe WK (2010) Expression of sweet pepper Hrap gene in banana enhances resistance to Xanthomonas campestris pv. musacearum. Mol Plant Pathol 11:721–731
van de Craen M, Goh PY, Logghe MG, Khu YL, Mortier K, Bogaert TAOE (2006) USA Patent Publication No. 2006/0247197A1
van der Hoom RAL, Kamoun S (2008) From guard to decoy: a new model for perception of plant pathogen effectors. Plant Cell 20:2009–2017
Varshney RK, Graner A, Sorrells ME (2005) Genomics-assisted breeding for crop improvement. Trends Plant Sci 10:621–630
Varshney RK, Nayak SN, May GD, Jackson SA (2010) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27:522–530
Varshney RK, Bansal KC, Aggarwal PK, Datta SK, Craufurd PQ (2011) Agricultural biotechnology for crop improvement in a variable climate: hope or hype? Trends Plant Sci 16:363–371
Venter M (2006) Synthetic promoters: genetic control through cis engineering. Trends Plant Sci 12:118–124
Vijayan S, Kirti PB (2012) Mungbean plants expressing BjNPR1 exhibit enhanced resistance against the seedling rot pathogen, Rhizoctonia solani. Transgenic Res 21:193–200
Vlot AC, Klessig DF, Park S-W (2008) Systemic acquired resistance: the elusive signal(s). Curr Opin Plant Biol 11:436–442
Vogel JP, Somerville SC (2000) Isolation and characterization of powdery mildew-resistant Arabidopsis mutants. Proc Natl Acad Sci USA 97:1897–1902
Vogel JP, Raab TK, Schiff C, Somerville SC (2002) PMR6, a pectate lyase-like gene required for powdery mildew susceptibility in Arabdiopsis. Plant Cell 14:1–13
Volpi C, Janni M, Lionetti V, Bellincampi D, Favaron F, D’Ovidio R (2011) The ectopic expression of a pectin methyl esterase inhibitor increases pectin methyl esterification and limits fungal diseases in wheat. Mol Plant Microbe Interact 24:1012–1019
Wally O, Jayaraj J, Punja ZK (2009) Broad-spectrum disease resistance to necrotrophic and biotrophic pathogens in transgenic carrots (Daucus carota L.) expressing an Arabidopsis NPR1 gene. Planta 231:131–141
Wan J, Zhang XC, Neece D, Ramonell KM, Clough S, Kim SY, Stacey MG, Stacey G (2008) A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20:471–481
Wang C, Chin CK, Chen A (1998) Expression of the yeast Δ-9 desaturase gene in tomato enhances its resistance to powdery mildew. Physiol Mol Plant Pathol 52:371–383
Wang C, Chin CK, Gianfagna T (2000) Relationship between cutin monomers and tomato resistance to powdery mildew infection. Physiological Mol Plant Pathol 57:55–61
Wang Z, Mao H, Dong C, Ji R, Cai L, Fu H, Liu S (2009) Overexpression of Brassica napus MPK4 enhances resistance to Sclerotinia sclerotiorum in oilseed rape. Mol Plant Microbe Interact 22:235–244
Wani SH, Sanghera GS, Singh NB (2010) Biotechnology and plant disease control-role of RNA interference. Am J Plant Sci 1:55–68
Weinthal D, Tovkach A, Zeevi V, Tzfira T (2010) Genome editing in plant cells by zinc finger nucleases. Trends Plant Sci 15:308–321
Weng LX, Deng HH, Xu JL, Li Q, Zhang YQ, Jiang ZD, Li QW, Chen JW, Zhang LH (2011) Transgenic sugarcane plants expressing high levels of modified cry1Ac provide effective control against stem borers in field trials. Transgenic Res 20:759–772
Wenzel G (2006) Molecular plant breeding: achievements in green biotechnology and future perspectives. Appl Microbiol Biotechnol 70:642–650
Wulff BBH, Horvath DM, Ward ER (2011) Improving immunity in crops: new tactics in an old game. Curr Opin Plant Biol 14:468–476
Xia Y, Gao QM, Yu K, Lapchyk L, Navarre D, Hildebrand D, Kachroo A, Kachroo P (2009) An intact cuticle in distal tissues is essential for the induction of systemic acquired resistance in plants. Cell Host Microbe 5:151–165
Xia Y, Yu K, Navarre D, Seebold K, Kachroo A, Kachroo P (2010) The glabra1 mutation affects cuticle formation and plant responses to microbes. Plant Physiol 154:833–846
Xiao F, Goodwin SM, Xiao Y, Sun Z, Baker D, Tang X, Jenks MA, Zhou JM (2004) Arabidopsis CYP86A2 represses Pseudomonas syringae type III genes and is required for cuticle development. EMBO J 23:2903–2913
Xiao YH, Li XB, Yang XY, Luo M, Hou L, Guo SH, Luo XY, Pei Y (2007) Cloning and characterization of a balsam pear class I chitinase gene (Mcchit1) and its ectopic expression enhances fungal resistance in transgenic plants. Biosci Biotechnol Biochem 71:1211–1219
Yamaguchi T, Yamada A, Hong N, Ogawa T, Ishii T, Shibuya N (2000) Differences in the recognition of glucan elicitors signals between rice and soybean: β-glucan fragments from the rice blast disease fungus Pyricularia oryzae that elicit phytoalexin biosynthesis in suspension cultured rice cells. Plant Cell 12:817–826
Yang S, Gao M, Xu C, Gao J, Deshpande S, Lin S, Roe BA, Zhu H (2008) Alfalfa benefits from Medicago truncatula: The RCT1 gene from M. truncatula confers broad-spectrum resistance to anthracnose in alfalfa. Proc Natl Acad Sci USA 105:12164-12169
Yarasi B, Sadumpati V, Immanni CP, Vudem DR, Khareedu VR (2008) Transgenic rice expressing Allium sativum leaf agglutinin (ASAL) exhibits high-level resistance against major sap-sucking pests. BMC Plant Biol 8:102
Ye SH, Chen S, Zhang F, Wang W, Tian Q, Liu JZ, Chen F, Bao JK (2009) Transgenic tobacco expressing Zephyranthes grandiflora agglutinin confers enhanced resistance to aphids. Appl Biochem Biotechnol 158:615–630
Yevtushenko DP, Misra S (2007) Comparison of pathogen-induced expression and efficacy of two amphibian antimicrobial peptides, MsrA2 and temporin A, for engineering wide-spectrum disease resistance in tobacco. Plant Biotechnol J 5:720–734
Yin C, Jurgenson JE, Hulbert SH (2011) Development of a host-induced RNAi system in the wheat stripe rust fungus Puccinia striiformis f. sp. tritici. Mol Plant Microbe Interact 24:554–561
Zhang J, Zhou J-M (2010) Plant immunity triggered by microbial molecular signatures. Mol Plant 3:783–793
Zhu YJ, Agbayani R, Moore PH (2007) Ectopic expression of Dahlia merckii defensin DmAMP1 improves papaya resistance to Phytophthora palmivora by reducing pathogen vigor. Planta 226:87–97
Zipfel C (2009) Early molecular events in PAMP-triggered immunity. Curr Opin Plant Biol 12:414–420
Zipfel C, Robatzek S (2010) Pathogen-associated molecular pattern-triggered immunity: Veni, vidi…? Plant Physiol 154:551–554
Acknowledgements
This work is supported partially by research grant provided by Department of Biotechnology, Government of India and National Institute of Plant Genome Research, New Delhi. We acknowledge Dr. K. D. Srivastava, Indian Agricultural Research Institute, New Delhi for valuable suggestions and critically editing the manuscript. K.K. acknowledges NIPGR for postdoctoral fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Kumar, K., Verma, P.K. (2013). Plant Pathogen Interactions: Crop Improvement Under Adverse Conditions. In: Tuteja, N., Singh Gill, S. (eds) Plant Acclimation to Environmental Stress. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-5001-6_16
Download citation
DOI: https://doi.org/10.1007/978-1-4614-5001-6_16
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-5000-9
Online ISBN: 978-1-4614-5001-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)