Abstract
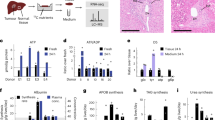
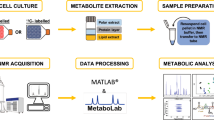
Human metabolic liver disease is dramatically increasing globally and presents an urgent clinical unmet need. Rodent models of non-alcoholic fatty liver disease (NAFLD) are available, but they fail to fully recreate the metabolic and cellular features of human disease. Thus, it is imperative to understand the metabolic interplay in human cells in the context of disease. We have applied nuclear magnetic resonance (NMR) spectroscopy approaches to enable the detection of numerous metabolites in human cells and within intact tissue in a single measurement. In this chapter, we describe the challenges of using isolated human hepatocytes vs perfused human liver tissue for metabolic tracer experiments and how experimental parameters can be refined to interrogate signals from intact tissue and cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Duarte IF (2011) Following dynamic biological processes through NMR-based metabonomics: a new tool in nanomedicine? J Control Release 153:34–39
Giraudeau P (2020) NMR-based metabolomics and fluxomics: developments and future prospects. Analyst 145:2457–2472
Beckonert O, Keun HC, Ebbels TM et al (2007) Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue. Nat Protoc 2:2692–2703
Carrigan JB, Reed MA, Ludwig C et al (2016) Tracer-based metabolic NMR-based flux analysis in a leukaemia cell line. ChemPlusChem 81:453–459
Reed MC, Roberts J, Gierth P et al (2019) Quantitative isotopomer rates in real-time metabolism of cells determined by NMR methods. Chembiochem 20:2207–2211
Saborano R, Eraslan Z, Roberts J et al (2019) A framework for tracer-based metabolism in mammalian cells by NMR. Sci Rep 9:2520
Schofield Z, Reed MA, Newsome PN et al (2017) Changes in human hepatic metabolism in steatosis and cirrhosis. World J Gastroenterol 23:2685–2695
Sheriff L, Khan RS, Saborano R et al (2020) Alcoholic hepatitis and metabolic disturbance in female mice: a more tractable model than Nrf2−/− animals. Dis Model Mech 13:dmm046383
Shepherd EL, Saborano R, Northall E et al (2021) Ketohexokinase inhibition improves NASH by reducing fructose-induced steatosis and fibrogenesis. JHEP Rep 3:100217
Izumi Y, Taniuchi Y, Tsuji T et al (1995) Characterization of human colon-carcinoma variant cells selected for sialyl le(x) carbohydrate antigen – liver colonization and adhesion to vascular endothelial-cells. Exp Cell Res 216:215–221
Andrews RC, Rooyackers O, Walker BR (2003) Effects of the 11 beta-hydroxysteroid dehydrogenase inhibitor carbenoxolone on insulin sensitivity in men with type 2 diabetes. J Clin Endocrinol Metab 88:285–291
Ludwig C, Gunther UL (2011) MetaboLab – advanced NMR data processing and analysis for metabolomics. BMC Bioinformatics 12:366
Gunther UL, Ludwig C, Ruterjans H (2000) NMRLAB – advanced NMR data processing in matlab. J Magn Reson 145:201–208
Wishart DS, Feunang YD, Marcu A et al (2018) HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res 46:D608–D617
Delaglio F, Grzesiek S, Vuister GW et al (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Hyberts SG, Milbradt AG, Wagner AB et al (2012) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson gap scheduling. J Biomol NMR 52:315–327
Bhogal RH, Hodson J, Bartlett DC et al (2011) Isolation of primary human hepatocytes from normal and diseased liver tissue: a one hundred liver experience. PLoS One 6:e18222
Mergental H, Laing RW, Kirkham AJ et al (2020) Transplantation of discarded livers following viability testing with normothermic machine perfusion. Nat Commun 11:2939
Acknowledgments
This study includes independent research supported by the Birmingham National Institute for Health and Care Research (NIHR) Birmingham Biomedical Research Centre, based at the University of Birmingham. The views expressed are those of the authors and not necessarily those of the NHS, the National Institute for Health and Care Research, or the Department of Health and Social Care. We are grateful to the physicians and patients at the Queen Elizabeth Hospital in Birmingham who donated their tissue for our investigations.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Saborano, R., Shepherd, E., Günther, U.L., Lalor, P.F. (2023). Tracer-Based Metabolic Analysis by NMR in Intact Perfused Human Liver Tissue. In: Papa, S., Bubici, C. (eds) Metabolic Reprogramming. Methods in Molecular Biology, vol 2675. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3247-5_13
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3247-5_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3246-8
Online ISBN: 978-1-0716-3247-5
eBook Packages: Springer Protocols