Abstract
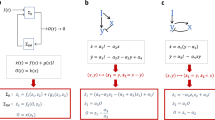
Biochemical networks are often characterized by tremendous complexity—both in terms of the sheer number of interacting molecules (“nodes”) and in terms of the varied and incompletely understood interactions among these molecules (“interconnections” or “edges”). Strikingly, the vast and intricate networks of interacting proteins that exist within each living cell have the capacity to perform remarkably robustly, and reproducibly, despite significant variations in concentrations of the interacting components from one cell to the next and despite mutability over time of biochemical parameters. Here we consider the ubiquitously observed and fundamentally important signalling response known as robust perfect adaptation (RPA). We have recently shown that all RPA-capable networks, even the most complex ones, must satisfy an extremely rigid set of design principles, and are modular, being decomposable into just two types of network building-blocks—opposer modules and balancer modules. Here we present an overview of the design principles that characterize all RPA-capable network topologies through a detailed examination of a collection of simple examples. We also introduce a diagrammatic method for studying the potential of a network to exhibit RPA, which may be applied without a detailed knowledge of the complex mathematical principles governing RPA.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wagner A (2013) Robustness and evolvability in living systems, vol 24. Princeton University Press
Araujo RP, Liotta LA, Petricoin EF (2007) Proteins, drug targets and the mechanisms they control: the simple truth about complex networks. Nat Rev Drug Discov 6(11):871–880
Whitacre JM (2012) Biological robustness: paradigms, mechanisms, and systems principles. Front Genet 3:67
Barkai N, Leibler S (1997) Robustness in simple biochemical networks. Nature 387(6636):913–917
Araujo RP, Liotta LA (2018) The topological requirements for robust perfect adaptation in networks of any size. Nat Commun 9(1):1–12
Yi TM, Huang Y, Simon MI, Doyle J (2000) Robust perfect adaptation in bacterial chemotaxis through integral feedback control. Proc Natl Acad Sci 97(9):4649–4653
Hoeller O, Gong D, Weiner OD (2014) How to understand and outwit adaptation. Dev Cell 28(6):607–616
Alon U, Surette MG, Barkai N, Leibler S (1999) Robustness in bacterial chemotaxis. Nature 397(6715):168–171
Muzzey D, Gómez-Uribe CA, Mettetal JT, van Oudenaarden A (2009) A systems-level analysis of perfect adaptation in yeast osmoregulation. Cell 138(1):160–171
El-Samad H, Goff JP, Khammash M (2002) Calcium homeostasis and parturient hypocalcemia: an integral feedback perspective. J Theor Biol 214(1):17–29
Kaupp UB (2010) Olfactory signalling in vertebrates and insects: differences and commonalities. Nat Rev Neurosci 11(3):188–200
Yau KW, Hardie RC (2009) Phototransduction motifs and variations. Cell 139(2):246–264
Ben-Zvi D, Barkai N (2010) Scaling of morphogen gradients by an expansion-repression integral feedback control. Proc Natl Acad Sci 107(15):6924–6929
Eldar A, Dorfman R, Weiss D, Ashe H, Shilo BZ, Barkai N (2002) Robustness of the BMP morphogen gradient in Drosophila embryonic patterning. Nature 419(6904):304–308
Yadid G, Overstreet DH, Zangen A (2001) Limbic dopaminergic adaptation to a stressful stimulus in a rat model of depression. Brain Res 896(1–2):43–47
Medina-Gomez G, Yetukuri L, Velagapudi V, Campbell M, Blount M, Jimenez-Linan M, Ros M, Orešič M, Vidal-Puig A (2009) Adaptation and failure of pancreatic β cells in murine models with different degrees of metabolic syndrome. Dis Model Mech 2(11–12):582–592
Sturgeon JA, Zautra AJ (2010) Resilience: a new paradigm for adaptation to chronic pain. Curr Pain Headache Rep 14(2):105–112
Fodale V, Pierobon M, Liotta L, Petricoin E (2011) Mechanism of cell adaptation: when and how do cancer cells develop chemoresistance? Cancer J 17(2):89–95
Araujo RP, Petricoin EF, Liotta LA (2007) Mathematical modeling of the cancer cell’s control circuitry: paving the way to individualized therapeutic strategies. Curr Signal Transduction Ther 2(2):145–155
Geho DH, Petricoin EF, Liotta LA, Araujo RP (2005) Modeling of protein signaling networks in clinical proteomics. In: Cold Spring Harbor symposia on quantitative biology, vol 70. Cold Spring Harbor Laboratory Press, pp 517–524
Ma W, Trusina A, El-Samad H, Lim WA, Tang C (2009) Defining network topologies that can achieve biochemical adaptation. Cell 138(4):760–773
Briat C, Gupta A, Khammash M (2016) Antithetic integral feedback ensures robust perfect adaptation in noisy biomolecular networks. Cell Syst 2(1):15–26
Aoki SK, Lillacci G, Gupta A, Baumschlager A, Schweingruber D, Khammash M (2019) A universal biomolecular integral feedback controller for robust perfect adaptation. Nature 570(7762):533–537
Qian Y, Del Vecchio D (2018) Realizing ‘integral control’ in living cells: how to overcome leaky integration due to dilution? J R Soc Interface 15(139):20170902
Del Vecchio D, Dy AJ, Qian Y (2016) Control theory meets synthetic biology. J R Soc Interface 13(120):20160380
Francis BA, Wonham WM (1975) The internal model principle for linear multivariable regulators. Appl Math Optim 2(2):170–194
Francis BA, Wonham WM (1976) The internal model principle of control theory. Automatica 12(5):457–465
Ferrell JE Jr (2016) Perfect and near-perfect adaptation in cell signaling. Cell Syst 2(2):62–67
Tyson JJ, Chen KC, Novak B (2003) Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Curr Opin Cell Biol 15(2):221–231
Shinar G, Feinberg M (2010) Structural sources of robustness in biochemical reaction networks. Science 327(5971):1389–1391
Xiao F, Doyle JC (2018, December) Robust perfect adaptation in biomolecular reaction networks. In: 2018 IEEE conference on decision and control (CDC). IEEE, pp 4345–4352
Acknowledgments
Robyn Araujo is the recipient of an Australian Research Council (ARC) Future Fellowship (project number FT190100645) funded by the Australian Government.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Araujo, R.P., Liotta, L.A. (2023). Design Principles Underlying Robust Adaptation of Complex Biochemical Networks. In: Nguyen, L.K. (eds) Computational Modeling of Signaling Networks. Methods in Molecular Biology, vol 2634. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3008-2_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3008-2_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3007-5
Online ISBN: 978-1-0716-3008-2
eBook Packages: Springer Protocols