Abstract
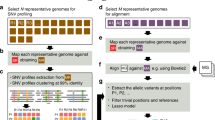
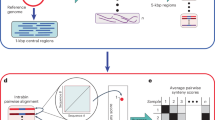
Microbial strains are interpreted as a lineage derived from a recent ancestor that have not experienced “too many” recombination events and can be successfully retrieved with culture-independent techniques using metagenomic sequencing. Such a strain variability has been increasingly shown to display additional phenotypic heterogeneities that affect host health, such as virulence, transmissibility, and antibiotics resistance. New statistical and computational methods have recently been developed to track the strains in samples based on shotgun metagenomics data either based on reference genome sequences or Metagenome-assembled genomes (MAGs). In this paper, we review some recent statistical methods for strain identifications based on frequency counts at a set of single nucleotide variants (SNVs) within a set of single-copy marker genes. These methods differ in terms of whether reference genome sequences are needed, how SNVs are called, what methods of deconvolution are used and whether the methods can be applied to multiple samples. We conclude our review with areas that require further research.
Short title: Resolving Microbial Strain Mixtures
This research is supported by NIH grants GM123056 and P30DK050306.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lloyd-Price J, Arze C, Ananthakrishnan AN et al (2019) Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569(7758):655–662
Zhou W, Sailani MR, Contrepois K et al (2019) Longitudinal multi-omics of host–microbe dynamics in prediabetes. Nature 569(7758):663–671
Van Rossum T, Ferretti P, Maistrenko OM, Bork P (2020) Diversity within species: interpreting strains in microbiomes. Nat Rev Microbiol 18(9):491–506
Fournier PE, Dubourg G, Raoult D (2014) Clinical detection and characterization of bacterial pathogens in the genomics era. Genome Med 6(11):1–15
Quince C, Nurk S, Raguideau S et al (2021) Strong: metagenomics strain resolution on assembly graphs. Genome Biol 22(1):1–34
van Dijk LR, Walker BJ, Straub TJ et al (2022) StrainGE: A toolkit to track and characterize low-abundance strains in complex microbial communities. Genome Biol 23(1):1–27
Smith BJ, Li X, Abate A et al (2022) Scalable microbial strain inference in metagenomic data using StrainFacts. bioRxiv
Lloyd-Price J, Mahurkar A, Rahnavard G et al (2017) Strains, functions and dynamics in the expanded human microbiome project. Nature 550(7674):61–66
Smillie CS, Sauk J, Gevers D et al (2018) Strain tracking reveals the determinants of bacterial engraftment in the human gut following fecal microbiota transplantation. Cell Host Microbe 23(2):229–240
Zhou W, Spoto M, Hardy R et al (2020) Host-specific evolutionary and transmission dynamics shape the functional diversification of staphylococcus epidermidis in human skin. Cell 180(3):454–470
Garrity GM, Parker CT, Tindall BJ (2015) International code of nomenclature of prokaryotes. Int J Syst Evol Microbiol 90(6)
Jain C, Rodriguez-R LM, Phillippy AM et al (2018) High throughput ani analysis of 90k prokaryotic genomes reveals clear species boundaries. Nat Commun 9(1):1–8
Yan Y, Nguyen LH, Franzosa EA, Huttenhower C (2020) Strain-level epidemiology of microbial communities and the human microbiome. Genome Med 12(1):1–16
Segata N, Waldron L, Ballarini A et al (2012) Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods 9(8):811–814
Wood DE, Salzberg SL (2014) Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol 15(3):1–12
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) Dada2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583
Brenner DJ, Fanning G, Steigerwalt A et al (1972) Polynucleotide sequence relatedness among three groups of pathogenic Escherichia coli strains. Infect Immun 6(3):308–315
Truong DT, Tett A, Pasolli E et al (2017) Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res 27(4):626–638
Albanese D, Donati C (2017) Strain profiling and epidemiology of bacterial species from metagenomic sequencing. Nat Commun 8(1):1–14
Li X, Saadat S, Hu H, Li X (2019) BHap: a novel approach for bacterial haplotype reconstruction. Bioinformatics 35(22):4624–4631
Anyansi C, Straub TJ, Manson AL et al (2020) Computational methods for strain-level microbial detection in colony and metagenome sequencing data. Front Microbiol 11:1925
Garg S (2021) Computational methods for chromosome-scale haplotype reconstruction. Genome Biol 22(1):1–24
Ghazi AR, Münch PC, Chen D et al (2022) Strain identification and quantitative analysis in microbial communities. J Mol Biol, 167582
Quince C, Delmont TO, Raguideau S et al (2017) Desman: a new tool for de novo extraction of strains from metagenomes. Genome Biol 18(1):1–22
Pasolli E, Asnicar F, Manara S et al (2019) Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176(3):649–662
Loh PR, Danecek P, Palamara PF et al (2016) Reference-based phasing using the haplotype reference consortium panel. Nat Genet 48(11):1443–1448
Scholz M, Ward DV, Pasolli E et al (2016) Strain-level microbial epidemiology and population genomics from shotgun metagenomics. Nat Methods 13(5):435–438
Wang S, Jiang Y, Li S (2021) Pstrain: an iterative microbial strains profiling algorithm for shotgun metagenomic sequencing data. Bioinformatics 36(22–23):5499–5506
Moon TK (1996) The expectation-maximization algorithm. IEEE Signal Process Mag 13(6):47–60
Cowles MK, Carlin BP (1996) Markov chain Monte Carlo convergence diagnostics: a comparative review. J Am Stat Assoc 91(434):883–904
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18(5):821–829
Dilthey AT, Jain C, Koren S, Phillippy AM (2019) Strain-level metagenomic assignment and compositional estimation for long reads with MetaMaps. Nat Commun 10(1):1–12
Luo C, Knight R, Siljander H et al (2015) Constrains identifies microbial strains in metagenomic datasets. Nat Biotechnol 33(10):1045–1052
Sakamoto Y, Ishiguro M, Kitagawa G (1986) Akaike information criterion statistics. Dordrecht, The Netherlands: D Reidel 81(10.5555):26853
Li H (2015) Microbiome, metagenomics, and high-dimensional compositional data analysis. Annual Review of Statistics and Its Application 2:73–94. https://doi.org/10.1146/annurevstatistics010814020351
Sinha R, Abu-Ali G, Vogtmann E et al (2017) Assessment of variation in microbial community amplicon sequencing by the microbiome quality control (MBQC) project consortium. Nat Biotechnol 35(11):1077–1086
Neale B, Rivas M, Voight B et al (2011) Testing for an unusual distribution of rare variants. PLoS Genet 7(3):e1001322
Wu M, Lee S, Cai T et al (2011) Rare-variant association testing for sequencing data with the sequence kernel association test. Am J Hum Genet 89:82–93
Moss EL, Maghini DG, Bhatt AS (2020) Complete, closed bacterial genomes from microbiomes using nanopore sequencing. Nat Biotechnol 38:701–707
Vicedomini R, Quince C, Darling AE, Chikhi R (2021) Strainberry: automated strain separation in low-complexity metagenomes using long reads. Nat Commun 12:4485
Emiola A, Zhou W, Oh J (2020) Metagenomic growth rate inferences of strains in situ. Sci Adv 6(17):eaaz2299
Acknowledgements
The authors would like to thank Andrew Ghazi, Phillip Münch, Di Chen, Jordan Jensen, Yancong Zhang, Curtis Huttenhower, and Christopher Quince for their helpful input and discussions on this review.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ma, S., Li, H. (2023). Statistical and Computational Methods for Microbial Strain Analysis. In: Fridley, B., Wang, X. (eds) Statistical Genomics. Methods in Molecular Biology, vol 2629. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2986-4_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2986-4_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2985-7
Online ISBN: 978-1-0716-2986-4
eBook Packages: Springer Protocols