Abstract
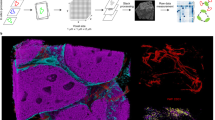
In situ profiling of the tumor-immune microenvironment (TiME) requires the ability to co-localize and detect multiple proteins simultaneously. Imaging mass cytometry (IMC), using the Hyperion™ imaging system is a novel multiplex imaging modality that currently enables detection of up to 50 markers on fixed tissues at subcellular resolution and thus has the potential to inform both pre-clinical and clinical research by providing investigators with spatially resolved information about the TiME. Here we provide an overview of the IMC workflow from sample fixation to analysis, with a focus on multiplex panel design and tissue staining.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Giesen C, Wang HA, Schapiro D et al (2014) Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat Methods 11:417–422
Leipold MD, Maecker HT (2012) Mass cytometry: protocol for daily tuning and running cell samples on a CyTOF mass cytometer. J Vis Exp 69:e4398
Chevrier S, Crowell HL, Zanotelli VRT et al (2018) Compensation of signal spillover in suspension and imaging mass cytometry. Cell Syst 6:612–20 e5
Takahashi C, Au-Yeung A, Fuh F et al (2017) Mass cytometry panel optimization through the designed distribution of signal interference. Cytometry A 91:39–47
Elaldi R, Hemon P, Petti L et al (2021) High dimensional imaging mass cytometry panel to visualize the tumor immune microenvironment contexture. Front Immunol 12:666233
Jackson HW, Fischer JR, Zanotelli VRT et al (2020) The single-cell pathology landscape of breast cancer. Nature 578:615–620
Thirumal S, Jamzad A, Cotechini T et al (2022) TITAN: an end-to-end data analysis environment for the Hyperion imaging system. Cytometry A 101:423–433
Economou M, Schoni L, Hammer C et al (2014) Proper paraffin slide storage is crucial for translational research projects involving immunohistochemistry stains. Clin Transl Med 3:4
DiVito KA, Charette LA, Rimm DL et al (2004) Long-term preservation of antigenicity on tissue microarrays. Lab Investig 84:1071–1078
Schapiro D, Jackson HW, Raghuraman S et al (2017) histoCAT: analysis of cell phenotypes and interactions in multiplex image cytometry data. Nat Methods 14:873–876
Greenwald NF, Miller G, Moen E et al (2021) Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat Biotechnol 40:555–565
Xiao X, Qiao Y, Jiao Y et al (2021) Dice-XMBD: deep learning-based cell segmentation for imaging mass cytometry. Front Genet 12:721229
Wang Z (2019) Cell segmentation for image cytometry: advances, insufficiencies, and challenges. Cytometry A 95:708–711
Zanotelli V, Bodenmiller B (2017) ImcSegmentationPipeline: A pixelclassification based multiplexed image segmentation pipeline. Zenodo. https://doi.org/10.5281/zenodo.3841961
Eling N, Damond N, Hoch T et al (2020) Cytomapper: an R/bioconductor package for visualisation of highly multiplexed imaging data. Bioinformatics 36:5706–5708
Levine JH, Simonds EF, Bendall SC et al (2015) Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162:184–197
Palla G, Spitzer H, Klein M et al (2022) Squidpy: a scalable framework for spatial omics analysis. Nat Methods 19:171–178
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Cotechini, T., Jones, O., Hindmarch, C.C.T. (2023). Imaging Mass Cytometry in Immuno-Oncology. In: Ursini-Siegel, J. (eds) The Tumor Microenvironment. Methods in Molecular Biology, vol 2614. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2914-7_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2914-7_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2913-0
Online ISBN: 978-1-0716-2914-7
eBook Packages: Springer Protocols