Abstract
Cancer-derived organoids and three-dimensional (3D) extracellular matrix (ECM) are taking center stage as in vitro models to study neoplastic cell behavior, since they recapitulate the heterogeneous cellular composition of tumors and their extracellular environment. In combination with imaging and molecular/biochemical techniques, 3D organoid models have contributed substantially to our knowledge about the cellular and molecular mechanisms that regulate the growth of tumors and invasion into the surrounding tissue. We here outline a set of protocols that describe culturing of cancer-derived organoids in 3D matrices and various strategies that allow modeling of tumor growth, tumor cell penetration into basement membranes, and invasion into Collagen I-rich ECM. Furthermore, we specify protocols for subsequent handling of organoids cultured in 3D ECM for confocal microscopy and analysis of gene expression at the protein and mRNA level. Although we here use breast cancer-derived organoids, these protocols can be directly applied or adapted for organoids derived from other cancer types or healthy tissues. Thus, in addition to investigating cell behavior of multiple cancer types, the combination of protocols described here may be used to study processes such as cell differentiation and migration during homeostasis and normal development.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- Cancer cell invasion
- Extracellular matrix
- ECM remodeling
- 3D cultures
- Organoids
- Immunofluorescence
- Confocal microscopy
- Protein extraction
- RNA extraction
1 Introduction
The extracellular matrix (ECM) is a network of proteins and proteoglycans that forms a major component of the tumor microenvironment. In addition to providing structural support, the ECM is a source of biochemical and physical cues controlling tumor growth and invasion into the peritumor tissue [1]. Besides the glycocalyx, tumor cells are in contact with two major types of ECM, the basement membrane and interstitial ECM [2]. The basement membrane is a thin layer of ECM, composed of several molecules including laminins and Collagen IV. The basement membrane surrounds most epithelia and separates the cells from the surrounding interstitial ECM [3]. The interstitial ECM is predominantly composed of the fibrillar collagen, Collagen I, and includes other molecules such as fibronectin and elastin [2, 4]. In epithelial tumors (including breast, colon, and head and neck squamous cell carcinomas), the presence of an intact basement membrane is a favorable prognostic factor. Tumors that retain basement membranes are confined; they show no to little invasion into the surrounding interstitial ECM [3, 5, 6]. Cancer cell invasion starts with the penetration (breaching) of the basement membrane whereafter cells become in contact with the Collagen I-containing interstitial matrix. It is now well established that contact with Collagen I induces many cancer cell types to become invasive and to spread into the tissue [7,8,9]. Particularly, elevated Collagen I density and mechanical stiffness/tension drive protrusive cell behavior and motility and are associated with metastasis in cancer patients [10,11,12,13].
Modeling cellular processes such as tumor growth and invasion in vitro has improved substantially with the establishment of cancer-derived organoids growing in 3D ECM. Organoids derived from tumors maintain (at least in part) the heterogeneous cellular composition of the tissues they originate from [14, 15]. This is in contrast to established cell lines that lose cellular heterogeneity due to long-term culturing [16]. In addition to the cellular component, the biophysical properties of the ECM are essential elements of a (patho) physiologically relevant in vitro model. Reconstituted ECM such as basement membrane extract (BME, or Matrigel) and Collagen I are now used as 3D lattices to embed cells in. Compared to the conventional 2D rigid ECM-coated surfaces, 3D ECM better mimics the topology and biomechanics of ECM in tissues [17, 18]. Many 3D patient-derived organoid models have been shown to recapitulate tumor characteristics in patients including cancer cell survival, proliferation, and resistance to therapy [14, 15, 19, 20]. Furthermore, 3D Collagen I-based models recapitulate both the modes by which cancer cells invade into the tissues as well as the invasion-associated remodeling of the ECM [18, 21,22,23,24,25,26]. In addition to enzymatic modifications (ex. degradation and crosslinking), ECM remodeling includes ECM pulling, resulting in the alignment of Collagen I fibers to orientations that are parallel to the direction of invasion. Such tumor-associated Collagen I signatures represent histopathological markers for invasive cancers (incl. breast and pancreas) and associate with poor clinical outcome [10, 13]. Consequently, models using cancer-derived organoids in 3D ECM are becoming more frequently used in cancer research.
We here describe various procedures to model in vitro tumor formation and growth in ECM rich in basement membrane proteins (Fig. 1a, b), breaching of the basement membrane (Fig. 2a, b), and invasion into Collagen I-rich ECM with different biomechanical properties (Fig. 2a, c–f). Moreover, we describe techniques for fixation and subsequent immunostaining that maintain ECM architecture and secure efficient antibody penetration and imaging of cells that are located in outer and inner layers of organoids (Fig. 3a, b). In addition to ECM proteins, this procedure enables the detection of cytoskeletal, membrane, and nuclear proteins in situ (Fig. 3a–d). Furthermore, this protocol describes the steps to isolate proteins and mRNA from 3D BME and Collagen I gels for biochemical and molecular analysis such as western blot and real-time quantitative PCR (RT-qPCR) (Fig. 4a–c). Although we use murine breast cancer organoids, this protocol is applicable for models using organoids derived from different tumor types (including colon cancer, pancreatic ductal carcinoma, and head and neck squamous cell carcinoma) as well as healthy tissues (including breast, colon, and pancreas). Combined with pharmacological and molecular interferences (e.g., lentiviral transductions and Crisp/Cas 9 gene deletions) and/or live-cell imaging, this collection of protocols will be useful for research groups working on drug screening or aiming to study the cellular and molecular mechanisms underlying a wide range of important processes during morphogenesis, differentiation, and carcinogenesis.
Passaging of murine breast cancer organoids in 3D BME. (a) Workflow for the dissociation of MMTV-PyMT and MMTV-NeuT organoids into single cells using trypsin-EDTA and propagation of the 3D cultures. (b) Representative bright-field images using 10× objective (NA = 0.4, EVOS microscope) showing the different stages of MMTV-PyMT organoid cultures: prior to and post dissociation into individual cells and after organoid formation from single cells. Insets, established multicellular organoids or individual cells. Scale bar, 100 μm
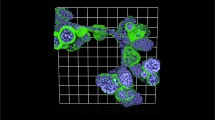
Variations in isolating organoids from cultures and embedding in attached or floating 3D ECM. (a) Workflow for isolating established organoids from cultures with or without ECM digestion (Dispase II, 1 mg/mL) followed by three washing steps. Isolated organoids are embedded in Collagen I or BME. To manipulate ECM biomechanics, gels may be kept attached to or detached from the bottom of the well. Whereas the ECM concentration is the same, floating gels have lower mechanical stiffness and tension buildup compared to the attached gels. (b, c) Images represent single confocal slices from the mid-region of MMTV-PyMT organoids embedded in 2 mg/mL Collagen I for 1 day. The organoids were isolated from BME cultures (b) without and (c) with Dispase II treatment. Detection of the ECM protein laminin-α1, cytoskeletal markers that identify different cancer cell subtypes keratin 14 (K14, basal cell marker) and keratin 8 (K8, luminal cell marker). Collagen I was detected by reflection mode using the 488 nm laser. White arrowheads: continuous layer of laminin-α1 around the organoid. White asterisks: very weak or absent laminin-α1 layer. (d) Bright-field images of MMTV-PyMT organoids embedded in Collagen I or BME (attached or floating) for 3 days. Black arrowheads: Multicellular invasive strands (collective invasion). (e) Percentage of organoids that show one or more invading strands. P value, two-tailed student t-test. (e) Bright-field images of MMTV-NeuT organoids growing in Collagen I or BME for 3 days. Unlike MMTV-PyMT organoids, MMTV-NeuT organoids do not show protrusive collective invasive strands in 2 mg/mL Collagen I. Scale bars 100 μm (d, f), 50 μm (b, c), 10 μm (b, c inset)
Different fixation methods and confocal imaging of cytoskeletal, membrane, and nuclear proteins. (a, b) Confocal imaging of keratin 14 (K14) and keratin 8 (K8) of MMTV-PyMT organoids embedded in (a) BME for 3 days and fixed using PFA+glutaraldehyde and (b) Collagen I or BME for 3 days and fixed using PFA only. White arrowheads: K14-positive cells located at the outer layers of the organoids and guiding collective invasion (in Collagen I condition). White arrows: Collagen I fibers aligned parallel to the direction of migration of the invasive multicellular strands. White asterisks: K8-positive cells at inner regions of the organoid. Detection of (c) active β1 integrins enriched in protrusions of cells guiding collective invasion (white arrowheads) and (d) nuclear proteins: transcription factor p63. Scale bars 50 μm (a–c), 10 μm (c, d inset)
Analysis of protein and RNA expression from organoids growing and invading in different 3D ECM. (a) Western blot showing the expression of keratin 14 (K14), keratin 17 (K17), and β-actin from whole cell lysates isolated from wild-type (WT) and K17 knockout (KO) MMTV-PyMT organoids embedded in 3D BME and Collagen I for 3 days. Organoids from both BME and Collagen I (COLI) gels were isolated using Dispase II and Collagenase I. (b) Western blot displaying the effect of Dispase II and Collagenase I enzymes on the detection of proteins located at ~60–70 kDa height, for example, pYap (Ser127). Without enzymatic digestion, a smear appears on the location of 60–70 kDa that prevents the detection of clear distinct bands. In contrast, β-actin (45 kDa) is clearly visible after both isolation methods. (c) RT-qPCR analysis of mRNA from MMTV-PyMT organoids embedded in BME and Collagen I and isolated using TRIzol. Data shows the fold change (FC) between BME and Collagen I for various genes and calculated using the 2−ΔΔCt method. Data was normalized over a set of housekeeping genes (average) Tuba1, Cyca, Pbgd, and Hnrnpa. P values, two-tailed student t-test
2 Materials
2.1 Propagation of Organoids
2.1.1 Thawing and Passaging
-
1.
Frozen breast cancer organoids (MMTV-PyMT and MMTV-NeuT organoids) [27] (kind gift from Dr. Jacco van Rheenen).
-
2.
Growth factor-reduced basement membrane extract (BME), type 2, PathClear (Cat# 3533-005-02, Cultrex) (see Note 1).
-
3.
Working medium: Advanced DMEM/F-12, 1% v/v penicillin-streptomycin (P/S), 10 mM HEPES buffer solution, 1× GlutaMAX.
-
4.
Growth medium: Working medium supplemented with 1× B27 (Cat# 17504001, Life Technologies), 1.25 mM N-acetyl cysteine (Cat# A9165-5G, Sigma-Aldrich), 1× Primocyn (Cat# ant-pm-2, Bio-connect), 2.5 nM FGF2.
-
5.
Trypsin-EDTA.
-
6.
Bright-field microscope.
-
7.
Ice.
-
8.
Water bath.
-
9.
15 mL conical tubes.
-
10.
50 mL conical tubes.
-
11.
Centrifuge.
-
12.
37 °C prewarmed 24-well plate.
-
13.
EVOS M5000 cell imaging system (ThermoFischer Scientific) and Fiji software (ImageJ, US National Institutes of Health, Bethesda, Maryland, USA).
2.1.2 Freezing
-
1.
Recovery™ Cell Culture Freezing Medium (Cat# 12648-010, Gibco).
-
2.
2 mL cryovials.
-
3.
−80 °C fridge.
-
4.
Liquid nitrogen.
2.2 Transfer of Established Organoids
2.2.1 Isolating Organoids from BME Gels with Dispase
-
1.
Dispase II (Cat# 17105041, Life Technologies).
-
2.
Working medium.
-
3.
15 mL conical tubes.
-
4.
Water bath.
2.2.2 Isolating Organoids from BME Gels Without Dispase
-
1.
Working medium.
-
2.
15 mL conical tubes.
-
3.
Water bath.
2.2.3 Embedding Organoids (from BME to Collagen I)
-
1.
Collagen I, rat tail (3.28 mg/mL, Cat# 354236, Corning®).
-
2.
1N NaOH.
-
3.
10× PBS (Mg2+- and Ca2+-free): 26.7 mM KCl, 14.7mM KH2PO4, 1.38 M NaCl, 80.6 mM Na2HPO4-7H2O.
-
4.
Milli-Q water.
-
5.
pH paper.
-
6.
15 mL conical tubes.
-
7.
1.5 mL Eppendorf tube.
-
8.
Water bath.
-
9.
Prewarmed 24-well plate.
-
10.
Growth medium.
2.2.4 Embedding of Organoids in BME (BME to BME)
-
1.
15 mL conical tubes.
-
2.
1.5 mL Eppendorf tube.
-
3.
Water bath.
-
4.
Prewarmed 24-well plate.
-
5.
Growth medium.
2.3 Fixation
2.3.1 Fixation with PFA and Glutaraldehyde
-
1.
8% paraformaldehyde (PFA) in Milli-Q.
-
2.
25% glutaraldehyde.
-
3.
1× PBS: (Mg2+- and Ca2+-free) 2.67 mM KCl, 1.47 mM KH2PO4, 138 mM NaCl, 8.06 mM Na2HPO4-7H2O.
-
4.
Sodium borohydride buffer (0.01 g sodium borohydride in 10 mL 1× PBS).
2.3.2 Fixation with PFA
-
1.
8% PFA in Milli-Q water.
-
2.
10× PBS.
2.4 Immunostaining for Confocal Microscopy
-
1.
Blocking buffer: 1× PBS, 10% (v/v) normal goat serum 0.3% (v/v) Triton X-100 (100%).
-
2.
Antibody dilution buffer: 1× PBS, 1% (w/v) bovine serum albumin, 0.3% (v/v) Triton X-100 (100%).
-
3.
Antibodies and dilutions: Rat anti-mouse keratin 8 (Cat# 531826, DSHB) 1:50, rabbit anti-human keratin 14 (Cat# 905301, BioLegend) 1:300, rat anti-mouse laminin alpha 1 (Cat# MAB4656, R & D Systems) 1:100, rat anti-mouse integrin beta-1 (9EG7 Cat# 553715, BD Pharmingen) 1:50, mouse anti-human p63 (Cat# GA66261-2) 1:100.
-
4.
Secondary Alexa fluorophore-conjugated antibodies.
-
5.
Dapi, phalloidin.
-
6.
Zeiss LSM 880 (40× objective, NA = 1.1).
2.5 Protein and RNA Extraction
2.5.1 Protein Extraction
-
1.
Collagenase I (Cat# C0130, Sigma-Aldrich).
-
2.
Dispase II (Cat# 17105041, Life Technologies).
-
3.
Laemmli sample buffer 5×: 10% (w/v) SDS, 50% (v/v) glycerol, 9% (v/v) β-mercaptoethanol, 0.3 M Tris–HCL pH 6.8, 0.5% (w/v) bromophenol blue in water.
-
4.
Sonicator.
-
5.
Heat block for Eppendorf tubes, preheated at 100 °C.
2.5.2 Western Blot
-
1.
40% acrylamide/Bis solution.
-
2.
Electrophoresis running buffer: 25 mM tris, 190 mM glycine, 3.5 mM SDS.
-
3.
Blotting buffer: 25 mM tris, 190 mM glycine with 10% v/v methanol.
-
4.
100% methanol.
-
5.
PVDF membranes.
-
6.
1× Tris buffer saline (TBS): 50 mM tris, 150 mM NaCl, pH 7.4.
-
7.
Blocking buffer: 5% w/v non-fat dry milk in 1× TBS.
-
8.
TBS-T: 1× TBS with 0.1% v/v Tween-20.
-
9.
Antibody dilution buffer: TBS-T with 5% w/v bovine serum albumin.
-
10.
Primary antibodies and dilutions: Rat anti-mouse keratin 8 (Cat# 531826, DSHB) 1:500, rabbit anti-human keratin 14 (Cat# 905301, BioLegend) 1:1000, rabbit anti-human keratin 17 (Cat# ab183330, Abcam) 1:1000, rabbit anti-human Phospho-YAP1 (S127) (Cat# 4911S, CST) 1:500, mouse anti-chicken actin (Cat# 1501, Millipore) 1:3000.
-
11.
Secondary fluorescent-labeled antibodies and dilutions: IRDye secondary antibodies (Li-COR) 1:2500.
-
12.
Fluorescence detector (Amersham Typhoon, GE Life Sciences).
2.5.3 RNA Extraction
-
1.
Ice-cooled TRIzol reagent.
-
2.
Ice-cooled chloroform.
-
3.
70% ethanol in Milli-Q water.
-
4.
Fume hood.
-
5.
RNaseZap™RNase Decontamination Solution (Cat# AM9780, Invitrogen™).
-
6.
RNeasy Lipid Tissue Mini Kit (Cat# 74804, Qiagen).
-
7.
Nanodrop 2000 spectrophotometer.
-
8.
Tabletop centrifuge for Eppendorf tubes, cooled to 4 °C.
2.5.4 cDNA Synthesis
-
1.
iScript™ cDNA Synthesis Kit (Cat# 1708890, Bio-Rad).
-
2.
Thermal cycler for optimized for PCR.
2.5.5 RT-qPCR
-
1.
cDNA diluted in Milli-Q water.
-
2.
Thermal cycler optimized for real-time quantitative PCR (RT-qPCR) (CFX96 Touch Real-Time PCR Detection System (Bio-Rad)).
-
3.
2× FastStart Universal SYBR Green Master mix (Rox) (Merck).
-
4.
Forward and reverse primers that detect the gene products of interest. Ensure that primers have a product length between 50 and 150 bp. Primer can be designed using the NCBI Primer-Blast tool, accessible at https://www.ncbi.nlm.nih.gov/tools/primer-blast/) (Table 1).
3 Methods
3.1 Propagation of Cancer-Derived Organoid Culture
This protocol describes the culture of breast cancer organoids (originally isolated from mice harboring MMTV-PyMT or MMTV-NeuT tumors) starting from frozen vials. This protocol does not describe the isolation of organoids from the primary tumor, and this has been thoroughly described elsewhere [14, 15, 28].
-
1.
Prewarm the tissue culture plates for at least 48 h in the incubator (37 °C, 5% CO2, humidified atmosphere). This ensures that the reconstituted ECM forms a drop of the appropriate height from the bottom of the plate.
-
2.
Prewarm 10 mL working medium in a 15 mL tube.
-
3.
Thaw BME on ice (see Note 1).
-
4.
Thaw frozen cryovial-containing organoids in a 37 °C water bath.
-
5.
Collect organoids and place in 10 mL medium in a 15 mL tube.
-
6.
Centrifuge for 4 min at 4 °C, 253 g.
-
7.
Inspect the pellet, which contains the organoids. In some cases a diffuse layer of BME (from previous culture) is visible above the organoid pellet.
-
8.
Remove the supernatant. Do not discard the diffuse BME layer since it might contain many organoids.
-
9.
Suspend the pellet in 100–150 μL BME. The volume of BME may be adapted depending on the number of organoids present in the pellet.
-
10.
Pipette the organoids/BME mixture as 25–45 μL drops on the prewarmed 24-well plate (see Note 2).
-
11.
Flip the plate carefully to form a hanging drop. This will reduce the number of organoids that will grow close to the basal surface of the culture plates.
-
12.
Incubate in 37 °C, 5% CO2, humidified atmosphere for 30 min until the gel is polymerized.
-
13.
Add 750 μL of prewarmed growth medium to the gels. Attention: adding cold medium will dissolve the BME gel.
-
14.
Place the plate in 37 °C, 5% CO2, humidified atmosphere.
-
15.
Change growth medium every 2nd day.
3.1.1 Passaging (Fig. 1a)
Passaging organoids allows the maintenance and expansion of cultures. Expansion of cultures is important to generate sufficient amount of cells needed for analysis such as western blot, RT-qPCR, and fluorescent-activated cell sorting (FACS).
-
1.
Inspect BME gels containing organoids and assess confluency by bright-field microscopy (see Note 3) (Fig. 1a, b).
-
2.
Gels containing organoids with more than ~100 cells and/or occupying more than 50% of the gels are taken for passaging.
-
3.
Remove growth medium from the well.
-
4.
Add 300 μL of cold trypsin/EDTA per gel (the low temperature of this solution will dissolve the BME gel).
-
5.
Suspend by successive pipetting up and down using a p200 pipette.
-
6.
Transfer the organoids into a 15 mL tube and mix well.
-
7.
Incubate the tube in a 37 °C water bath.
-
8.
Inspect regularly (every 15–20 s) whether the organoids are dissociated into small clusters and single cells by checking the bottom of the 15 mL tube with a bright-field microscope.
-
9.
After every inspection, mix up and down with a p200 pipette. This will speed up the disintegration of the organoids into small clusters and/or single cells.
-
10.
Depending on the desired extent of organoid disintegration, proceed to step 11. Here, we fully dissociated the organoids into single cells (Fig. 1a, b).
-
11.
Pipette up and down a few times with a p200 pipette.
-
12.
Spin down for 4–5 min at 253 g at 4 °C.
-
13.
Inspect the pellet and discard the supernatant.
-
14.
Add 50–300 μL of cold BME per gel, depending on the confluency of the organoids and the desired number of cells. Mix by pipetting up and down (ten times). Put on ice while you get the prewarmed 24-well plate.
-
15.
Pipette 40–50 μL drops on the prewarmed plastic well and flip the plate upside down. You should now have hanging drops. Place in the incubator (37 °C, 5% CO2, humidified atmosphere) for 30 min, until the BME drops polymerized and formed droplets.
-
16.
Add 750–1000 μL of the prewarmed growth medium.
-
17.
Microscopical imaging and analysis (see Note 4).
3.1.2 Freezing
-
1.
Suspend BME gels and organoids in 1 mL cold working medium.
-
2.
Add to 10 mL cold working medium in a 15 mL tube. To avoid the formation of a large BME layer above the pellet (after centrifugation, step 3), do not pool many gels in one tube (maximum 150 μL of BME gels per 15 mL tube).
-
3.
Spin down 253 g, 4 °C for 5 min.
-
4.
Discard the supernatant.
-
5.
Add slowly 250–1000 μL of recovery freezing medium depending on confluency of the organoids and desired dilution of organoids in the cryovials. Typically, the contents of one confluent gel (40 μL) can be frozen in 4 cryovials.
-
6.
Transfer on ice to −80 °C.
-
7.
After at least 48 h, transfer the cryovial to liquid nitrogen.
3.2 Isolation and Embedding of Organoids in Collagen I or BME
3.2.1 Isolating Organoids from Cultures Using Cold Medium (Fig. 2a, b)
Organoids grown in BME cultures can be isolated from the gels using only cold working medium. Using this method, basement membrane components remain present and form a layer around the isolated organoid (Fig. 2, arrowheads). This protocol is useful when a layer of basement membrane components is desired to be present around the organoids that will be embedded in Collagen I or other matrices. Embedding organoids that are surrounded by a layer of basement membrane components in Collagen I may be used to study cancer cell penetration of the basement membrane and early contacts with and subsequent invasion into the Collagen I-rich stroma.
-
1.
Remove medium surrounding the BME droplet.
-
2.
Add 500 μL cold working medium.
-
3.
Suspend the organoids in the BME by pipetting up and down.
-
4.
Transfer the suspended organoids to a 15 mL tube.
-
5.
Fill up to 10 mL with cold working medium.
-
6.
Spin down for 4 min 253 g.
-
7.
Discard supernatant.
-
8.
Suspend the organoids with the required volume of cold working medium.
-
9.
Now the organoids are ready to be embedded in Collagen or other hydrogels.
3.2.2 Isolating Organoids from Cultures Using Dispase II (See Note 5) (Fig. 2a, c)
Treatment with Dispase II and the following washes remove a large portion of the basement membrane components present around the organoids. This protocol is useful when no or little basement membrane components are desired to be present around organoids when they are embedded in Collagen I or other matrices (Fig. 2c). Embedding such organoids in Collagen I models tumors that have already lost basement membrane coverage and allows studying cell proliferation/survival and invasion when the cancer cells are already present within the Collagen I-rich tissue.
-
1.
Remove medium surrounding the well.
-
2.
Add 500 μL cold working medium, and gently suspend the organoids in the BME.
-
3.
Transfer the suspended organoids to a 15 mL tube.
-
4.
Fill up to 10 mL with cold working medium.
-
5.
Spin down for 4 min at 253 g.
-
6.
Discard supernatant.
-
7.
Add 750 μL of 1 mg/mL Dispase II diluted in working medium.
-
8.
Incubate the gel + Dispase II at 37 °C in a water bath for 15 min.
-
9.
Add 10 mL of working medium to the 15 mL tube.
-
10.
Spin down 253 g for 1 min.
-
11.
Discard supernatant.
-
12.
Wash three times by adding 10 mL of cold working medium to the 15 mL tube, spin down, and remove supernatant.
-
13.
Now the organoids are ready to be embedded in Collagen I or other hydrogels (see Note 6).
3.2.3 Preparation of 3D Collagen I Gels
-
1.
In a 1.5 mL Eppendorf tube (final concentration 2 mg/mL), add 45 μL of 10× PBS, 32.6 μL H2O, 6.6 μL 1 N NaOH (see Note 7), and 365.8 μL Collagen I (see Notes 7 and 8).
-
2.
Incubate on ice for 2 h.
3.2.4 Embedding Organoids in 3D Collagen I (Fig. 2a)
For a total gel volume of 600 μL:
-
1.
Add 150 μL of cells (organoids or single cells, see Note 6) in working medium to 450 μL of Collagen mixture.
-
2.
Mix thoroughly with a p1000 pipette.
-
3.
Incubate at room temperature for 5 min.
-
4.
After 5 min, pipette up and down one to two times (Collagen is already polymerizing so do not mix too long at this stage).
-
5.
Plate as 45 μL drops in prewarmed 24-well plate (see Note 9).
-
6.
Keep the gel for 1 min at room temperature.
-
7.
To reduce the growth of organoids close to the basal surface of the plate, flip the plate carefully (so that it is upside-down) and put it at 37 °C for 1 min.
-
8.
Repeat step 7 twice.
-
9.
Incubate at 37 °C for at least 15 min.
-
10.
After polymerization add pre-warmed growth medium or fix (if timepoint 0 in Collagen is required).
-
11.
Detach the Collagen gel from the plate in case Collagen I gel with less stiffness and tension buildup is required [29, 30] (Fig. 2a, d) (see Note 8).
3.2.5 Embedding of Organoids Back to BME (See Note 10)
-
1.
Suspend isolated organoids from (Subheadings 3.2.1 or 3.2.2) in working medium.
-
2.
Add suspended organoids to BME.
-
3.
Mix thoroughly.
-
4.
Per well, plate a 45 μL drop in prewarmed 24-well plate.
-
5.
Keep the gel for 1 min at room temperature.
-
6.
To reduce the growth of organoids close to the basal surface of the plate, flip the plate carefully (so that it is upside-down) and put it at 37 °C for 1 min.
-
7.
Repeat step 8 twice.
-
8.
Incubate at 37 °C for at least 15 min.
-
9.
After polymerization add prewarmed growth medium or fix (if timepoint 0 in Collagen I is required).
-
10.
Microscopical imaging and analysis. We used the EVOS cell imaging system and Fiji software, respectively (see Note 4) (Fig. 2d–f).
3.3 Fixation and Immunostaining for Confocal Microscopy
3.3.1 Fixation with PFA and Glutaraldehyde (Fig. 3a)
Fixing organoids in BME gels with PFA causes the gel to dissolve partially or completely. If the ECM is desired to be maintained, the dissolution of the gel is avoided by the addition of glutaraldehyde (see Note 11). This can be used with both BME and Collagen I gels.
-
1.
Add up to 500 μL of 4% PFA in 1× PBS containing 0.25% (v/v) glutaraldehyde to the BME or Collagen I gels.
-
2.
Incubate for 10–15 min at room temperature.
-
3.
Wash 2× with 1× PBS.
-
4.
Wash 3× with sodium borohydride (see Note 11).
-
5.
Wash 2× with 1× PBS.
3.3.2 Fixation with PFA (See Note 12) (Fig. 3b)
-
1.
Add up to 500 μL of 4% PFA in 1× PBS.
-
2.
Incubate for 10–15 min at room temperature.
-
3.
Wash 2× with 1× PBS.
Fixed gels may be stored in PBS at 4 °C for several weeks when properly sealed (paraffin). To prevent microbial contamination and for longer storage periods, add sodium azide at a final concentration of 0.3% (in 1× PBS).
3.4 (Immuno)staining for Confocal Microscopy (See Notes 13 and 14)
-
1.
Add 1 mL of PBS to a 1.5 mL Eppendorf tube.
-
2.
Transfer fixed BME or Collagen I gels to a 1.5 mL Eppendorf tube using a spatula.
-
3.
Remove PBS.
-
4.
Transfer gels to blocking buffer and let them shake for 60–90 min.
-
5.
Aspirate blocking solution, apply diluted primary antibody.
-
6.
Incubate for 20 h at 4 °C.
-
7.
Wash at least four times with 1× PBS for at least 10–15 min each with shaking.
-
8.
Incubate specimen in secondary antibody (1:500), Dapi (5 μg/mL), and phalloidin (1:100) for 20 h in the dark at 4 °C.
-
9.
Wash at least four times with 1× PBS for at least 10–15 min each.
-
10.
This procedure allows the visualization of cytoskeletal (Fig. 3a, b, d), membrane (Fig. 3c) and nuclear (Fig. 3d) proteins during growth and invasion in BME and Collagen I gels (Fig. 3b, c, d).
-
11.
Label-free confocal imaging of Collagen I is performed using 488 nm laser, the main dichroic beam splitter (MBS) T80/R20, and detection range of 469–496 nm.
3.5 Extraction of Organoids for Protein and RNA Isolation
This protocol describes the isolation of proteins and RNA from whole cell lysates, ready to be used for biochemical and molecular analysis such as western blotting (Fig. 4a, b) and RT-qPCR (Fig. 4c), respectively.
3.5.1 Using Dispase II and Collagenase I
-
1.
Add the appropriate amount of Collagenase I (20 mg/mL) and or Dispase II (1 mg/mL) in PBS in a 15 mL tube. At least 100 μL of enzyme (Collagenase and or Dispase) per gel (40–50 μL) should be used.
-
2.
Pick up Collagen I or BME drop with a spatula and place in the 15 mL tube
-
3.
Incubate in 37 °C water bath with manual shaking for 3 min.
-
4.
Fill the 15 mL tube to 10 mL with 1× PBS.
-
5.
Centrifuge for 5 min at 253 g at 4 °C.
-
6.
Aspirate supernatant (see Note 15).
-
7.
Add to the pellet 1–2.5× Laemmli sample buffer; 40–50 μL per 1 confluent gel (see Note 16).
-
8.
Lyse cells by pipetting up and down.
-
9.
Incubate at least 15 min at room temperature.
-
10.
Sonicate for at least 5 cycles at 4 °C.
-
11.
Boil samples for 5 min using a heat block at 95 °C.
-
12.
Samples are now ready to be analyzed by western blot (see Note 17 and Fig. 4a, b). Continue at Subheading 3.5.3 for the protocol used here.
3.5.2 Without Enzymes (See Note 18) (Fig. 4b)
-
1.
BME and Collagen I gels may be placed immediately in the lysis buffer.
-
2.
Perform 15 cycles of sonication.
-
3.
Boil the samples for 5 min using a heat block at 95 °C.
-
4.
Samples are ready for western blot.
3.5.3 Western Blot
-
1.
Load 20 μL of lysate onto a 10% SDS polyacrylamide gel.
-
2.
Separate the proteins by gel electrophoresis at 90 V for ~1 h.
-
3.
Soak the PVDF membranes in methanol for 10 s and then in water for 5 min.
-
4.
Electrotransfer (wet transfer) the proteins to activated PVDF membranes at 0.35 A in blotting buffer for 90 min at 4 °C.
-
5.
Block membranes for 1 h at room temperature.
-
6.
Incubate the membranes with primary antibody overnight at 4 °C.
-
7.
Wash 4× with TBS-T, 10 min each.
-
8.
Incubate membrane with secondary antibodies for 1 h at room temperature.
-
9.
Wash 4× with TBS-T, 10 min each.
-
10.
Scan the membranes using a fluorescence detector.
3.5.4 RNA Isolation
-
1.
Clean the working space with RNaseZap™ Decontamination solution before performing RNA isolation.
-
2.
Remove medium surrounding the 3D cultures (Collagen I or BME gels).
-
3.
Add 500 μL TRIzol reagent per gel.
-
4.
Homogenize gels in TRIzol by extensive pipetting up and down using a cutoff tip (cut with scissors that are decontaminated by RNaseZap™ Decontamination Solution).
-
5.
Additional extensive pipetting up and down to allow cell lysis, until the sample (gel and cells) is homogeneous.
-
6.
Vortex.
-
7.
Incubate for 5 min at room temperature.
-
8.
Add 100 μL chloroform per 500 μL of TRIzol that was added at step 2. Shake the tube vigorously. Incubate for 2–3 min at room temperature.
-
9.
Centrifuge the samples for 15 min at max speed (15,000 g) at 4 °C. The mixture now separates into a lower red phenol-chloroform phase, a white-colored intermediate phase, and a colorless upper aqueous phase.
-
10.
Transfer the aqueous phase containing the RNA to a new tube. Hold the tube at an angle to ensure that all of the aqueous phase is transferred. Avoid pipetting any of the intermediate and organic phase. Important: it is better to sacrifice aqueous material than risk pipetting the layers below, as they will contaminate the samples.
-
11.
Add an equal volume of 70% ethanol to the aqueous phase. Vortex.
-
12.
Proceed with the ‘on column gDNA digestion’ step of the QIAgen RNeasy kit by transferring up to 700 μL of the RNA mixture to a filter with spin column and follow the steps of the kit. In these steps, RNA will be washed using 70% ethanol and eluted in 30–50 μL Milli-Q.
-
13.
Measure nucleotide concentration and the 260/280 ratio using a spectrophotometer. The 260/280 ratio is a measure of nucleic acid purity, which for RNA should be between 1.9 and 2.1 (see Note 19).
-
14.
Snap-freeze all RNA that is not used for subsequent cDNA synthesis in liquid nitrogen. Store all RNA samples at −80 °C. mRNA is now ready to be used for subsequent analysis (Fig. 4c). Here it is used for cDNA synthesis and RT-qPCR.
3.5.5 cDNA Synthesis
-
1.
Synthesize cDNA following the iScript cDNA synthesis protocol with 1000 ng RNA as input.
3.5.6 RT-qPCR
-
1.
For the RT-qPCR reaction, mix (to a final volume of 15 μL): 10–100× diluted cDNA, 400 nM of each forward and reverse primer, and 1× FastStart Universal SYBR Green Master mix.
-
2.
Use the following program on the thermal cycler: 95 °C for 10 min, then a 40× repeated cycle of 95 °C for 10 s (denaturation), 55 °C for 10 s (annealing), 72 °C for 30 s (extension). A melt peak program of 95 °C for 10 s and increments of 0.5 °C from 65 °C–95 °C was used to confirm whether the primers amplify multiple products.
4 Notes
-
1.
In all the steps described in this chapter, Matrigel may be used as an alternative to BME. Both Matrigel and BME originate from murine tumor cells and are used in in vitro models to mimic basement membranes [17, 31]. The major ECM components present in Matrigel and BME are laminin 1, Collagen IV, and heparan sulfate [32]. The ECM mixture is liquid at low temperatures (4 °C) and polymerizes into a gel at higher temperatures (including 25 °C and 37 °C).
-
2.
When a bigger volume of BME is used (50–100 μL) organoids in the middle of the gels tend to be stressed and grow much slower than the organoids located at the borders. The use of smaller volumes (20–40 μL) secures that the organoids in the middle and border regions of the BME gels receive comparable amounts of nutrients and oxygen. With small volumes, multiple gels can be pipetted in one well of a 6- or 12-well plate.
-
3.
Confluence of cultures in 3D is less straightforward to assess compared to 2D cultures. In 3D, confluency is determined based on 2 criteria: size of the organoids and free space in the BME gel. When the organoids are big (more than 100 cells, with dark core) the use of trypsin-EDTA to dissociate the cells is required prior to splitting of the cultures. If the gels contain many but small organoids then dissolve the gels with cold medium and split the cells without trypsin-EDTA.
-
4.
Organoid growth may be analyzed by measuring the area or volume of the organoids over time. Invasion can be measured by calculating the percentage of invasive organoids (Fig. 2e) or by quantifying the number and length of invasive strands [25] or the number of invasive single cells per organoid [33]. In addition, morphological analysis of invasion can be performed. Depending on the invasion mode (collective or single cell) the morphology of invasion can be determined. For instance, collective invasive strands may be classified as protrusive (pointed morphology) or blunt (round morphology); Invasive single cells may be classified as rounded or spindle-shaped.
-
5.
The enzyme Dispase II degrades the basement membrane efficiently by cleaving Collagen IV and not laminin 1 [34].
-
6.
If individual cells are needed, organoids should be dissociated into single cells using trypsin-EDTA. This is followed by treatment with trypsin inhibitors and embedding into Collagen I.
-
7.
Rat tail Collagen I constitutes monomeric Collagen I dissolved in acetic acid. The polymerization into Collagen I fibers depends on pH and temperature. Neutralization is performed by the addition of a base such as NaOH. To ensure a Collagen I gel with pH 7–7.5, the amount of 1N NaOH to be added should be adjusted with every new Collagen I lot (due to lot variations in Collagen I concentration and acid content). According to manufacturer’s instructions (Corning), the volume of NaOH is calculated based on a factor (0.03) multiplied by the volume of Collagen I added. However, the factor needs to be adjusted with every new lot, depending on the volume of NaOH required to get a Collagen I mix with a pH of 7–7.5.
-
8.
The mechanics of Collagen I can also be modulated by changing the final Collagen I concentration, time/temperature used for polymerization, or inducing non-enzymatic cross-linking through glycation prior to gel polymerization [35, 36].
-
9.
Use microscopy-compatible plates instead of standard tissue culture plates for time-lapse imaging.
-
10.
Organoids that form from single cells can be isolated from the BME gels and embedded in BME again. This may be important for studying how established organoids behave in BME or may be used as control conditions for organoids growing in other matrices such as Collagen I.
-
11.
Glutaraldehyde fixation keeps several free aldehyde groups in the cross-linked proteins within the fixed samples. The free aldehyde groups bind to many proteins including the antibodies, which increases the nonspecific background signal (autofluorescence). Washing the samples with freshly prepared sodium borohydride buffer renders the free aldehyde groups nonreactive. Consequently, nonspecific binding to antibodies and background signal are reduced.
-
12.
In case it is not required to maintain the architecture of the BME, PFA fixation may be used. PFA-fixed organoids can be isolated from the disrupted/dissolved BME (due to PFA) by centrifugation and embedded in other gels such as Collagen I. This is followed by fixation of the new Collagen gel and subsequent immunostaining steps. Such a procedure may be important if antibody penetration is different in BME versus Collagen I and whether expression levels are to be compared between the two conditions. Alternatively, warm PFA (37 °C) may be used to fix BME gels in microscopy-compatible multiwell chambers. To reduce the dissociation of the BME gels, immunostaining steps are performed at room temperature in the presence of sodium azide. Warm temperatures during PFA fixation and immunostaining reduce the dissolution of the fixed BME gel (enhanced in low temperatures).
-
13.
All the steps of immunostainings (blocking, antibody incubation, and washing) of organoids in 3D gels should be performed with shaking. Shaking enhances the penetration of the blocking peptides, antibodies into the gels, and over the multicellular layers of the organoids and the washing of nonspecific antibody interactions.
-
14.
Depending on the protein of interest to be visualized, Triton X-100 can be replaced with other detergents such as saponin or Tween-20.
-
15.
Organoids that are isolated from the BME and Collagen I gels can alternatively be dissociated into single cells using trypsin-EDTA followed by single-cell-based analyses such as FACS and/or single-cell sequencing.
-
16.
Isolated organoids can also be lysed in Ripa buffer and analyzed for protein concentration prior to loading into acrylamide gels.
-
17.
Three to six confluent gels (volume of the gel = 40 μL) of BME or Collagen I provide whole cell lysates sufficient for at least three rounds of western blots (30 μL loading/per round).
-
18.
The extraction of whole cell lysates without enzymes is a fast method to extract proteins and may be essential whenever fast regulation of proteins is to be assessed. However, this procedure has limitations since there will be a large amount of ECM proteins in the lysate. When analyzed by western blot, thick bands (blobs) without a distinct border are detected at the ~60 kDa height. The identity of these proteins remains unknown (Fig. 4b), but they are probably partially digested (due to Laemmli buffer and sonication) ECM proteins.
-
19.
RNA isolation is performed without enzymatic digestion, to ensure a fast procedure and that RNA is kept as intact as possible. Constantly keep samples on ice during RNA isolation. With the RNA extraction method, 6 mid-confluent BME and Collagen I gels yield 10–50 μg of RNA.
References
Schwartz MA (2010) Integrins and extracellular matrix in mechanotransduction. Cold Spring Harb Perspect Biol 2(12):a005066–a005066. https://doi.org/10.1101/cshperspect.a005066
Theocharis AD, Skandalis SS, Gialeli C, Karamanos NK (2016) Extracellular matrix structure. Adv Drug Deliv Rev 97:4-27. doi:https://doi.org/https://doi.org/10.1016/j.addr.2015.11.001
Sekiguchi R, Yamada KM (2018) Basement membranes in development and disease. Curr Top Dev Biol 130:143–191. https://doi.org/10.1016/bs.ctdb.2018.02.005
Mouw JK, Ou G, Weaver VM (2014) Extracellular matrix assembly: a multiscale deconstruction. Nat Rev Mol Cell Biol 15(12):771–785. https://doi.org/10.1038/nrm3902
Chang J, Chaudhuri O (2019) Beyond proteases: basement membrane mechanics and cancer invasion. J Cell Biol 218(8):2456–2469. https://doi.org/10.1083/jcb.201903066
Mylonas CC, Lazaris AC (2014) Colorectal cancer and basement membranes: clinicopathological correlations. Gastroenterol Res Pract 2014:580159. https://doi.org/10.1155/2014/580159
Vellinga TT, den Uil S, Rinkes IH, Marvin D, Ponsioen B, Alvarez-Varela A, Fatrai S, Scheele C, Zwijnenburg DA, Snippert H, Vermeulen L, Medema JP, Stockmann HB, Koster J, Fijneman RJ, de Rooij J, Kranenburg O (2016) Collagen-rich stroma in aggressive colon tumors induces mesenchymal gene expression and tumor cell invasion. Oncogene 35(40):5263–5271. https://doi.org/10.1038/onc.2016.60
Nguyen-Ngoc K-V, Cheung KJ, Brenot A, Shamir ER, Gray RS, Hines WC, Yaswen P, Werb Z, Ewald AJ (2012) ECM microenvironment regulates collective migration and local dissemination in normal and malignant mammary epithelium. Proc Natl Acad Sci 109(39):E2595–E2604. https://doi.org/10.1073/pnas.1212834109
Lai SL, Tan ML, Hollows RJ, Robinson M, Ibrahim M, Margielewska S, Parkinson EK, Ramanathan A, Zain RB, Mehanna H, Spruce RJ, Wei W, Chung I, Murray PG, Yap LF, Paterson IC (2019) Collagen induces a more proliferative, migratory and chemoresistant phenotype in head and neck cancer via DDR1. Cancer 11(11):1766. https://doi.org/10.3390/cancers11111766
Conklin MW, Eickhoff JC, Riching KM, Pehlke CA, Eliceiri KW, Provenzano PP, Friedl A, Keely PJ (2011) Aligned collagen is a prognostic signature for survival in human breast carcinoma. Am J Pathol 178(3):1221–1232. https://doi.org/10.1016/j.ajpath.2010.11.076
Provenzano PP, Inman DR, Eliceiri KW, Knittel JG, Yan L, Rueden CT, White JG, Keely PJ (2008) Collagen density promotes mammary tumor initiation and progression. BMC Med 6:11–11. https://doi.org/10.1186/1741-7015-6-11
Le CC, Bennasroune A, Langlois B, Salesse S, Boulagnon-Rombi C, Morjani H, Dedieu S, Appert-Collin A (2020) Functional interplay between collagen network and cell behavior within tumor microenvironment in colorectal cancer. Front Oncol 10:527–527. https://doi.org/10.3389/fonc.2020.00527
Drifka CR, Loeffler AG, Mathewson K, Keikhosravi A, Eickhoff JC, Liu Y, Weber SM, Kao WJ, Eliceiri KW (2016) Highly aligned stromal collagen is a negative prognostic factor following pancreatic ductal adenocarcinoma resection. Oncotarget 7(46):76197–76213. https://doi.org/10.18632/oncotarget.12772
Sachs N, de Ligt J, Kopper O, Gogola E, Bounova G, Weeber F, Balgobind AV, Wind K, Gracanin A, Begthel H, Korving J, van Boxtel R, Duarte AA, Lelieveld D, van Hoeck A, Ernst RF, Blokzijl F, Nijman IJ, Hoogstraat M, van de Ven M, Egan DA, Zinzalla V, Moll J, Boj SF, Voest EE, Wessels L, van Diest PJ, Rottenberg S, Vries RGJ, Cuppen E, Clevers H (2018) A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172(1–2):373–386.e310. https://doi.org/10.1016/j.cell.2017.11.010
Sato T, Clevers H (2013) Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science (New York, NY) 340(6137):1190–1194. https://doi.org/10.1126/science.1234852
Gunti S, Hoke ATK, Vu KP, London NR Jr (2021) Organoid and spheroid tumor models: techniques and applications. Cancer 13(4):874. https://doi.org/10.3390/cancers13040874
Benton G, Kleinman HK, George J, Arnaoutova I (2011) Multiple uses of basement membrane-like matrix (BME/Matrigel) in vitro and in vivo with cancer cells. Int J Cancer 128(8):1751–1757. https://doi.org/10.1002/ijc.25781
Wolf K, Alexander S, Schacht V, Coussens LM, von Andrian UH, van Rheenen J, Deryugina E, Friedl P (2009) Collagen-based cell migration models in vitro and in vivo. Semin Cell Dev Biol 20(8):931–941. https://doi.org/10.1016/j.semcdb.2009.08.005
Sachs N, Clevers H (2014) Organoid cultures for the analysis of cancer phenotypes. Curr Opin Genet Dev 24:68–73. https://doi.org/10.1016/j.gde.2013.11.012
Ponsioen B, Post JB, Buissant des Amorie JR, Laskaris D, van Ineveld RL, Kersten S, Bertotti A, Sassi F, Sipieter F, Cappe B, Mertens S, Verlaan-Klink I, Boj SF, RGJ V, Rehmann H, Vandenabeele P, Riquet FB, Trusolino L, Bos JL, HJG S (2021) Quantifying single-cell ERK dynamics in colorectal cancer organoids reveals EGFR as an amplifier of oncogenic MAPK pathway signalling. Nat Cell Biol 23(4):377–390. https://doi.org/10.1038/s41556-021-00654-5
Gjorevski NS, Piotrowski A, Varner VD, Nelson CM (2015) Dynamic tensile forces drive collective cell migration through three-dimensional extracellular matrices. Sci Rep 5(1):11458. https://doi.org/10.1038/srep11458
Wolf K, Wu YI, Liu Y, Geiger J, Tam E, Overall C, Stack MS, Friedl P (2007) Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat Cell Biol 9(8):893–904. https://doi.org/10.1038/ncb1616
Provenzano PP, Eliceiri KW, Campbell JM, Inman DR, White JG, Keely PJ (2006) Collagen reorganization at the tumor-stromal interface facilitates local invasion. BMC Med 4(1):38–38. https://doi.org/10.1186/1741-7015-4-38
Riching KM, Cox BL, Salick MR, Pehlke C, Riching AS, Ponik SM, Bass BR, Crone WC, Jiang Y, Weaver AM, Eliceiri KW, Keely PJ (2014) 3D collagen alignment limits protrusions to enhance breast cancer cell persistence. Biophys J 107(11):2546–2558. https://doi.org/10.1016/j.bpj.2014.10.035
Khalil AA, Ilina O, Vasaturo A, Venhuizen J-H, Vullings M, Venhuizen V, Bilos A, Figdor CG, Span PN, Friedl P (2020) Collective invasion induced by an autocrine purinergic loop through connexin-43 hemichannels. J Cell Biol 219(10). https://doi.org/10.1083/jcb.201911120
Cheung KJ, Gabrielson E, Werb Z, Ewald AJ (2013) Collective invasion in breast cancer requires a conserved basal epithelial program. Cell 155(7):1639–1651. https://doi.org/10.1016/j.cell.2013.11.029
Beerling E, Seinstra D, de Wit E, Kester L, van der Velden D, Maynard C, Schäfer R, van Diest P, Voest E, van Oudenaarden A, Vrisekoop N, van Rheenen J (2016) Plasticity between epithelial and mesenchymal states unlinks EMT from metastasis-enhancing stem cell capacity. Cell Rep 14(10):2281–2288. https://doi.org/10.1016/j.celrep.2016.02.034
Padmanaban V, Grasset EM, Neumann NM, Fraser AK, Henriet E, Matsui W, Tran PT, Cheung KJ, Georgess D, Ewald AJ (2020) Organotypic culture assays for murine and human primary and metastatic-site tumors. Nat Protoc 15(8):2413–2442. https://doi.org/10.1038/s41596-020-0335-3
Kural MH, Billiar KL (2013) Regulating tension in three-dimensional culture environments. Exp Cell Res 319(16):2447–2459. https://doi.org/10.1016/j.yexcr.2013.06.019
Grinnell F, Petroll WM (2010) Cell motility and mechanics in three-dimensional collagen matrices. Annu Rev Cell Dev Biol 26:335–361. https://doi.org/10.1146/annurev.cellbio.042308.113318
Kleinman HK, McGarvey ML, Hassell JR, Star VL, Cannon FB, Laurie GW, Martin GR (1986) Basement membrane complexes with biological activity. Biochemistry 25(2):312–318. https://doi.org/10.1021/bi00350a005
Kleinman HK, McGarvey ML, Liotta LA, Robey PG, Tryggvason K, Martin GR (1982) Isolation and characterization of type IV procollagen, laminin, and heparan sulfate proteoglycan from the EHS sarcoma. Biochemistry 21(24):6188–6193. https://doi.org/10.1021/bi00267a025
Ilina O, Gritsenko PG, Syga S, Lippoldt J, La Porta CAM, Chepizhko O, Grosser S, Vullings M, Bakker G-J, Starruß J, Bult P, Zapperi S, Käs JA, Deutsch A, Friedl P (2020) Cell-cell adhesion and 3D matrix confinement determine jamming transitions in breast cancer invasion. Nat Cell Biol 22(9):1103–1115. https://doi.org/10.1038/s41556-020-0552-6
Stenn KS, Link R, Moellmann G, Madri J, Kuklinska E (1989) Dispase, a neutral protease from Bacillus polymyxa, is a powerful fibronectinase and type IV collagenase. J Invest Dermatol 93(2):287–290. https://doi.org/10.1111/1523-1747.ep12277593
Wolf K, te Lindert M, Krause M, Alexander S, te Riet J, Willis AL, Hoffman RM, Figdor CG, Weiss SJ, Friedl P (2013) Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force. J Cell Biol 201(7):1069–1084. https://doi.org/10.1083/jcb.201210152
Mason BN, Starchenko A, Williams RM, Bonassar LJ, Reinhart-King CA (2013) Tuning three-dimensional collagen matrix stiffness independently of collagen concentration modulates endothelial cell behavior. Acta Biomater 9(1):4635–4644. https://doi.org/10.1016/j.actbio.2012.08.007
Acknowledgments
This work is sponsored by the European Union’s Horizon 2020 FET Proactive program under the grant agreement No. 731957 (MECHANO-CONTROL) and by the Dutch Cancer Society (KWF) Young Investigator Grant 2020-13552 (AAK). We are grateful for Dr. Johan de Rooij for the helpful discussions and comments about the protocols and manuscript. We thank Colinda Scheele and Laura Bornes (van Rheenen Lab) for the isolation of breast cancer organoids from mice and Dr. Fried Zwartkruis and Denise Westland for assistance with Crispr/Cas9-mediated gene knockout. We also thank Marjolein Vliem, Marjolein Lugtigheid, Livio Kleij, Ingrid Verlaan, and Dr. Gerard van der Krogt for technical support and Kitty van Zwieten for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2023 The Authors
About this protocol
Cite this protocol
Smits, D., Khalil, A.A. (2023). Multimodal Techniques to Study Tumor Growth, Basement Membrane Breaching, and Invasion in 3D Matrices. In: Margadant, C. (eds) Cell Migration in Three Dimensions. Methods in Molecular Biology, vol 2608. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2887-4_17
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2887-4_17
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2886-7
Online ISBN: 978-1-0716-2887-4
eBook Packages: Springer Protocols