Abstract
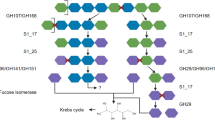
Sustainable use of natural products is one of the key challenges for the future. An increasing focus is on marine organic matter, mostly algae. New biotechnological tools for processing high amounts of micro- and macroalgae are necessary for efficient industrial degradation of marine matter. Secreted glycosyl hydrolases can be enriched and tested on the specific algae cell wall polymers of all algae groups (Rhodophyta; Phaeophyceae; Chlorophyta/Charophyta). Metagenomic analyses established new possibilities to screen algae-associated microbiomes for novel degrading enzymes in combination with sequence-based function prediction.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bórawski P, Bełdycka-Bórawska A, Szymańska EJ et al (2019) Development of renewable energy sources market and biofuels in the European Union. J Clean Prod 228:467–484
Daroń M, Wilk M (2021) Management of energy sources and the development potential in the energy production sector—a comparison of EU countries. Energies 14:685
Smith P, Gregory PJ (2013) Climate change and sustainable food production. Proc Nutr Soc 72:21–28
Pfeuty B, Thommen Q, Corellou F et al (2012) Circadian clocks in changing weather and seasons: lessons from the picoalga Ostreococcus tauri. BioEssays 34:781–790
Bolton JJ (2010) The biogeography of kelps (Laminariales, Phaeophyceae): a global analysis with new insights from recent advances in molecular phylogenetics. Helgol Mar Res 64:263–279
Field B, Randerson JT et al (1998) Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281:237–240
Arrigo KR (2005) Marine microorganisms and global nutrient cycles. Nature 437:349–355
Adeniyi OM, Azimov U, Burluka A (2018) Algae biofuel: current status and future applications. Renew Sust Energ Rev 90:316–335
Schmidt TM, DeLong EF, Pace NR (1991) Analysis of a marine picoplankton community by 16S rRNA gene cloning and sequencing. J Bacteriol 1991:4371–4378
Ferrer M, Golyshina OV, Chernikova TN et al (2005) Novel hydrolase diversity retrieved from a metagenome library of bovine rumen microflora. Environ Microbiol 7:1996–2010
Ferrer M, Golyshina OV, Plou FJ et al (2005) A novel alpha-glucosidase from the acidophilic archaeon Ferroplasma acidiphilum strain Y with high transglycosylation activity and an unusual catalytic nucleophile. Biochem J 391:269–276
Beloqui A, Pita M, Polaina J et al (2006) Novel polyphenol oxidase mined from a metagenome expression library of bovine rumen: biochemical properties, structural analysis, and phylogenetic relationships. J Biol Chem 281:22933–22942
Voget S, Leggewie C, Uesbeck A et al (2003) Prospecting for novel biocatalysts in a soil metagenome. Appl Environ Microbiol 69:6235–6242
Streit WR, Schmitz RA (2004) Metagenomics - the key to the uncultured microbes. Curr Opin Microbiol 7:492–498
Schmeisser C, Steele H, Streit WR (2007) Metagenomics, biotechnology with non-culturable microbes. Appl Microbiol Biotechnol 75:955–962
Daniel R (2004) The soil metagenome – a rich resource for the discovery of novel natural products. Curr Opin Biotechnol 15:199–204
Krohn I, Bergmann L, Qi M et al (2021) Deep (Meta)genomics and (Meta)transcriptome analyses of fungal and bacteria consortia from aircraft tanks and kerosene identify key genes in fuel and tank corrosion. Front Microbiol 12:722259
Krohn-Molt I, Wemheuer B, Alawi M et al (2013) Metagenome survey of a multispecies and alga-associated biofilm revealed key elements of bacterial-algal interactions in photobioreactors. Appl Environ Microbiol 79:6196–6206
Bari L, Yeasmin S (2021) Microbes culture methods. In: Chaplan MJ (ed) Reference module in biomedical sciences. Elsevier
Bäumgen M, Dutschei T, Bornscheuer UT (2021) Marine polysaccharides: occurrence, enzymatic degradation and utilization. Chembiochem 22:2247–2256
Deniaud-Bouët E, Kervarec N, Michel G et al (2014) Chemical and enzymatic fractionation of cell walls from Fucales: insights into the structure of the extracellular matrix of brown algae. Ann Bot 114:1203–1216
Quatrano RS, Stevens PT (1976) Cell wall assembly in fucus zygotes. Plant Pysiol 1976:224–231
Vreeland V, Waite JH, Epstein L (1998) Polyphenols and oxidases in substratum adhesion by marine algae and mussels. J Phycol 1998:1–8
Wiencke C, Clayton MN, Schoenwaelder M (2004) Sensitivity and acclimation to UV radiation of zoospores from five species of Laminariales from the Arctic. Mar Biol 145:31–39
Verhaeghe EF, Fraysse A, Guerquin-Kern J-L et al (2008) Microchemical imaging of iodine distribution in the brown alga Laminaria digitata suggests a new mechanism for its accumulation. J Biol Inorg Chem 13:257–269
Drula E, Garron M-L, Dogan S et al (2022) The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577
Berteau O, Mulloy B (2003) Sulfated fucans, fresh perspectives: structures, functions, and biological properties of sulfated fucans and an overview of enzymes active toward this class of polysaccharide. Glycobiology 13:29R–40R
Kadam SU, Tiwari BK, O’Donnell CP (2015) Extraction, structure and biofunctional activities of laminarin from brown algae. Int J Food Sci Technol 50:24–31
Lee KY, Mooney DJ (2012) Alginate: properties and biomedical applications. Prog Polym Sci 37:106–126
Mohnen D (2008) Pectin structure and biosynthesis. Curr Opin Plant Biol 11:266–277
Moreira LRS, Filho EXF (2008) An overview of mannan structure and mannan-degrading enzyme systems. Appl Microbiol Biotechnol 79:165–178
Bastawde KB (1992) Xylan structure, microbial xylanases, and their mode of action. World J Microbiol Biotechnol 1992:353–368
Béguin P, Aubert J-P (1994) The biological degradation of cellulose. FEMS Microbiol Rev 1994:25–58
Necas J, Bartosikova L (2013) Carageenan: a review. Vet Med 2013:187–205
Juanes JA (1991) International workshop on Gelidium: proceedings of the international workshop on Gelidium Held in Santander, Spain, September 3–8 1990, Developments in Hydrobiology Series, vol 68. Springer, Dordrecht
Morrice LM, McLean MW, Long WF et al (1984) Porphyran primary structure. In: Bird CJ, Ragan MA (eds) Eleventh international seaweed symposium. Springer, Dordrecht, pp 572–575
Jönsson M, Allahgholi L, Sardari RR et al (2020) Extraction and modification of macroalgal polysaccharides for current and Next-Generation applications. Molecules 2020:930
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Macdonald, J.F.H., Krohn, I., Streit, W.R. (2023). Screening Metagenomes for Algae Cell Wall Carbohydrates Degrading Hydrolases in Enrichment Cultures. In: Streit, W.R., Daniel, R. (eds) Metagenomics. Methods in Molecular Biology, vol 2555. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2795-2_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2795-2_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2794-5
Online ISBN: 978-1-0716-2795-2
eBook Packages: Springer Protocols