Abstract
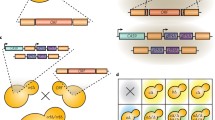
Complex haploinsufficiency refers to the genetic interaction that occurs in strains with heterozygous mutations at two different loci (a double heterozygous deletion mutant). Double heterozygous deletion mutants can be used to identify gene partners that act within the same pathway or to determine expression-dependent genetic interactions that result in phenotypic changes outside of what would be expected based on the phenotypes of the single heterozygous deletion mutants. The approach outlined here uses a lithium acetate transformation method on a parental “query” strain to introduce a transcription factor deletion DNA construct that is derived from the Homann et al. Candida albicans transcription factor deletion library (Homann et al. PLoS Genet 5(12):e1000783, 2009). We also outline the steps to confirming the genotype of the resulting transformants as well as an example of the use of double heterozygous deletion mutants for complex haploinsufficiency analysis of biofilm formation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Costanzo M et al (2019) Global genetic networks and the genotype-to-phenotype relationship. Cell 177(1):85–100. https://doi.org/10.1016/j.cell.2019.01.033
Hickman MA et al (2013) The “obligate diploid” Candida albicans forms mating-competent haploids. Nature 494(7435):55–59. https://doi.org/10.1038/nature11865
Shapiro RS et al (2018) A CRISPR–Cas9-based gene drive platform for genetic interaction analysis in Candida albicans. Nat Microbiol 3(1):73–82. https://doi.org/10.1038/s41564-017-0043-0
Glazier VE, Krysan DJ (2020) Genetic interaction analysis comes to the diploid human pathogen Candida albicans. PLoS Pathog 16(4):e1008399. https://doi.org/10.1371/journal.ppat.1008399
Glazier VE et al (2017) Genetic analysis of the Candida albicans biofilm transcription factor network using simple and complex haploinsufficiency. PLOS Genet 13(8):e1006948. https://doi.org/10.1371/journal.pgen.1006948
Forsberg SKG, Bloom JS, Sadhu MJ, Kruglyak L, Carlborg Ö (2017) Accounting for genetic interactions improves modeling of individual quantitative trait phenotypes in yeast. Nat Genet 49(4):497–503. https://doi.org/10.1038/ng.3800
Li SC, Diakov TT, Rizzo JM, Kane PM (2012) Vacuolar H+-ATPase works in parallel with the HOG pathway to adapt Saccharomyces cerevisiae cells to osmotic stress. Eukaryot Cell 11(3):282–291. https://doi.org/10.1128/EC.05198-11
Bharucha N et al (2011) A large-scale complex haploinsufficiency-based genetic interaction screen in Candida albicans: analysis of the RAM network during morphogenesis. PLOS Genet 7(4):e1002058. https://doi.org/10.1371/journal.pgen.1002058
Glazier VE et al (2018) Systematic complex haploinsufficiency-based genetic analysis of Candida albicans transcription factors: tools and applications to virulence-associated phenotypes. G3 (Bethesda) 8(4):1299–1314. https://doi.org/10.1534/g3.117.300515
Homann OR, Dea J, Noble SM, Johnson AD (2009) A phenotypic profile of the Candida albicans regulatory network. PLOS Genet 5(12):e1000783. https://doi.org/10.1371/journal.pgen.1000783
Glazier VE, Krysan DJ (2018) Transcription factor network efficiency in the regulation of Candida albicans biofilms: it is a small world. Curr Genet 64(4):883–888. https://doi.org/10.1007/s00294-018-0804-1
Noble SM, Johnson AD (2005) Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot Cell 4(2):298–309. https://doi.org/10.1128/EC.4.2.298-309.2005
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Glazer, V., Krysan, D. (2022). Construction of Double Heterozygous Deletion Strains for Complex Haploinsufficiency-Based Genetic Analysis in Candida albicans. In: Calderone, R. (eds) Candida Species. Methods in Molecular Biology, vol 2542. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2549-1_6
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2549-1_6
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2548-4
Online ISBN: 978-1-0716-2549-1
eBook Packages: Springer Protocols