Abstract
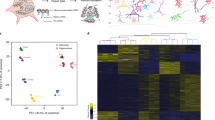
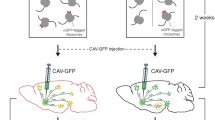
Over the past years, technological advances in transcriptomics provided deep insights into gene expression programs and their role in tissue organization and cellular functions. The isolation of ribosome-associated transcripts is a powerful approach for deep profiling of cell type–specific transcripts, and particularly well-suited for quantitative analysis of transcript isoforms. This method employs conditional ribosome epitope-tagging in genetically defined cell types, followed by affinity-isolation of ribosome-associated mRNAs. Advantages of this approach are twofold: first, the method enables rapid retrieval of mRNAs without tissue dissociation and cell sorting steps. Second, capturing of ribosome-associated mRNAs, enriches for transcripts recruited for active translation, therefore providing an approximation to the cellular translatome. Here, we describe one application of this method for the identification of the transcriptome of excitatory neuronal cells (mitral and tufted cells) of the mouse olfactory bulb, through RiboTag isolation from the vGlut2-IRES-cre mouse line as genetic driver of endogenously tagged ribosome expression.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Zeisel A, Hochgerner H, Lonnerberg P, Johnsson A, Memic F, van der Zwan J, Haring M, Braun E, Borm LE, La Manno G, Codeluppi S, Furlan A, Lee K, Skene N, Harris KD, Hjerling-Leffler J, Arenas E, Ernfors P, Marklund U, Linnarsson S (2018) Molecular architecture of the mouse nervous system. Cell 174(4):999–1014 e1022
Zeng H, Sanes JR (2017) Neuronal cell-type classification: challenges, opportunities and the path forward. Nat Rev Neurosci 18(9):530–546
Nguyen TM, Schreiner D, Xiao L, Traunmuller L, Bornmann C, Scheiffele P (2016) An alternative splicing switch shapes neurexin repertoires in principal neurons versus interneurons in the mouse hippocampus. eLife 5:e22757
Wamsley B, Jaglin XH, Favuzzi E, Quattrocolo G, Nigro MJ, Yusuf N, Khodadadi-Jamayran A, Rudy B, Fishell G (2018) Rbfox1 mediates cell-type-specific splicing in cortical interneurons. Neuron 100(4):846–859.e7
Gonatopoulos-Pournatzis T, Niibori R, Salter EW, Weatheritt RJ, Tsang B, Farhangmehr S, Liang X, Braunschweig U, Roth J, Zhang S, Henderson T, Sharma E, Quesnel-Vallieres M, Permanyer J, Maier S, Georgiou J, Irimia M, Sonenberg N, Forman-Kay JD, Gingras AC, Collingridge GL, Woodin MA, Cordes SP, Blencowe BJ (2020) Autism-Misregulated eIF4G microexons control synaptic translation and higher order cognitive functions. Mol Cell 77(6):1176–1192 e1116
Parikshak NN, Swarup V, Belgard TG, Irimia M, Ramaswami G, Gandal MJ, Hartl C, Leppa V, Ubieta LT, Huang J, Lowe JK, Blencowe BJ, Horvath S, Geschwind DH (2016) Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism. Nature 540(7633):423–427
Furlanis E, Scheiffele P (2018) Regulation of neuronal differentiation, function, and plasticity by alternative splicing. Annu Rev Cell Dev Biol 34:451–469
Vogel C, Marcotte EM (2012) Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet 13(4):227–232
Liu Y, Beyer A, Aebersold R (2016) On the dependency of cellular protein levels on mRNA abundance. Cell 165(3):535–550
Weatheritt RJ, Sterne-Weiler T, Blencowe BJ (2016) The ribosome-engaged landscape of alternative splicing. Nat Struct Mol Biol 23(12):1117–1123
Crouch EE, Doetsch F (2018) FACS isolation of endothelial cells and pericytes from mouse brain microregions. Nat Protoc 13(4):738–751
Tischfield DJ, Anderson SA (2017) Differentiation of mouse embryonic stem cells into cortical interneuron precursors. J Vis Exp 130:56358
van den Brink SC, Sage F, Vertesy A, Spanjaard B, Peterson-Maduro J, Baron CS, Robin C, van Oudenaarden A (2017) Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat Methods 14(10):935–936
Heiman M, Schaefer A, Gong S, Peterson JD, Day M, Ramsey KE, Suarez-Farinas M, Schwarz C, Stephan DA, Surmeier DJ, Greengard P, Heintz N (2008) A translational profiling approach for the molecular characterization of CNS cell types. Cell 135(4):738–748
Sanz E, Yang L, Su T, Morris DR, McKnight GS, Amieux PS (2009) Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc Natl Acad Sci U S A 106(33):13939–13944
Thomas A, Lee PJ, Dalton JE, Nomie KJ, Stoica L, Costa-Mattioli M, Chang P, Nuzhdin S, Arbeitman MN, Dierick HA (2012) A versatile method for cell-specific profiling of translated mRNAs in drosophila. PLoS One 7(7):e40276
Gregory JA, Hoelzli E, Abdelaal R, Braine C, Cuevas M, Halpern M, Barretto N, Schrode N, Akbalik G, Kang K, Cheng E, Bowles K, Lotz S, Goderie S, Karch CM, Temple S, Goate A, Brennand KJ, Phatnani H (2020) Cell type-specific in vitro gene expression profiling of stem cell-derived neural models. Cell 9(6):1406
Mardinly AR, Spiegel I, Patrizi A, Centofante E, Bazinet JE, Tzeng CP, Mandel-Brehm C, Harmin DA, Adesnik H, Fagiolini M, Greenberg ME (2016) Sensory experience regulates cortical inhibition by inducing IGF1 in VIP neurons. Nature 531(7594):371–375
Thomson SR, Seo SS, Barnes SA, Louros SR, Muscas M, Dando O, Kirby C, Wyllie DJA, Hardingham GE, Kind PC, Osterweil EK (2017) Cell-type-specific translation profiling reveals a novel strategy for treating fragile X syndrome. Neuron 95(3):550–563 e555
Gonzalez C, Sims JS, Hornstein N, Mela A, Garcia F, Lei L, Gass DA, Amendolara B, Bruce JN, Canoll P, Sims PA (2014) Ribosome profiling reveals a cell-type-specific translational landscape in brain tumors. J Neurosci 34(33):10924–10936
Furlanis E, Traunmuller L, Fucile G, Scheiffele P (2019) Landscape of ribosome-engaged transcript isoforms reveals extensive neuronal-cell-class-specific alternative splicing programs. Nat Neurosci 22(10):1709–1717
Lee H, Fenster RJ, Pineda SS, Gibbs WS, Mohammadi S, Davila-Velderrain J, Garcia FJ, Therrien M, Novis HS, Gao F, Wilkinson H, Vogt T, Kellis M, LaVoie MJ, Heiman M (2020) Cell type-specific transcriptomics reveals that mutant huntingtin leads to mitochondrial RNA release and neuronal innate immune activation. Neuron 107(5):891–908 e898
Luo L, Ambrozkiewicz MC, Benseler F, Chen C, Dumontier E, Falkner S, Furlanis E, Gomez AM, Hoshina N, Huang WH, Hutchison MA, Itoh-Maruoka Y, Lavery LA, Li W, Maruo T, Motohashi J, Pai EL, Pelkey KA, Pereira A, Philips T, Sinclair JL, Stogsdill JA, Traunmuller L, Wang J, Wortel J, You W, Abumaria N, Beier KT, Brose N, Burgess HA, Cepko CL, Cloutier JF, Eroglu C, Goebbels S, Kaeser PS, Kay JN, Lu W, Luo L, Mandai K, McBain CJ, Nave KA, Prado MAM, Prado VF, Rothstein J, Rubenstein JLR, Saher G, Sakimura K, Sanes JR, Scheiffele P, Takai Y, Umemori H, Verhage M, Yuzaki M, Zoghbi HY, Kawabe H, Craig AM (2020) Optimizing nervous system-specific gene targeting with Cre driver lines: prevalence of germline recombination and influencing factors. Neuron 106(1):37–65 e35
Acknowledgments
We are thankful to O. Mauger and E. Furlanis for constructive comments on the chapter manuscript and to T.M. Nguyen, L. Traunmüller, E. Furlanis, and S. Falkner for discussions and help in setting up the protocol for RiboTRAP purifications. Work in the laboratory was supported by funds to P.S. from the Swiss National Science Foundation, a European Research Council Advanced Grant (SPLICECODE), EU-AIMS and AIMS-2-TRIALS which are supported by the Innovative Medicines Initiatives from the European Commission. The results leading to this publication has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement no. 777394. This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA and AUTISM SPEAKS, Autistica, SFARI. The Scheiffele Laboratory is an associate member of the Swiss National Science Foundation’s National Competence Centre for Research (NCCR) RNA and Disease.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Di Bartolomei, G., Scheiffele, P. (2022). An Optimized Protocol for the Mapping of Cell Type–Specific Ribosome-Associated Transcript Isoforms from Small Mouse Brain Regions. In: Scheiffele, P., Mauger, O. (eds) Alternative Splicing. Methods in Molecular Biology, vol 2537. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2521-7_3
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2521-7_3
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2520-0
Online ISBN: 978-1-0716-2521-7
eBook Packages: Springer Protocols