Abstract
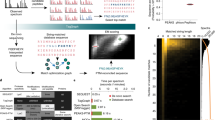
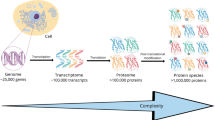
Post-translational modifications (PTMs) regulate complex biological processes through the modulation of protein activity, stability, and localization. Insights into the specific modification type and localization within a protein sequence can help ascertain functional significance. Computational models are increasingly demonstrated to offer a low-cost, high-throughput method for comprehensive PTM predictions. Algorithms are optimized using existing experimental PTM data, thus accurate prediction performance relies on the creation of robust datasets. Herein, advancements in mass spectrometry–based proteomics technologies to maximize PTM coverage are reviewed. Further, requisite experimental validation approaches for PTM predictions are explored to ensure that follow-up mechanistic studies are focused on accurate modification sites.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Qi HH, Ongusaha PP, Myllyharju J et al (2008) Prolyl 4-hydroxylation regulates argonaute 2 stability. Nature 455(7211):421–424. https://doi.org/10.1038/nature07186
Sahar S, Zocchi L, Kinoshita C et al (2010) Regulation of BMAL1 protein stability and circadian function by GSK3β-mediated phosphorylation. PLoS One 5(1):e8561. https://doi.org/10.1371/journal.pone.0008561
Deribe YL, Pawson T, Dikic I (2010) Post-translational modifications in signal integration. Nat Struct Mol Biol 17(6):666–672. https://doi.org/10.1038/nsmb.1842
Liu J, Qian C, Cao X (2016) Post-translational modification control of innate immunity. Immunity 45(1):15–30. https://doi.org/10.1016/j.immuni.2016.06.020
Ahearn IM, Haigis K, Bar-Sagi D et al (2012) Regulating the regulator: post-translational modification of RAS. Nat Rev Mol Cell Biol 13(1):39–51. https://doi.org/10.1038/nrm3255
Xie Y, Kang R, Sun X et al (2015) Posttranslational modification of autophagy-related proteins in macroautophagy. Autophagy 11(1):28–45. https://doi.org/10.4161/15548627.2014.984267
Smith LM, Kelleher NL (2018) Proteoforms as the next proteomics currency: identifying precise molecular forms of proteins can improve our understanding of function. Science 359(6380):1106–1107. https://doi.org/10.1126/science.aat1884
Khoury GA, Baliban RC, Floudas CA (2011) Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci Rep 1(1):1–5. https://doi.org/10.1038/srep00090
Audagnotto M, Dal Peraro M (2017) Protein post-translational modifications: in silico prediction tools and molecular modeling. Comput Struct Biotechnol J 15:307–319. https://doi.org/10.1016/j.csbj.2017.03.004
Choudhary C, Mann M (2010) Decoding signalling networks by mass spectrometry-based proteomics. Nat Rev Mol Cell Biol 11(6):427–439. https://doi.org/10.1038/nrm2900
Olsen JV, Mann M (2013) Status of large-scale analysis of posttranslational modifications by mass spectrometry. Mol Cell Proteomics 12(12):3444–3452. https://doi.org/10.1074/mcp.O113.034181
Zhang Y, Zhang C, Jiang H et al (2015) Fishing the PTM proteome with chemical approaches using functional solid phases. Chem Soc Rev 44(22):8260–8287. https://doi.org/10.1039/c4cs00529e
Zhang Y, Fonslow BR, Shan B et al (2013) Protein analysis by shotgun/bottom-up proteomics. Chem Rev 113(4):2343–2394. https://doi.org/10.1021/cr3003533
Gillet LC, Leitner A, Aebersold R (2016) Mass spectrometry applied to bottom-up proteomics: entering the high-throughput era for hypothesis testing. Annu Rev Anal Chem 9(1):449–472. https://doi.org/10.1146/annurev-anchem-071015-041535
Stahl DC, Swiderek KM, Davis MT et al (1996) Data-controlled automation of liquid chromatography/tandem mass spectrometry analysis of peptide mixtures. J Am Soc Mass Spectrom 7(6):532–540. https://doi.org/10.1016/1044-0305(96)00057-8
Singh C, Zampronio CG, Creese AJ et al (2012) Higher energy collision dissociation (HCD) product ion-triggered electron transfer dissociation (ETD) mass spectrometry for the analysis of N-linked glycoproteins. J Proteome Res 11(9):4517–4525. https://doi.org/10.1021/pr300257c
Frese CK, Altelaar AFM, Hennrich ML et al (2011) Improved peptide identification by targeted fragmentation using CID, HCD and ETD on an LTQ-orbitrap velos. J Proteome Res 10(5):2377–2388. https://doi.org/10.1021/pr1011729
Perkins DN, Pappin DJC, Creasy DM et al (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20(18):3551–3567. 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2
Tabb DL, Eng JK, Yates JR (2001) Protein identification by SEQUEST. Springer, Berlin, pp 125–142
Cox J, Neuhauser N, Michalski A et al (2011) Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res 10(4):1794–1805. https://doi.org/10.1021/pr101065j
Nesvizhskii AI (2010) A survey of computational methods and error rate estimation procedures for peptide and protein identification in shotgun proteomics. J Proteomics 73(11):2092–2123. https://doi.org/10.1016/j.jprot.2010.08.009
Chalkley RJ, Clauser KR (2012) Modification site localization scoring: strategies and performance. Mol Cell Proteomics 11(5):3–14. https://doi.org/10.1074/mcp.R111.015305
Needham EJ, Parker BL, Burykin T et al (2019) Illuminating the dark phosphoproteome. Sci Signal 12(565):8645. https://doi.org/10.1126/scisignal.aau8645
Xiao H, Chen W, Smeekens JM et al (2018) An enrichment method based on synergistic and reversible covalent interactions for large-scale analysis of glycoproteins. Nat Commun 9(1):1–12. https://doi.org/10.1038/s41467-018-04081-3
Dang L, Jia L, Zhi Y et al (2019) Mapping human N-linked glycoproteins and glycosylation sites using mass spectrometry. Trends Anal Chem 114:143–150. https://doi.org/10.1016/j.trac.2019.02.009
Al-Barakati HJ, EW MC, Hicks LM et al (2018) SVM-SulfoSite: a support vector machine based predictor for sulfenylation sites. Sci Rep 8(1):1–9. https://doi.org/10.1038/s41598-018-29,126-x
Gnad F, Ren S, Cox J et al (2007) PHOSIDA (phosphorylation site database): management, structural and evolutionary investigation, and prediction of phosphosites. Genome Biol 8(11):R250. https://doi.org/10.1186/gb-2007-8-11-r250
Wang D, Liang Y, Xu D (2019) Capsule network for protein post-translational modification site prediction. Bioinformatics 35(14):2386–2394. https://doi.org/10.1093/bioinformatics/bty977
Hasan MM (2017) Prediction of protein post-translational modification sites: an overview. Ann Proteomics Bioinforma 2(1):49–57. https://doi.org/10.29328/journal.apb.1001005
Zhang N, Li BQ, Gao S et al (2012) Computational prediction and analysis of protein γ-carboxylation sites based on a random forest method. Mol Biosyst 8(11):2946–2955. https://doi.org/10.1039/c2mb25185j
Wang J-R, Huang W-L, Tsai M-J et al (2017) ESA-UbiSite: accurate prediction of human ubiquitination sites by identifying a set of effective negatives. Bioinformatics 33(5):btw701. https://doi.org/10.1093/bioinformatics/btw701
Chen QY, Tang J, Du PF (2017) Predicting protein lysine phosphoglycerylation sites by hybridizing many sequence based features. Mol Biosyst 13(5):874–882. https://doi.org/10.1039/c6mb00875e
Wysocki VH, Resing KA, Zhang Q et al (2005) Mass spectrometry of peptides and proteins. Methods 35(3):211–222. https://doi.org/10.1016/j.ymeth.2004.08.013
Issaq HJ, Conrads TP, Janini GM et al (2002) Methods for fractionation, separation and profiling of proteins and peptides. Electrophoresis 23(17):3048–3061. https://doi.org/10.1002/1522-2683(200209)23:17 < 3048::AID-ELPS3048 > 3.0.CO;2-L
Canbay V, auf dem Keller U (2021) New strategies to identify protease substrates. Curr Opin Chem Biol 60:89–96. https://doi.org/10.1016/j.cbpa.2020.09.009
Iannetta AA, Rogers HT, Al-Mohanna T et al (2021) Profiling thimet oligopeptidase-mediated proteolysis in Arabidopsis thaliana. Plant J 106(2):336–350. https://doi.org/10.1111/tpj.15165
Olsen JV, Ong SE, Mann M (2004) Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol Cell Proteomics 3(6):608–614. https://doi.org/10.1074/mcp.T400003-MCP200
Lodish H, Berk A, Zipursky SL et al (2000) Hierarchical structure of proteins. In: Molecular cell biology. W. H. Freeman, New York
Keil-Dlouhá V, Zylber N, Imhoff JM et al (1971) Proteolytic activity of pseudotrypsin. FEBS Lett 16(4):291–295. https://doi.org/10.1016/0014-5793(71)80373-3
Rice RH, Means GE, Brown WD (1977) Stabilization of bovine trypsin by reductive methylation. BBA Protein Struct 492(2):316–321. https://doi.org/10.1016/0005-2795(77)90082-4
Ma J, Liang Z, Qiao X et al (2008) Organic-inorganic hybrid silica monolith based immobilized trypsin reactor with high enzymatic activity. Anal Chem 80(8):2949–2956. https://doi.org/10.1021/ac702343a
Sun L, Zhu G, Yan X et al (2014) Uncovering immobilized trypsin digestion features from large-scale proteome data generated by high-resolution mass spectrometry. J Chromatogr A 1337:40–47. https://doi.org/10.1016/j.chroma.2014.02.014
Tran BQ, Hernandez C, Waridel P et al (2011) Addressing trypsin bias in large scale (pPhospho)proteome analysis by size exclusion chromatography and secondary digestion of large post-trypsin peptides. J Proteome Res 10(2):800–811. https://doi.org/10.1021/pr100951t
Imre T, Schlosser G, Pocsfalvi G et al (2005) Glycosylation site analysis of human alpha-1-acid glycoprotein (AGP) by capillary liquid chromatography—electrospray mass spectrometry. J Mass Spectrom 40(11):1472–1483. https://doi.org/10.1002/jms.938
Guo X, Trudgian DC, Lemoff A et al (2014) Confetti: a multiprotease map of the HeLa proteome for comprehensive proteomics. Mol Cell Proteomics 13(6):1573–1584. https://doi.org/10.1074/mcp.M113.035170
Swaney DL, Wenger CD, Coon JJ (2010) Value of using multiple proteases for large-scale mass spectrometry-based proteomics. J Proteome Res 9(3):1323–1329. https://doi.org/10.1021/pr900863u
Wiśniewski JR, Mann M (2012) Consecutive proteolytic digestion in an enzyme reactor increases depth of proteomic and phosphoproteomic analysis. Anal Chem 84(6):2631–2637. https://doi.org/10.1021/ac300006b
Bian Y, Ye M, Song C et al (2012) Improve the coverage for the analysis of phosphoproteome of HeLa cells by a tandem digestion approach. J Proteome Res 11(5):2828–2837. https://doi.org/10.1021/pr300242w
Zhao Y, Jensen ON (2009) Modification-specific proteomics: strategies for characterization of post-translational modifications using enrichment techniques. Proteomics 9(20):4632–4641. https://doi.org/10.1002/pmic.200900398
Huang J, Wang F, Ye M et al (2014) Enrichment and separation techniques for large-scale proteomics analysis of the protein post-translational modifications. J Chromatogr A 1372:1–17. https://doi.org/10.1016/j.chroma.2014.10.107
Xu H, Wang Y, Lin S et al (2018) PTMD: a database of human disease-associated post-translational modifications. Genomics Proteomics Bioinformatics 16(4):244–251. https://doi.org/10.1016/j.gpb.2018.06.004
Ward PS, Thompson CB (2012) Signaling in control of cell growth and metabolism. Cold Spring Harb Perspect Biol 4(7):1–15. https://doi.org/10.1101/cshperspect.a006783
Dhanasekaran DN, Premkumar Reddy E (2017) JNK-signaling: a multiplexing hub in programmed cell death. Genes Cancer 8(9–10):682–694. https://doi.org/10.18632/genesandcancer.155
Humphrey SJ, James DE, Mann M (2015) Protein phosphorylation: a major switch mechanism for metabolic regulation. Trends Endocrinol Metab 26(12):676–687. https://doi.org/10.1016/j.tem.2015.09.013
Dennis MD, Jefferson LS, Kimball SR (2012) Role of p70S6K1-mediated phosphorylation of eIF4B and PDCD4 proteins in the regulation of protein synthesis. J Biol Chem 287(51):42890–42899. https://doi.org/10.1074/jbc.M112.404822
Beilharz K, Nováková L, Fadda D et al (2012) Control of cell division in Streptococcus pneumoniae by the conserved Ser/Thr protein kinase StkP. Proc Natl Acad Sci U S A 109(15):E905–E913. https://doi.org/10.1073/pnas.1119172109
Lin S, Wang C, Zhou J et al (2021) EPSD: a well-annotated data resource of protein phosphorylation sites in eukaryotes. Brief Bioinform 22(1):298–307. https://doi.org/10.1093/bib/bbz169
Rush J, Moritz A, Lee KA et al (2005) Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat Biotechnol 23(1):94–101. https://doi.org/10.1038/nbt1046
Kaneko T, Huang H, Cao X et al (2012) Superbinder SH2 domains act as antagonists of cell signaling. Sci Signal 5(243):ra68. https://doi.org/10.1126/scisignal.2003021
Fuhs SR, Hunter T (2017) pHisphorylation: the emergence of histidine phosphorylation as a reversible regulatory modification. Curr Opin Cell Biol 45:8–16. https://doi.org/10.1016/j.ceb.2016.12.010
Cieśla J, Fraczyk T, Rode W (2011) Phosphorylation of basic amino acid residues in proteins: important but easily missed. Acta Biochim Pol 58(2):137–148. https://doi.org/10.18388/abp.2011_2258
Falke JJ, Bass RB, Butler SL et al (1997) The two-component signaling pathway of bacterial chemotaxis: a molecular view of signal transduction by receptors, kinases, and adaptation enzymes. Annu Rev Cell Dev Biol 13:457–512. https://doi.org/10.1146/annurev.cellbio.13.1.457
Hardman G, Perkins S, Brownridge PJ et al (2019) Strong anion exchange-mediated phosphoproteomics reveals extensive human non-canonical phosphorylation. EMBO J 38(21):e100847. https://doi.org/10.15252/embj.2018100847
Humphrey SJ, Azimifar SB, Mann M (2015) High-throughput phosphoproteomics reveals in vivo insulin signaling dynamics. Nat Biotechnol 33(9):990–995. https://doi.org/10.1038/nbt.3327
Liu JJ, Sharma K, Zangrandi L et al (2018) In vivo brain GPCR signaling elucidated by phosphoproteomics. Science 360(6395):eaao4927. https://doi.org/10.1126/science.aao4927
Tan H, Yang K, Li Y et al (2017) Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46(3):488–503. https://doi.org/10.1016/j.immuni.2017.02.010
Aasebø E, Mjaavatten O, Vaudel M et al (2016) Freezing effects on the acute myeloid leukemia cell proteome and phosphoproteome revealed using optimal quantitative workflows. J Proteomics 145:214–225. https://doi.org/10.1016/j.jprot.2016.03.049
Andersson L, Porath J (1986) Isolation of phosphoproteins by immobilized metal (Fe3+) affinity chromatography. Anal Biochem 154(1):250–254. https://doi.org/10.1016/0003-2697(86)90523-3
Jensen SS, Larsen MR (2007) Evaluation of the impact of some experimental procedures on different phosphopeptide enrichment techniques. Rapid Commun Mass Spectrom 21(22):3635–3645. https://doi.org/10.1002/rcm.3254
Ficarro SB, McCleland ML, Stukenberg PT et al (2002) Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat Biotechnol 20(3):301–305. https://doi.org/10.1038/nbt0302-301
Lai AC-Y, Tsai C-F, Hsu C-C et al (2012) Complementary Fe3+ and Ti4+ immobilized metal ion affinity chromatography for purification of acidic and basic phosphopeptides. Rapid Commun Mass Spectrom 26(18):2186–2194. https://doi.org/10.1002/rcm.6327
Tsai CF, Hsu CC, Hung JN et al (2014) Sequential phosphoproteomic enrichment through complementary metal-directed immobilized metal ion affinity chromatography. Anal Chem 86(1):685–693. https://doi.org/10.1021/ac4031175
Iliuk AB, Martin VA, Alicie BM et al (2010) In-depth analyses of kinase-dependent tyrosine phosphoproteomes based on metal ion-functionalized soluble nanopolymers. Mol Cell Proteomics 9(10):2162–2172. https://doi.org/10.1074/mcp.M110.000091
Xue L, Wang WH, Iliuk A et al (2012) Sensitive kinase assay linked with phosphoproteomics for identifying direct kinase substrates. Proc Natl Acad Sci U S A 109(15):5615–5620. https://doi.org/10.1073/pnas.1119418109
Leitner A (2010) Phosphopeptide enrichment using metal oxide affinity chromatography. Trends Anal Chem 29(2):177–185. https://doi.org/10.1016/j.trac.2009.08.007
Pinkse MWH, Uitto PM, Hilhorst MJ et al (2004) Selective isolation at the femtomole level of phosphopeptides from proteolytic digests using 2D-NanoLC-ESI-MS/MS and titanium oxide precolumns. Anal Chem 76(14):3935–3943. https://doi.org/10.1021/ac0498617
Werth EG, McConnell EW, Couso Lianez I et al (2019) Investigating the effect of target of rapamycin kinase inhibition on the Chlamydomonas reinhardtii phosphoproteome: from known homologs to new targets. New Phytol 221(1):247–260. https://doi.org/10.1111/nph.15339
Ma WF, Zhang C, Zhang YT et al (2014) Magnetic MSP@ZrO2 microspheres with yolk-shell structure: designed synthesis and application in highly selective enrichment of phosphopeptides. Langmuir 30(22):6602–6611. https://doi.org/10.1021/la501381v
Huang SY, Chen YC (2013) Magnetic nanoparticle-based platform for characterization of histidine-rich proteins and peptides. Anal Chem 85(6):3347–3354. https://doi.org/10.1021/ac4000128
Li Y, Leng T, Lin H et al (2007) Preparation of Fe3O4@ZrO2 core—shell microspheres as affinity probes for selective enrichment and direct determination of phosphopeptides using matrix-assisted laser desorption ionization mass spectrometry. J Proteome Res 6(11):4498–4510. https://doi.org/10.1021/pr070167s
Mann M, Ong SE, Grønborg M et al (2002) Analysis of protein phosphorylation using mass spectrometry: deciphering the phosphoproteome. Trends Biotechnol 20(6):261–268. https://doi.org/10.1016/S0167-7799(02)01944-3
Bodenmiller B, Mueller LN, Mueller M et al (2007) Reproducible isolation of distinct, overlapping segments of the phosphoproteome. Nat Methods 4(3):231–237. https://doi.org/10.1038/nmeth1005
Thingholm TE, Jensen ON, Robinson PJ et al (2008) SIMAC (sequential elution from IMAC), a phosphoproteomics strategy for the rapid separation of monophosphorylated from multiply phosphorylated peptides. Mol Cell Proteomics 7(4):661–671. https://doi.org/10.1074/mcp.M700362-MCP200
Peng J, Zhang H, Li X et al (2016) Dual-metal centered zirconium-organic framework: a metal-affinity probe for highly specific interaction with phosphopeptides. ACS Appl Mater Interfaces 8(51):35012–35020. https://doi.org/10.1021/acsami.6b12630
Ruprecht B, Koch H, Medard G et al (2015) Comprehensive and reproducible phosphopeptide enrichment using iron immobilized metal ion affinity chromatography (Fe-IMAC) columns. Mol Cell Proteomics 14(1):205–215. https://doi.org/10.1074/mcp.M114.043109
Beltran L, Cutillas PR (2012) Advances in phosphopeptide enrichment techniques for phosphoproteomics. Amino Acids 43(3):1009–1024. https://doi.org/10.1007/s00726-012-1288-9
Sugiyama N, Masuda T, Shinoda K et al (2007) Phosphopeptide enrichment by aliphatic hydroxy acid-modified metal oxide chromatography for nano-LC-MS/MS in proteomics applications. Mol Cell Proteomics 6(6):1103–1109. https://doi.org/10.1074/mcp.T600060-MCP200
Dubrovska A, Souchelnytskyi S (2005) Efficient enrichment of intact phosphorylated proteins by modified immobilized metal-affinity chromatography. Proteomics 5(18):4678–4683. https://doi.org/10.1002/pmic.200500002
Hwang L, Ayaz-Guner S, Gregorich ZR et al (2015) Specific enrichment of phosphoproteins using functionalized multivalent nanoparticles. J Am Chem Soc 137(7):2432–2435. https://doi.org/10.1021/ja511833y
Liu W, Zheng J, Li S et al (2015) Aluminium glycinate functionalized silica nanoparticles for highly specific separation of phosphoproteins. J Mater Chem B 3(31):6528–6535. https://doi.org/10.1039/c5tb01055a
Schieber M, Chandel NS (2014) ROS function in redox signaling and oxidative stress. Curr Biol 24(10):R453–R462. https://doi.org/10.1016/j.cub.2014.03.034
Poole LB, Nelson KJ (2008) Discovering mechanisms of signaling-mediated cysteine oxidation. Curr Opin Chem Biol 12(1):18–24. https://doi.org/10.1016/j.cbpa.2008.01.021
Couturier J, Chibani K, Jacquot JP et al (2013) Cysteine-based redox regulation and signaling in plants. Front Plant Sci 4:105. https://doi.org/10.3389/fpls.2013.00105
Paulech J, Solis N, Edwards AVG et al (2013) Large-scale capture of peptides containing reversibly oxidized cysteines by thiol-disulfide exchange applied to the myocardial redox proteome. Anal Chem 85(7):3774–3780. https://doi.org/10.1021/ac400166e
Guo J, Gaffrey MJ, Su D et al (2014) Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat Protoc 9(1):64–75. https://doi.org/10.1038/nprot.2013.161
Leonard SE, Carroll KS (2011) Chemical “omics” approaches for understanding protein cysteine oxidation in biology. Curr Opin Chem Biol 15(1):88–102. https://doi.org/10.1016/j.cbpa.2010.11.012
Murray CI, Van Eyk JE (2012) Chasing cysteine oxidative modifications: proteomic tools for characterizing cysteine redox status. Circ Cardiovasc Genet 5(5):591. https://doi.org/10.1161/CIRCGENETICS.111.961425
Jaffrey SR, Erdjument-Bromage H, Ferris CD et al (2001) Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat Cell Biol 3(2):193–197. https://doi.org/10.1038/35055104
McConnell EW, Werth EG, Hicks LM (2018) The phosphorylated redox proteome of Chlamydomonas reinhardtii: revealing novel means for regulation of protein structure and function. Redox Biol 17:35–46. https://doi.org/10.1016/j.redox.2018.04.003
Ford MM, Smythers AL, McConnell EW et al (2019) Inhibition of TOR in Chlamydomonas reinhardtii leads to rapid cysteine oxidation reflecting sustained physiological changes. Cells 8(10):1171. https://doi.org/10.3390/cells8101171
Smythers AL, McConnell EW, Lewis HC et al (2020) Photosynthetic metabolism and nitrogen reshuffling are regulated by reversible cysteine thiol oxidation following nitrogen deprivation in Chlamydomonas. Plants 9(6):784. https://doi.org/10.3390/plants9060784
Murray CI, Uhrigshardt H, O’Meally RN et al (2012) Identification and quantification of S-nitrosylation by cysteine reactive tandem mass tag switch assay. Mol Cell Proteomics 11(2):1–12. https://doi.org/10.1074/mcp.M111.013441
Poole LB, Zeng BB, Knaggs SA et al (2005) Synthesis of chemical probes to map sulfenic acid modifications on proteins. Bioconjug Chem 16(6):1624–1628. https://doi.org/10.1021/bc050257s
Akter S, Fu L, Jung Y et al (2018) Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. Nat Chem Biol 14(11):995–1004. https://doi.org/10.1038/s41589-018-0116-2
Seneviratne U, Nott A, Bhat VB et al (2016) S-nitrosation of proteins relevant to Alzheimer’s disease during early stages of neurodegeneration. Proc Natl Acad Sci U S A 113(15):4152–4157. https://doi.org/10.1073/pnas.1521318113
Martin BR, Cravatt BF (2009) Large-scale profiling of protein palmitoylation in mammalian cells. Nat Methods 6(2):135–138. https://doi.org/10.1038/nmeth.1293
Held JM (2020) Redox systems biology: harnessing the sentinels of the cysteine redoxome. Antioxid Redox Signal 32(10):659–676. https://doi.org/10.1089/ars.2019.7725
Shental-Bechor D, Levy Y (2008) Effect of glycosylation on protein folding: a close look at thermodynamic stabilization. Proc Natl Acad Sci U S A 105(24):8256–8261. https://doi.org/10.1073/pnas.0801340105
Rudd PM, Elliott T, Cresswell P et al (2001) Glycosylation and the immune system. Science 291(5512):2370–2376. https://doi.org/10.1126/science.291.5512.2370
Moremen KW, Tiemeyer M, Nairn AV (2012) Vertebrate protein glycosylation: diversity, synthesis and function. Nat Rev Mol Cell Biol 13(7):448–462. https://doi.org/10.1038/nrm3383
Zhu Y, Willems LI, Salas D et al (2020) Tandem bioorthogonal labeling uncovers endogenous cotranslationally O-GlcNAc modified nascent proteins. J Am Chem Soc 142(37):15729–15739. https://doi.org/10.1021/jacs.0c04121
Zhang H, Li X-J, Martin DB et al (2003) Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat Biotechnol 21(6):660–666. https://doi.org/10.1038/nbt827
Kameyama A, Thet Tin WW, Toyoda M et al (2019) A practical method of liberating O-linked glycans from glycoproteins using hydroxylamine and an organic superbase. Biochem Biophys Res Commun 513(1):186–192. https://doi.org/10.1016/j.bbrc.2019.03.144
Cao J, Shen C, Wang H et al (2009) Identification of N-glycosylation sites on secreted proteins of human hepatocellular carcinoma cells with a complementary proteomics approach. J Proteome Res 8(2):662–672. https://doi.org/10.1021/pr800826u
Xia C, Jiao F, Gao F et al (2018) Two-dimensional MoS2-based zwitterionic hydrophilic interaction liquid chromatography material for the specific enrichment of glycopeptides. Anal Chem 90(11):6651–6659. https://doi.org/10.1021/acs.analchem.8b00461
Nilsson J, Rüetschi U, Halim A et al (2009) Enrichment of glycopeptides for glycan structure and attachment site identification. Nat Methods 6(11):809–811. https://doi.org/10.1038/nmeth.1392
Calvano CD, Zambonin CG, Jensen ON (2008) Assessment of lectin and HILIC based enrichment protocols for characterization of serum glycoproteins by mass spectrometry. J Proteomics 71(3):304–317. https://doi.org/10.1016/j.jprot.2008.06.013
Zhang B, Yu RZ, Yu YH et al (2018) Lectin inspired polymers based on the dipeptide Ser-Asp for glycopeptide enrichment. Analyst 143(21):5090–5093. https://doi.org/10.1039/c8an01258j
Waniwan JT, Chen YJ, Capangpangan R et al (2018) Glycoproteomic alterations in drug-resistant nonsmall cell lung cancer cells revealed by lectin magnetic nanoprobe-based mass spectrometry. J Proteome Res 17(11):3761–3773. https://doi.org/10.1021/acs.jproteome.8b00433
Zeng Z, Hincapie M, Pitteri SJ et al (2011) A proteomics platform combining depletion, multi-lectin affinity chromatography (M-LAC), and isoelectric focusing to study the breast cancer proteome. Anal Chem 83(12):4845–4854. https://doi.org/10.1021/ac2002802
Plavina T, Wakshull E, Hancock WS et al (2007) Combination of abundant protein depletion and multi-lectin affinity chromatography (M-LAC) for plasma protein biomarker discovery. J Proteome Res 6(2):662–671. https://doi.org/10.1021/pr060413k
Imberty A, Mitchell EP, Wimmerová M (2005) Structural basis of high-affinity glycan recognition by bacterial and fungal lectins. Curr Opin Struct Biol 15(5):525–534. https://doi.org/10.1016/j.sbi.2005.08.003
Sparbier K, Koch S, Kessler I et al (2005) Selective isolation of glycoproteins and glycopeptides for MALDI-TOF MS detection supported by magnetic particles. J Biomol Tech 16(4):407–411
Qu Y, Liu J, Yang K et al (2012) Boronic acid functionalized core-shell polymer nanoparticles prepared by distillation precipitation polymerization for glycopeptide enrichment. Chem A Eur J 18(29):9056–9062. https://doi.org/10.1002/chem.201103514
Wohlgemuth J, Karas M, Jiang W et al (2010) Enhanced glyco-profiling by specific glycopeptide enrichment and complementary monolithic nano-LC (ZIC-HILIC/RP18e)/ESI-MS analysis. J Sep Sci 33(6–7):880–890. https://doi.org/10.1002/jssc.200900771
Gaunitz S, Nagy G, Pohl NLB et al (2017) Recent advances in the analysis of complex glycoproteins. Anal Chem 89(1):389–413. https://doi.org/10.1021/acs.analchem.6b04343
Ahn YH, Kim JY, Yoo JS (2015) Quantitative mass spectrometric analysis of glycoproteins combined with enrichment methods. Mass Spectrom Rev 34(2):148–165. https://doi.org/10.1002/mas.21428
Ma J, Hart GW (2014) O-GlcNAc profiling: from proteins to proteomes. Clin Proteomics 11(1):8. https://doi.org/10.1186/1559-0275-11-8
Steentoft C, Vakhrushev SY, Joshi HJ et al (2013) Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J 32(10):1478–1488. https://doi.org/10.1038/emboj.2013.79
Akmal MA, Rasool N, Khan YD (2017) Prediction of N-linked glycosylation sites using position relative features and statistical moments. PLoS One 12(8):e0181966. https://doi.org/10.1371/journal.pone.0181966
Hassan H, Badr A, Abdelhalim MB (2015) Prediction of O-glycosylation sites using random forest and GA-tuned PSO technique. Bioinform Biol Insights 9:103–109. https://doi.org/10.4137/BBI.S26864
Chauhan JS, Rao A, Raghava GPS (2013) In silico platform for prediction of N-, O- and C-glycosites in eukaryotic protein sequences. PLoS One 8(6):e67008. https://doi.org/10.1371/journal.pone.0067008
Hamby SE, Hirst JD (2008) Prediction of glycosylation sites using random forests. BMC Bioinformatics 9(1):500. https://doi.org/10.1186/1471-2105-9-500
Johnson ES (2002) Ubiquitin branches out. Nat Cell Biol 4(12):295. https://doi.org/10.1038/ncb1202-e295
Sun L, Chen ZJ (2004) The novel functions of ubiquitination in signaling. Curr Opin Cell Biol 16(2):119–126. https://doi.org/10.1016/j.ceb.2004.02.005
Akimov V, Rigbolt KTG, Nielsen MM et al (2011) Characterization of ubiquitination dependent dynamics in growth factor receptor signaling by quantitative proteomics. Mol Biosyst 7(12):3223–3233. https://doi.org/10.1039/c1mb05185g
Hjerpe R, Aillet F, Lopitz-Otsoa F et al (2009) Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep 10(11):1250–1258. https://doi.org/10.1038/embor.2009.192
Scott D, Oldham NJ, Strachan J et al (2015) Ubiquitin-binding domains: mechanisms of ubiquitin recognition and use as tools to investigate ubiquitin-modified proteomes. Proteomics 15(5–6):844–861. https://doi.org/10.1002/pmic.201400341
Hicke L, Schubert HL, Hill CP (2005) Ubiquitin-binding domains. Nat Rev Mol Cell Biol 6(8):610–621. https://doi.org/10.1038/nrm1701
Danielsen JMR, Sylvestersen KB, Bekker-Jensen S et al (2011) Mass spectrometric analysis of lysine ubiquitylation reveals promiscuity at site level. Mol Cell Proteomics 10(3):1–12. https://doi.org/10.1074/mcp.M110.003590
Peng J, Schwartz D, Elias JE et al (2003) A proteomics approach to understanding protein ubiquitination. Nat Biotechnol 21(8):921–926. https://doi.org/10.1038/nbt849
Pirone L, Xolalpa W, Sigursson JO et al (2017) A comprehensive platform for the analysis of ubiquitin-like protein modifications using in vivo biotinylation. Sci Rep 7:40756. https://doi.org/10.1038/srep40756
Mattern M, Sutherland J, Kadimisetty K et al (2019) Using ubiquitin binders to decipher the ubiquitin code. Trends Biochem Sci 44(7):599–615. https://doi.org/10.1016/j.tibs.2019.01.011
Kim W, Bennett EJ, Huttlin EL et al (2011) Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol Cell 44(2):325–340. https://doi.org/10.1016/j.molcel.2011.08.025
Wagner SA, Beli P, Weinert BT et al (2012) Proteomic analyses reveal divergent ubiquitylation site patterns in murine tissues. Mol Cell Proteomics 11(12):1578–1585. https://doi.org/10.1074/mcp.M112.017905
Wagner SA, Beli P, Weinert BT et al (2011) A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol Cell Proteomics 10(10):M111.013284. https://doi.org/10.1074/mcp.m111.013284
Udeshi ND, Svinkina T, Mertins P et al (2013) Refined preparation and use of anti-diglycine remnant (k-ε-gg) antibody enables routine quantification of 10,000s of ubiquitination sites in single proteomics experiments. Mol Cell Proteomics 12(3):825–831. https://doi.org/10.1074/mcp.O112.027094
Drazic A, Myklebust LM, Ree R et al (2016) The world of protein acetylation. Biochim Biophys Acta Proteins Proteomics 1864(10):1372–1401. https://doi.org/10.1016/j.bbapap.2016.06.007
Inuzuka H, Gao D, Finley LWS et al (2012) Acetylation-dependent regulation of Skp2 function. Cell 150(1):179–193. https://doi.org/10.1016/j.cell.2012.05.038
Li T, Diner BA, Chen J et al (2012) Acetylation modulates cellular distribution and DNA sensing ability of interferon-inducible protein IFI16. Proc Natl Acad Sci U S A 109(26):10558–10563. https://doi.org/10.1073/pnas.1203447109
Cao W, Bao C, Padalko E et al (2008) Acetylation of mitogen-activated protein kinase phosphatase-1 inhibits Toll-like receptor signaling. J Exp Med 205(6):1491–1503. https://doi.org/10.1084/jem.20071728
Narita T, Weinert BT, Choudhary C (2019) Functions and mechanisms of non-histone protein acetylation. Nat Rev Mol Cell Biol 20(3):156–174. https://doi.org/10.1038/s41580-018-0081-3
Dillon SC, Zhang X, Trievel RC et al (2005) The SET-domain protein superfamily: protein lysine methyltransferases. Genome Biol 6(8):1–10. https://doi.org/10.1186/gb-2005-6-8-227
Dhayalan A, Kudithipudi S, Rathert P et al (2011) Specificity analysis-based identification of new methylation targets of the SET7/9 protein lysine methyltransferase. Chem Biol 18(1):111–120. https://doi.org/10.1016/j.chembiol.2010.11.014
Mazur PK, Reynoird N, Khatri P et al (2014) SMYD3 links lysine methylation of MAP3K2 to Ras-driven cancer. Nature 510(7504):283–287. https://doi.org/10.1038/nature13320
Bedford MT, Clarke SG (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell 33(1):1–13. https://doi.org/10.1016/j.molcel.2008.12.013
Kwak YT, Guo J, Prajapati S et al (2003) Methylation of SPT5 regulates its interaction with RNA polymerase II and transcriptional elongation properties. Mol Cell 11(4):1055–1066. https://doi.org/10.1016/S1097-2765(03)00101-1
Guo A, Gu H, Zhou J et al (2014) Immunoaffinity enrichment and mass spectrometry analysis of protein methylation. Mol Cell Proteomics 13(1):372–387. https://doi.org/10.1074/mcp.O113.027870
Cao XJ, Arnaudo AM, Garcia BA (2013) Large-scale global identification of protein lysine methylation in vivo. Epigenetics 8(5):477–485. https://doi.org/10.4161/epi.24547
Nielsen ML, Savitski MM, Zubarev RA (2006) Extent of modifications in human proteome samples and their effect on dynamic range of analysis in shotgun proteomics. Mol Cell Proteomics 5(12):2384–2391. https://doi.org/10.1074/mcp.M600248-MCP200
Liu T, Qian WJ, Gritsenko MA et al (2005) Human plasma N-glycoproteome analysis by immunoaffinity subtraction, hydrazide chemistry, and mass spectrometry. J Proteome Res 4(6):2070–2080. https://doi.org/10.1021/pr0502065
Gilar M, Olivova P, Daly AE et al (2005) Orthogonality of separation in two-dimensional liquid chromatography. Anal Chem 77(19):6426–6434. https://doi.org/10.1021/ac050923i
Song C, Ye M, Han G et al (2010) Reversed-phase-reversed-phase liquid chromatography approach with high orthogonality for multidimensional separation of phosphopeptides. Anal Chem 82(1):53–56. https://doi.org/10.1021/ac9023044
Yang F, Shen Y, Camp DG et al (2012) High-pH reversed-phase chromatography with fraction concatenation for 2D proteomic analysis. Expert Rev Proteomics 9(2):129–134. https://doi.org/10.1586/epr.12.15
Wang Y, Yang F, Gritsenko MA et al (2011) Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics 11(10):2019–2026. https://doi.org/10.1002/pmic.201000722
Washburn MP, Wolters D, Yates JR (2001) Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat Biotechnol 19(3):242–247. https://doi.org/10.1038/85686
Pankow S, Bamberger C, Yates JR (2019) A posttranslational modification code for CFTR maturation is altered in cystic fibrosis. Sci Signal 12(562):eaan7984. https://doi.org/10.1126/scisignal.aan7984
Delmotte N, Lasaosa M, Tholey A et al (2007) Two-dimensional reversed-phase x ion-pair reversed-phase HPLC: an alternative approach to high-resolution peptide separation for shotgun proteome analysis. J Proteome Res 6(11):4363–4373. https://doi.org/10.1021/pr070424t
Di Palma S, Boersema PJ, Heck AJR et al (2011) Zwitterionic hydrophilic interaction liquid chromatography (ZIC-HILIC and ZIC-cHILIC) provide high resolution separation and increase sensitivity in proteome analysis. Anal Chem 83(9):3440–3447. https://doi.org/10.1021/ac103312e
Boersema PJ, Divecha N, Heck AJR et al (2007) Evaluation and optimization of ZIC-HILIC-RP as an alternative MudPIT strategy. J Proteome Res 6(3):937–946. https://doi.org/10.1021/pr060589m
Dehghani A, Gödderz M, Winter D (2018) Tip-based fractionation of batch-enriched phosphopeptides facilitates easy and robust phosphoproteome analysis. J Proteome Res 17(1):46–54. https://doi.org/10.1021/acs.jproteome.7b00256
Humphrey SJ, Karayel O, James DE et al (2018) High-throughput and high-sensitivity phosphoproteomics with the EasyPhos platform. Nat Protoc 13(9):1897–1916. https://doi.org/10.1038/s41596-018-0014-9
Scheltema RA, Hauschild JP, Lange O et al (2014) The Q exactive HF, a benchtop mass spectrometer with a pre-filter, high-performance quadrupole and an ultra-high-field orbitrap analyzer. Mol Cell Proteomics 13(12):3698–3708. https://doi.org/10.1074/mcp.M114.043489
Andrews GL, Simons BL, Young JB et al (2011) Performance characteristics of a new hybrid quadrupole time-of-flight tandem mass spectrometer (TripleTOF 5600). Anal Chem 83(13):5442–5446. https://doi.org/10.1021/ac200812d
Scigelova M, Makarov A (2006) Orbitrap mass analyzer—overview and applications in proteomics. Proteomics 1(1–2):16–21. https://doi.org/10.1002/pmic.200600528
Michalski A, Damoc E, Hauschild JP et al (2011) Mass spectrometry-based proteomics using Q exactive, a high-performance benchtop quadrupole orbitrap mass spectrometer. Mol Cell Proteomics 10(9):M111.011015. https://doi.org/10.1074/mcp.M111.011015
Michalski A, Damoc E, Lange O et al (2012) Ultra high resolution linear ion trap orbitrap mass spectrometer (orbitrap elite) facilitates top down LC MS/MS and versatile peptide fragmentation modes. Mol Cell Proteomics 11(3):1–11. https://doi.org/10.1074/mcp.O111.013698
Batth TS, Francavilla C, Olsen JV (2014) Off-line high-pH reversed-phase fractionation for in-depth phosphoproteomics. J Proteome Res 13(12):6176–6186. https://doi.org/10.1021/pr500893m
Kelstrup CD, Bekker-Jensen DB, Arrey TN et al (2018) Performance evaluation of the Q exactive HF-X for shotgun proteomics. J Proteome Res 17(1):727–738. https://doi.org/10.1021/acs.jproteome.7b00602
Boersema PJ, Mohammed S, Heck AJR (2009) Phosphopeptide fragmentation and analysis by mass spectrometry. J Mass Spectrom 44(6):861–878. https://doi.org/10.1002/jms.1599
Engholm-Keller K, Larsen MR (2013) Technologies and challenges in large-scale phosphoproteomics. Proteomics 13(6):910–931. https://doi.org/10.1002/pmic.201200484
Villén J, Beausoleil SA, Gygi SP (2008) Evaluation of the utility of neutral-loss-dependent MS3 strategies in large-scale phosphorylation analysis. Proteomics 8(21):4444–4452. https://doi.org/10.1002/pmic.200800283
Riley NM, Malaker SA, Driessen MD et al (2020) Optimal dissociation methods differ for N- and O-glycopeptides. J Proteome Res 19(8):3286–3301. https://doi.org/10.1021/acs.jproteome.0c00218
Syka JEP, Coon JJ, Schroeder MJ et al (2004) Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc Natl Acad Sci U S A 101(26):9528–9533. https://doi.org/10.1073/pnas.0402700101
Molina H, Horn DM, Tang N et al (2007) Global proteomic profiling of phosphopeptides using electron transfer dissociation tandem mass spectrometry. Proc Natl Acad Sci U S A 104(7):2199–2204. https://doi.org/10.1073/pnas.0611217104
Good DM, Wirtala M, McAlister GC et al (2007) Performance characteristics of electron transfer dissociation mass spectrometry. Mol Cell Proteomics 6(11):1942–1951. https://doi.org/10.1074/mcp.M700073-MCP200
Rose CM, Rush MJP, Riley NM et al (2015) A calibration routine for efficient ETD in large-scale proteomics. J Am Soc Mass Spectrom 26(11):1848–1857. https://doi.org/10.1007/s13361-015-1183-1
Frese CK, Altelaar AFM, Van Den Toorn H et al (2012) Toward full peptide sequence coverage by dual fragmentation combining electron-transfer and higher-energy collision dissociation tandem mass spectrometry. Anal Chem 84(22):9668–9673. https://doi.org/10.1021/ac3025366
Kong AT, Leprevost FV, Avtonomov DM et al (2017) MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry-based proteomics. Nat Methods 14(5):513–520. https://doi.org/10.1038/nmeth.4256
Dorfer V, Pichler P, Stranzl T et al (2014) MS Amanda, a universal identification algorithm optimized for high accuracy tandem mass spectra. J Proteome Res 13(8):3679–3684. https://doi.org/10.1021/pr500202e
Craig R, Beavis RC (2004) TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20(9):1466–1467. https://doi.org/10.1093/bioinformatics/bth092
Kim S, Pevzner PA (2014) MS-GF+ makes progress towards a universal database search tool for proteomics. Nat Commun 5:5277. https://doi.org/10.1038/ncomms6277
Solntsev SK, Shortreed MR, Frey BL et al (2018) Enhanced global post-translational modification discovery with MetaMorpheus. J Proteome Res 17(5):1844–1851. https://doi.org/10.1021/acs.jproteome.7b00873
Eng JK, Jahan TA, Hoopmann MR (2013) Comet: an open-source MS/MS sequence database search tool. Proteomics 13(1):22–24. https://doi.org/10.1002/pmic.201200439
Beausoleil SA, Villén J, Gerber SA et al (2006) A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat Biotechnol 24(10):1285–1292. https://doi.org/10.1038/nbt1240
Bailey CM, Sweet SMM, Cunningham DL et al (2009) SLoMo: automated site localization of modifications from ETD/ECD mass spectra. J Proteome Res 8(4):1965–1971. https://doi.org/10.1021/pr800917p
Fermin D, Avtonomov D, Choi H et al (2015) LuciPHOr2: site localization of generic post-translational modifications from tandem mass spectrometry data. Bioinformatics 31(7):1141–1143. https://doi.org/10.1093/bioinformatics/btu788
Savitski MM, Lemeer S, Boesche M et al (2011) Confident phosphorylation site localization using the mascot delta score. Mol Cell Proteomics 10(2):S1–S12. https://doi.org/10.1074/mcp.M110.003830
Taus T, Köcher T, Pichler P et al (2011) Universal and confident phosphorylation site localization using phosphoRS. J Proteome Res 10(12):5354–5362. https://doi.org/10.1021/pr200611n
Elias JE, Gygi SP (2007) Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat Methods 4(3):207–214. https://doi.org/10.1038/nmeth1019
Elias JE, Gygi SP (2010) Target-decoy search strategy for mass spectrometry-based proteomics. Methods Mol Biol 60:455–471. https://doi.org/10.1007/978-1-60,761-444-9_5
Olsen JV, Vermeulen M, Santamaria A et al (2010) Quantitative phosphoproteomics reveals widespread full phosphorylation site occupancy during mitosis. Sci Signal 3(104):ra3. https://doi.org/10.1126/scisignal.2000475
Udeshi ND, Mani DR, Eisenhaure T et al (2012) Methods for quantification of in vivo changes in protein ubiquitination following proteasome and deubiquitinase inhibition. Mol Cell Proteomics 11(5):148–159. https://doi.org/10.1074/mcp.M111.016857
Dupree EJ, Jayathirtha M, Yorkey H et al (2020) A critical review of bottom-up proteomics: the good, the bad, and the future of this field. Proteomes 8(3):1–26. https://doi.org/10.3390/proteomes8030014
Chick JM, Kolippakkam D, Nusinow DP et al (2015) A mass-tolerant database search identifies a large proportion of unassigned spectra in shotgun proteomics as modified peptides. Nat Biotechnol 33(7):743–749. https://doi.org/10.1038/nbt.3267
Liu K, Li S, Wang L et al (2020) Full-spectrum prediction of peptides tandem mass spectra using deep neural network. Anal Chem 92(6):4275–4283. https://doi.org/10.1021/acs.analchem.9b04867
Olsen JV, Blagoev B, Gnad F et al (2006) Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127(3):635–648. https://doi.org/10.1016/j.cell.2006.09.026
Locard-Paulet M, Bouyssié D, Froment C et al (2020) Comparing 22 popular phosphoproteomics pipelines for peptide identification and site localization. J Proteome Res 19(3):1338–1345. https://doi.org/10.1021/acs.jproteome.9b00679
Chen Z, Chen YZ, Wang XF et al (2011) Prediction of ubiquitination sites by using the composition of K-Spaced amino acid pairs. PLoS One 6(7):e22930. https://doi.org/10.1371/journal.pone.0022930
Swaney DL, Beltrao P, Starita L et al (2013) Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation. Nat Methods 10(7):676–682. https://doi.org/10.1038/nmeth.2519
Sidoli S, Garcia BA (2017) Middle-down proteomics: a still unexploited resource for chromatin biology. Expert Rev Proteomics 14(7):617–626. https://doi.org/10.1080/14789450.2017.1345632
Zhou M, Paša-Tolić L, Stenoien DL (2017) Profiling of histone post-translational modifications in mouse brain with high-resolution top-down mass spectrometry. J Proteome Res 16(2):599–608. https://doi.org/10.1021/acs.jproteome.6b00694
Leney AC, El Atmioui D, Wu W et al (2017) Elucidating crosstalk mechanisms between phosphorylation and O-GlcNAcylation. Proc Natl Acad Sci U S A 114(35):E7255–E7261. https://doi.org/10.1073/pnas.1620529114
Hasan MM, Khatun MS, Mollah MNH et al (2017) A systematic identification of species-specific protein succinylation sites using joint element features information. Int J Nanomedicine 126:303–6315. https://doi.org/10.2147/IJN.S140875
Liu Z, Dong W, Jiang W et al (2019) csDMA: an improved bioinformatics tool for identifying DNA 6 mA modifications via Chou’s 5-step rule. Sci Rep 9(1):1–9. https://doi.org/10.1038/s41598-019-49,430-4
Roitinger E, Hofer M, Köcher T et al (2015) Quantitative phosphoproteomics of the ataxia telangiectasia-mutated (ATM) and ataxia telangiectasia-mutated and Rad3-related (ATR) dependent DNA damage response in Arabidopsis thaliana. Mol Cell Proteomics 14(3):556–571. https://doi.org/10.1074/mcp.M114.040352
McConnell EW, Berg P, Westlake TJ et al (2019) Proteome-wide analysis of cysteine reactivity during effector-triggered immunity. Plant Physiol 179(4):1248–1264. https://doi.org/10.1104/pp.18.01194
Huang J, Willems P, Wei B et al (2019) Mining for protein S-sulfenylation in Arabidopsis uncovers redox-sensitive sites. Proc Natl Acad Sci U S A 116(42):20256–20261. https://doi.org/10.1073/pnas.1906768116
Hu J, Huang X, Chen L et al (2015) Site-specific nitrosoproteomic identification of endogenously S-nitrosylated proteins in Arabidopsis. Plant Physiol 167(4):1731–1746. https://doi.org/10.1104/pp.15.00026
Zielinska DF, Gnad F, Schropp K et al (2012) Mapping N-glycosylation sites across seven evolutionarily distant species reveals a divergent substrate proteome despite a common core machinery. Mol Cell 46(4):542–548. https://doi.org/10.1016/j.molcel.2012.04.031
Walton A, Stes E, Cybulski N et al (2016) It’s time for some “site”-seeing: novel tools to monitor the ubiquitin landscape in Arabidopsis thaliana. Plant Cell 28(1):6–16. https://doi.org/10.1105/tpc.15.00878
Hartl M, Füßl M, Boersema PJ et al (2017) Lysine acetylome profiling uncovers novel histone deacetylase substrate proteins in Arabidopsis. Mol Syst Biol 13(10):949. https://doi.org/10.15252/msb.20177819
Liang Q, Geng Q, Jiang L et al (2020) Protein methylome analysis in Arabidopsis reveals regulation in RNA-related processes. J Proteomics 213:103601. https://doi.org/10.1016/j.jprot.2019.103601
Huang J, Wu Z, Zhang X (2020) Short-term mild temperature-stress-induced alterations in the C. elegans phosphoproteome. Int J Mol Sci 21(17):6409. https://doi.org/10.3390/ijms21176409
Wang H, Gau B, Slade WO et al (2014) The global phosphoproteome of chlamydomonas reinhardtii reveals complex organellar phosphorylation in the flagella and thylakoid membrane. Mol Cell Proteomics 13(9):2337–2353. https://doi.org/10.1074/mcp.M114.038281
Morisse S, Zaffagnini M, Gao XH et al (2014) Insight into protein S-nitrosylation in Chlamydomonas reinhardtii. Antioxid Redox Signal 21(9):1271–1284. https://doi.org/10.1089/ars.2013.5632
Schulze S, Oltmanns A, Machnik N et al (2018) N-glycoproteomic characterization of mannosidase and xylosyltransferase mutant strains of Chlamydomonas reinhardtii. Plant Physiol 176(3):1952–1964. https://doi.org/10.1104/pp.17.01450
Yan J, Long Y, Zhou T et al (2020) Dynamic phosphoproteome profiling of zebrafish embryonic fibroblasts during cold acclimation. Proteomics 20(2):1900257. https://doi.org/10.1002/pmic.201900257
Kwon OK, Kim S, Lee S (2016) Global proteomic analysis of lysine acetylation in zebrafish (Danio rerio) embryos. Electrophoresis 37(23–24):3137–3145. https://doi.org/10.1002/elps.201600210
Bo Zhai JV, Beausoleil SA, Mintseris J et al (2008) Phosphoproteome analysis of drosophila melanogaster embryos. J Proteome Res 7(4):1675–1682. https://doi.org/10.1021/pr700696a
Menger KE, James AM, Cochemé HM et al (2015) Fasting, but not aging, dramatically alters the redox status of cysteine residues on proteins in Drosophila melanogaster. Cell Rep 11(12):1856–1865. https://doi.org/10.1016/j.celrep.2015.05.033
Sap KA, Bezstarosti K, Dekkers DHW et al (2017) Quantitative proteomics reveals extensive changes in the ubiquitinome after perturbation of the proteasome by targeted dsRNA-mediated subunit knockdown in drosophila. J Proteome Res 16(8):2848–2862. https://doi.org/10.1021/acs.jproteome.7b00156
Weinert BT, Wagner SA, Horn H et al (2011) Proteome-wide mapping of the Drosophila acetylome demonstrates a high degree of conservation of lysine acetylation. Sci Signal 4(183):ra48. https://doi.org/10.1126/scisignal.2001902
Semanjski M, Germain E, Bratl K et al (2018) The kinases HipA and HipA7 phosphorylate different substrate pools in Escherichia coli to promote multidrug tolerance. Sci Signal 11(547):5750. https://doi.org/10.1126/scisignal.aat5750
Shakir S, Vinh J, Chiappetta G (2017) Quantitative analysis of the cysteine redoxome by iodoacetyl tandem mass tags. Anal Bioanal Chem 409(15):3821–3830. https://doi.org/10.1007/s00216-017-0326-6
Weinert BT, Iesmantavicius V, Wagner SA et al (2013) Acetyl-phosphate is a critical determinant of lysine acetylation in E. coli. Mol Cell 51(2):265–272. https://doi.org/10.1016/j.molcel.2013.06.003
Zhang M, Xu J-Y, Hu H et al (2018) Systematic proteomic analysis of protein methylation in prokaryotes and eukaryotes revealed distinct substrate specificity. Proteomics 18(1):1700300. https://doi.org/10.1002/pmic.201700300
Sharma K, D’Souza RCJ, Tyanova S et al (2014) Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep 8(5):1583–1594. https://doi.org/10.1016/j.celrep.2014.07.036
Huang H, Petersen MH, Ibañez-Vea M et al (2016) Simultaneous enrichment of cysteine containing peptides and phosphopeptides using a cysteine-specific phosphonate adaptable tag (CysPAT) in combination with titanium dioxide (TiO2) chromatography. Mol Cell Proteomics 15(10):3282–3296. https://doi.org/10.1074/mcp.M115.054551
Mnatsakanyan R, Markoutsa S, Walbrunn K et al (2019) Proteome-wide detection of S-nitrosylation targets and motifs using bioorthogonal cleavable-linker-based enrichment and switch technique. Nat Commun 10(1):1–12. https://doi.org/10.1038/s41467-019-10,182-4
Zhu J, Sun Z, Cheng K et al (2014) Comprehensive mapping of protein N-glycosylation in human liver by combining hydrophilic interaction chromatography and hydrazide chemistry. J Proteome Res 13(3):1713–1721. https://doi.org/10.1021/pr401200h
Akimov V, Barrio-Hernandez I, Hansen SVF et al (2018) Ubisite approach for comprehensive mapping of lysine and n-terminal ubiquitination sites. Nat Struct Mol Biol 25(7):631–640. https://doi.org/10.1038/s41594-018-0084-y
Schölz C, Weinert BT, Wagner SA et al (2015) Acetylation site specificities of lysine deacetylase inhibitors in human cells. Nat Biotechnol 33(4):415–425. https://doi.org/10.1038/nbt.3130
Larsen SC, Sylvestersen KB, Mund A et al (2016) Proteome-wide analysis of arginine monomethylation reveals widespread occurrence in human cells. Sci Signal 9(443):rs9. https://doi.org/10.1126/scisignal.aaf7329
Wang Z, Ma J, Miyoshi C et al (2018) Quantitative phosphoproteomic analysis of the molecular substrates of sleep need. Nature 558(7710):435–439. https://doi.org/10.1038/s41586-018-0218-8
Wang J, Choi H, Chung NC et al (2018) Integrated dissection of cysteine oxidative post-translational modification proteome during cardiac hypertrophy. J Proteome Res 17(12):4243–4257. https://doi.org/10.1021/acs.jproteome.8b00372
Zareba-Koziol M, Bartkowiak-Kaczmarek A, Figiel I et al (2019) Stress-induced changes in the S-palmitoylation and S-nitrosylation of synaptic proteins. Mol Cell Proteomics 18(10):1916–1938. https://doi.org/10.1074/mcp.RA119.001581
Fang P, Wang XJ, Xue Y et al (2016) In-depth mapping of the mouse brain N-glycoproteome reveals widespread N-glycosylation of diverse brain proteins. Oncotarget 7(25):38796–38809. https://doi.org/10.18632/oncotarget.9737
Rardin MJ, Newman JC, Held JM et al (2013) Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc Natl Acad Sci U S A 110(16):6601–6606. https://doi.org/10.1073/pnas.1302961110
Li J, Paulo JA, Nusinow DP et al (2019) Investigation of proteomic and phosphoproteomic responses to signaling network perturbations reveals functional pathway organizations in yeast. Cell Rep 29(7):2092–2104.e4. https://doi.org/10.1016/j.celrep.2019.10.034
Neubert P, Halim A, Zauser M et al (2016) Mapping the O-mannose glycoproteome in saccharomyces cerevisiae. Mol Cell Proteomics 15(4):1323–1337. https://doi.org/10.1074/mcp.M115.057505
Iesmantavicius V, Weinert BT, Choudhary C (2014) Convergence of ubiquitylation and phosphorylation signaling in rapamycin-treated yeast cells. Mol Cell Proteomics 13(8):1979–1992. https://doi.org/10.1074/mcp.O113.035683
Henriksen P, Wagner SA, Weinert BT et al (2012) Proteome-wide analysis of lysine acetylation suggests its broad regulatory scope in saccharomyces cerevisiae. Mol Cell Proteomics 11(11):1510–1522. https://doi.org/10.1074/mcp.M112.017251
Zhang Y, Pan Y, Liu W et al (2016) In vivo protein allylation to capture protein methylation candidates. Chem Commun 52(40):6689–6692. https://doi.org/10.1039/c6cc02386j
Perez-Riverol Y, Csordas A, Bai J et al (2019) The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47(D1):D442–D450. https://doi.org/10.1093/nar/gky1106
Zhang Z, Tan M, Xie Z et al (2011) Identification of lysine succinylation as a new post-translational modification. Nat Chem Biol 7(1):58–63. https://doi.org/10.1038/nchembio.495
Newton K, Matsumoto ML, Wertz IE et al (2008) Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell 134(4):668–678. https://doi.org/10.1016/j.cell.2008.07.039
Ghosh R, Gilda JE, Gomes AV (2014) The necessity of and strategies for improving confidence in the accuracy of western blots. Expert Rev Proteomics 11(5):549–560. https://doi.org/10.1586/14789450.2014.939635
Pillai-Kastoori L, Schutz-Geschwender AR, Harford JA (2020) A systematic approach to quantitative Western blot analysis. Anal Biochem 593(15):113608. https://doi.org/10.1016/j.ab.2020.113608
Yu Y, Anjum R, Kubota K et al (2009) A site-specific, multiplexed kinase activity assay using stable-isotope dilution and high-resolution mass spectrometry. Proc Natl Acad Sci U S A 106(28):11606–11611. https://doi.org/10.1073/pnas.0905165106
Gillette MA, Carr SA (2013) Quantitative analysis of peptides and proteins in biomedicine by targeted mass spectrometry. Nat Methods 10(1):28–34. https://doi.org/10.1038/NMETH.2309
Morris M, Knudsen GM, Maeda S et al (2015) Tau post-translational modifications in wild-type and human amyloid precursor protein transgenic mice. Nat Neurosci 18(8):1183–1189. https://doi.org/10.1038/nn.4067
Shi T, Song E, Nie S et al (2016) Advances in targeted proteomics and applications to biomedical research. Proteomics 16(15–16):2160–2182. https://doi.org/10.1002/pmic.201500449
Ebhardt HA, Root A, Sander C et al (2015) Applications of targeted proteomics in systems biology and translational medicine. Proteomics 15(18):3193–3208. https://doi.org/10.1002/pmic.201500004
Zhao Y, Brasier AR (2013) Applications of selected reaction monitoring (SRM)-mass spectrometry (MS) for quantitative measurement of signaling pathways. Methods 61(3):313–322. https://doi.org/10.1016/j.ymeth.2013.02.001
Gallien S, Duriez E, Crone C et al (2012) Targeted proteomic quantification on quadrupole-orbitrap mass spectrometer. Mol Cell Proteomics 11(12):1709–1723. https://doi.org/10.1074/mcp.O112.019802
Ronsein GE, Pamir N, von Haller PD et al (2015) Parallel reaction monitoring (PRM) and selected reaction monitoring (SRM) exhibit comparable linearity, dynamic range and precision for targeted quantitative HDL proteomics. J Proteomics 113:388–399. https://doi.org/10.1016/j.jprot.2014.10.017
Rauniyar N (2015) Parallel reaction monitoring: a targeted experiment performed using high resolution and high mass accuracy mass spectrometry. Int J Mol Sci 16(12):28566–28581. https://doi.org/10.3390/ijms161226120
Schilling B, MacLean B, Held JM et al (2015) Multiplexed, scheduled, high-resolution parallel reaction monitoring on a full scan QqTOF instrument with integrated data-dependent and targeted mass spectrometric workflows. Anal Chem 87(20):10222–10229. https://doi.org/10.1021/acs.analchem.5b02983
Peterson AC, Russell JD, Bailey DJ et al (2012) Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol Cell Proteomics 11(11):1475–1488. https://doi.org/10.1074/mcp.O112.020131
Xu G, Jaffrey SR (2013) Proteomic identification of protein ubiquitination events. Biotechnol Genet Eng Rev 29(1):73–109. https://doi.org/10.1080/02648725.2013.801232
Blom N, Sicheritz-Pontén T, Gupta R et al (2004) Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4(6):1633–1649. https://doi.org/10.1002/pmic.200300771
Unwin RD, Griffiths JR, Whetton AD (2010) Simultaneous analysis of relative protein expression levels across multiple samples using iTRAQ isobaric tags with 2D nano LC-MS/MS. Nat Protoc 51:1574–1582. https://doi.org/10.1038/nprot.2010.123
Thompson A, Wölmer N, Koncarevic S et al (2019) TMTpro: design, synthesis, and initial evaluation of a proline-based isobaric 16-Plex Tandem Mass Tag Reagent Set. Anal Chem 91(24):15941–15950. https://doi.org/10.1021/acs.analchem.9b04474
Acknowledgments
This work was supported by a National Science Foundation CAREER award (MCB-1552522) awarded to L.M.H. We thank Prof. Dukka KC, Amanda Smythers, and Patric Sadecki for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Iannetta, A.A., Hicks, L.M. (2022). Maximizing Depth of PTM Coverage: Generating Robust MS Datasets for Computational Prediction Modeling. In: KC, D.B. (eds) Computational Methods for Predicting Post-Translational Modification Sites. Methods in Molecular Biology, vol 2499. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2317-6_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2317-6_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2316-9
Online ISBN: 978-1-0716-2317-6
eBook Packages: Springer Protocols