Abstract
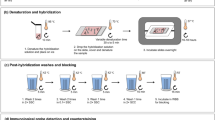
The opportunity to map genes and noncoding DNA sequences on the chromosomes by fluorescence in situ hybridization (FISH) has greatly enhanced the potential for fish karyotyping and comparative cytogenetics. The use of FISH allowed for significant advances in our understanding of the fish genome architecture, especially when applied to the study of the repetitive component of the genome, that is generally underestimated in the bioinformatic assembly. Here we provide a step-by-step protocol for FISH of repeated sequences onto chromosomes of fish species.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pardue ML, Gall JG (1969) Molecular hybridization of radioactive DNA to the DNA of cytological preparations. Proc Natl Acad Sci U S A 64:600–604. https://doi.org/10.1073/pnas.64.2.600
Phillips RB (2007) Application of fluorescence in situ hybridization (FISH) to genome mapping in fishes. In: Pisano E, Ozouf-Costaz C, Foresti F, Kapoor BG (eds) Fish cytogenetics. Science Publishers, Enfield, pp 455–471
Phillips RB, Keatley KA, Morasch MR et al (2009) Assignment of Atlantic salmon (Salmo salar) linkage groups to specific chromosomes: conservation of large syntenic blocks corresponding to whole chromosome arms in rainbow trout (Oncorhynchus mykiss). BMC Genet 10(1):1–11. https://doi.org/10.1186/1471-2156-10-46
Zhang P, Priebe B (2009) FISH on plant chromosomes. In: Fluorescence in situ hybridization (FISH)—application guide. Springer, Berlin, Heidelberg, pp 365–394. https://doi.org/10.1007/978-3-540-70581-9_32
Solinhac R, Leroux S, Galkina S et al (2010) Integrative mapping analysis of chicken microchromosome 16 organization. BMC Genomics 11(1):1–12. https://doi.org/10.1186/1471-2164-11-616
Abbasi FM, Khan MT, Perveen F et al (2010) Historical perspective of in situ hybridization for the analysis of genomic constitution of plants. Afr J Biotechnol 9(54):9142–9147
Jiang J (2019) Fluorescence in situ hybridization in plants: recent developments and future applications. Chrom Res 27(3):153–165. https://doi.org/10.1007/s10577-019-09607-z
Fortes GG, Bouza C, Viñas A et al (2007) Diversity in isochore structure and chromosome banding in fish. In: Pisano E, Ozouf-Costaz C, Foresti F, Kapoor BG (eds) Fish cytogenetics. Science Publishers, Enfield, pp 405–420
Pisano E, Ozouf-Costaz C, Foresti F, Kapoor BG (eds) (2007) Fish cytogenetics. Science Publishers, Enfield
Mazzuchelli J, Martins C (2009) Genomic organization of repetitive DNAs in the cichlid fish Astronotus ocellatus. Genetica 136(3):461–469. https://doi.org/10.1007/s10709-008-9346-7
Reichwald K, Lauber C, Nanda I et al (2009) High tandem repeat content in the genome of the short-lived annual fish Nothobranchius furzeri: a new vertebrate model for aging research. Genome Biol 10(2):1–17. https://doi.org/10.1186/gb-2009-10-2-r16
Majtánová Z, Moy KG, Unmack PJ et al (2019) Characterization of the karyotype and accumulation of repetitive sequences in Australian Darling hardyhead Craterocephalus amniculus (Atheriniformes, Teleostei). PeerJ 7:e7347. https://doi.org/10.7717/peerj.7347
Biscotti MA, Olmo E, Heslop-Harrison JS (2015) Repetitive DNA in eukaryotic genomes. Chromosom Res 23:415–420. https://doi.org/10.1007/s10577-015-9499-z
Crollius HR, Jaillon O, Dasilva C et al (2000) Characterization and repeat analysis of the compact genome of the freshwater pufferfish Tetraodon nigroviridis. Genome Res 10(7):939–949. https://doi.org/10.1101/gr.10.7.939
Freeman JL, Adeniyi A, Banerjee R et al (2007) Definition of the zebrafish genome using flow cytometry and cytogenetic mapping. BMC Genomics 8(1):1–10. https://doi.org/10.1186/1471-2164-8-195
Nicodemus-Johnson J, Silic S, Ghigliotti L et al (2011) Assembly of the antifreeze glycoprotein/trypsinogen-like protease genomic locus in the Antarctic toothfish Dissostichus mawsoni (Norman). Genomics 98(3):194–201. https://doi.org/10.1016/j.ygeno.2011.06.002
Zhuang X, Murphy KR, Ghigliotti L et al (2018) Reconstruction of the repetitive antifreeze glycoprotein genomic loci in the cold-water gadids Boreogadus saida and Microgadus tomcod. Mar Genomics 39:73–84. https://doi.org/10.1016/j.margen.2018.02.003
Pisano E, Ghigliotti L (2009) Ribosomal genes in notothenioid fishes: focus on the chromosomal organization. Mar Genomics 2:75–80. https://doi.org/10.1016/j.margen.2009.03.006
Gornung E (2013) Twenty years of physical mapping of major ribosomal RNA genes across the teleosts: a review of research. Cytogenet Genome Res 141(2–3):90–102. https://doi.org/10.1159/000354832
Rebordinos L, Cross I, Merlo A (2013) High evolutionary dynamism in 5S rDNA of fish: state of the art. Cytogenet Genome Res 141(2–3):103–113. https://doi.org/10.1159/000354871
Sochorová J, Garcia S, Gálvez F et al (2018) Evolutionary trends in animal ribosomal DNA loci: introduction to a new online database. Chromosoma 127(1):141–150. https://doi.org/10.1007/s00412-017-0651-8
Ocalewicz K (2012) Genomic distribution of telomeric DNA sequences—what do we learn from fish about telomere evolution. In: Li B (ed) Reviews on selected topics of telomere biology. InTech, Rijeka, pp 271–294
Auvinet J, Graça P, Ghigliotti L, Pisano E et al (2019) Insertion hot spots of DIRS1 retrotransposon and chromosomal diversifications among the Antarctic Teleosts Nototheniidae. Int J Mol Sci 20:701. https://doi.org/10.3390/ijms20030701
Schemberger MO, Nascimento VD, Coan R et al (2019) DNA transposon invasion and microsatellite accumulation guide W chromosome differentiation in a Neotropical fish genome. Chromosoma 128(4):547–560. https://doi.org/10.1007/s00412-019-00721-9
Ghigliotti L, Mazzei F, Ozouf-Costaz C et al (2007) The two giant sister species of the Southern Ocean, Dissostichus eleginoides and Dissostichus mawsoni, differ in karyotype and chromosomal pattern of ribosomal RNA genes. Polar Biol 30:625–634. https://doi.org/10.1007/s00300-006-0222-6
Ijdo JW, Wells RA, Baldini A et al (1991) Improved telomere detection using a telomere repeat probe (TTAGGG) n generated by PCR. Nucleic Acids Res 19(17):4780
Ghigliotti L, Cheng CHC, Ozouf-Costaz C et al (2020) Cytogenetic characterization of the Antarctic silverfish Pleuragramma antarctica (Boulenger 1902) through analysis of mitotic chromosomes from early larvae. Mar Genomics 52:100737
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ghigliotti, L., Auvinet, J., Pisano, E. (2022). Physical Mapping of Repeated Sequences on Fish Chromosomes by Fluorescence In Situ Hybridization (FISH). In: Verde, C., Giordano, D. (eds) Marine Genomics. Methods in Molecular Biology, vol 2498. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2313-8_21
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2313-8_21
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2312-1
Online ISBN: 978-1-0716-2313-8
eBook Packages: Springer Protocols