Abstract
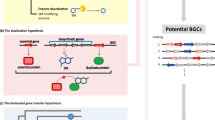
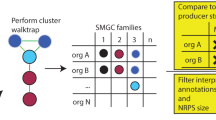
Predicting secondary metabolite biosynthetic gene clusters is a routine analysis performed for each newly sequenced fungal genome. Yet, the usefulness of such predictions remains restricted as they provide total numbers of biosynthetic pathways with only very limited biological significance. In this chapter, we describe a workflow to predict and analyze biosynthetic gene clusters in fungal genomes. It relies on similarity networking and phylogeny to perform genetic dereplication and to prioritize candidate gene clusters that potentially produce new compounds. This basic workflow includes the generation of high-quality figures for publication.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hyde KD, Xu J, Rapior S et al (2019) The amazing potential of fungi: 50 ways we can exploit fungi industrially. Fungal Divers 97:1–136
Mosunova O, Navarro-Muñoz JC, Collemare J (2020) The biosynthesis of fungal secondary metabolites: from fundamentals to biotechnological applications. In: Reference module in life sciences. Elsevier, Amsterdam
Keller NP, Hohn TM (1997) Metabolic pathway gene clusters in filamentous fungi. Fungal Genet Biol 21:17–29
Greco C, Keller NP, Rokas A (2019) Unearthing fungal chemodiversity and prospects for drug discovery. Curr Opin Microbiol 51:22–29
Medema MH, Blin K, Cimermancic P et al (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346
Blin K, Shaw S, Steinke K et al (2019) AntiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Khaldi N, Seifuddin FT, Turner G et al (2010) SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genet Biol 47:736–741
Wolf T, Shelest V, Nath N et al (2016) CASSIS and SMIPS: promoter-based prediction of secondary metabolite gene clusters in eukaryotic genomes. Bioinformatics 32:1138–1143
Umemura M, Koike H, Nagano N et al (2013) MIDDAS-M: motif-independent de novo detection of secondary metabolite gene clusters through the integration of genome sequencing and transcriptome data. PLoS One 8:e84028
Vesth TC, Brandl J, Andersen MR (2016) FunGeneClusterS: predicting fungal gene clusters from genome and transcriptome data. Synth Syst Biotechnol 1:122–129
Takeda I, Umemura M, Koike H et al (2014) Motif-independent prediction of a secondary metabolism gene cluster using comparative genomics: application to sequenced genomes of Aspergillus and ten other filamentous fungal species. DNA Res 21:447–457
Almeida H, Palys S, Tsang A et al (2020) TOUCAN: a framework for fungal biosynthetic gene cluster discovery. NAR Genom Bioinform 2:1–11
Blin K, Shaw S, Kautsar SA et al (2021) The antiSMASH database version 3: increased taxonomic coverage and new query features for modular enzymes. Nucleic Acids Res 49:D639–D643
Kautsar SA, Blin K, Shaw S et al (2019) MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res 48:D454–D458
Kautsar SA, Blin K, Shaw S et al (2021) BiG-FAM: the biosynthetic gene cluster families database. Nucleic Acids Res 49:D490–D497
Weber T, Blin K, Duddela S et al (2015) antiSMASH 3.0--a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:1–7
Adamek M, Alanjary M, Ziemert N (2019) Applied evolution: phylogeny-based approaches in natural products research. Nat Prod Rep 36:1295–1312
Navarro-Muñoz JC, Selem-Mojica N, Mullowney MW et al (2020) A computational framework to explore large-scale biosynthetic diversity. Nat Chem Biol 16:60–68
Gilchrist CLM, Chooi Y-H (2021) Clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics btab007
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973
Minh BQ, Schmidt HA, Chernomor O et al (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534
Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30:3276–3278
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296
Grigoriev IV, Nikitin R, Haridas S et al (2014) MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res 42:D699–D704
Kroken S, Glass NL, Taylor JW et al (2003) Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc Natl Acad Sci U S A 100:15670–15675
Gallo A, Ferrara M, Perrone G (2013) Phylogenetic study of polyketide synthases and nonribosomal peptide synthetases involved in the biosynthesis of mycotoxins. Toxins (Basel) 5:717–742
Bushley KE, Turgeon BG (2010) Phylogenomics reveals subfamilies of fungal nonribosomal peptide synthetases and their evolutionary relationships. BMC Evol Biol 10:26
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Sievers F, Higgins DG (2018) Clustal omega for making accurate alignments of many protein sequences. Protein Sci 27:135–145
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Steenwyk JL, Buida TJ, Li Y et al (2020) ClipKIT: a multiple sequence alignment trimming software for accurate phylogenomic inference. PLoS Biol 18:e3001007
Kalyaanamoorthy S, Minh BQ, Wong TKF et al (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195
Guindon S, Dufayard J-F, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Huerta-Cepas J, Serra F, Bork P (2016) ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol Biol Evol 33:1635–1638
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Navarro-Muñoz, J.C., Collemare, J. (2022). A Bioinformatics Workflow for Investigating Fungal Biosynthetic Gene Clusters. In: Skellam, E. (eds) Engineering Natural Product Biosynthesis. Methods in Molecular Biology, vol 2489. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2273-5_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2273-5_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2272-8
Online ISBN: 978-1-0716-2273-5
eBook Packages: Springer Protocols