Abstract
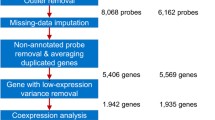
Gene co-expression analysis is a data analysis technique that helps identify groups of genes with similar expression patterns across several different conditions. By means of these techniques, different groups have been able to assign putative metabolic pathways and functions to understudied genes and to identify novel metabolic regulation networks for different metabolites. Some groups have even used network comparative studies to understand the evolution of these networks from green algae to land plants. In this chapter, we will go over the basic definitions required to understand network topology and gene module identification. Additionally, we offer the reader a walk-through a standard analysis pipeline as implemented in the package WGCNA that takes as input raw fastq files and obtains co-expressed gene clusters and representative gene expression patterns from each module for downstream applications.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Weston DJ, Gunter LE, Rogers A, Wullschleger SD (2008) Connecting genes, coexpression modules, and molecular signatures to environmental stress phenotypes in plants. BMC Syst Biol 2(1):16
Das S, Meher PK, Rai A, Bhar LM, Mandal BN (2017) Statistical approaches for gene selection, hub gene identification and module interaction in gene co-expression network analysis: an application to aluminum stress in soybean (Glycine max L.). PLoS One 12(1):e0169605
Emamjomeh A, Saboori Robat E, Zahiri J, Solouki M, Khosravi P (2017) Gene co-expression network reconstruction: a review on computational methods for inferring functional information from plant-based expression data. Plant Biotechnol Rep 11(2):71–86
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9(1):559
Labrou NE, Papageorgiou AC, Pavli O, Flemetakis E (2015) Plant GSTome: structure and functional role in xenome network and plant stress response. Curr Opin Biotechnol 32:186–194
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K (2017) Transcriptional regulatory network of plant heat stress response. Trends Plant Sci 22(1):53–65
Lv L, Zhang W, Sun L, Zhao A, Zhang Y, Wang L et al (2020) Gene co-expression network analysis to identify critical modules and candidate genes of drought-resistance in wheat. PLoS One 15(8):e0236186
Tai Y, Liu C, Yu S, Yang H, Sun J, Guo C et al (2018) Gene co-expression network analysis reveals coordinated regulation of three characteristic secondary biosynthetic pathways in tea plant (Camellia sinensis). BMC Genomics 19(1):616
Wisecaver JH, Borowsky AT, Tzin V, Jander G, Kliebenstein DJ, Rokas A (2017) A global coexpression network approach for connecting genes to specialized metabolic pathways in plants. Plant Cell 29(5):944–959
Guerin C, Joët T, Serret J, Lashermes P, Vaissayre V, Agbessi MDT et al (2016) Gene coexpression network analysis of oil biosynthesis in an interspecific backcross of oil palm. Plant J 87(5):423–441
DCJ W, Matus JT (2017) Constructing integrated networks for identifying new secondary metabolic pathway regulators in grapevine: recent applications and future opportunities. Front Plant Sci 8:505
Ferreira SS, Hotta CT, Poelking VG, DCC L, Buckeridge MS, Loureiro ME et al (2016) Co-expression network analysis reveals transcription factors associated to cell wall biosynthesis in sugarcane. Plant Mol Biol 91(1–2):15–35
Ruprecht C, Proost S, Hernandez-Coronado M, Ortiz-Ramirez C, Lang D, Rensing SA et al (2017) Phylogenomic analysis of gene co-expression networks reveals the evolution of functional modules. Plant J 90(3):447–465
Ruprecht C, Vaid N, Proost S, Persson S, Mutwil M (2017) Beyond genomics: studying evolution with gene Coexpression networks. Trends Plant Sci 22(4):298–307
Masalia RR, Bewick AJ, Burke JM (2017) Connectivity in gene coexpression networks negatively correlates with rates of molecular evolution in flowering plants. PLoS One 12(7):e0182289
Schaefer RJ, Michno J-M, Myers CL (2017) Unraveling gene function in agricultural species using gene co-expression networks. Biochim Biophys Acta Gene Regul Mech 1860(1):53–63
Gupta C, Pereira A (2019) Recent advances in gene function prediction using context-specific coexpression networks in plants. F1000Res 8
Zhang H, Zhang J, Xu Q, Wang D, Di H, Huang J et al (2020) Identification of candidate tolerance genes to low-temperature during maize germination by GWAS and RNA-seq approaches. BMC Plant Biol 20(1):333
Zhang H, Wang ML, Schaefer R, Dang P, Jiang T, Chen C (2019) GWAS and coexpression network reveal Ionomic variation in cultivated peanut. J Agric Food Chem 67(43):12026–12036
Schaefer RJ, Michno J-M, Jeffers J, Hoekenga O, Dilkes B, Baxter I et al (2018) Integrating coexpression networks with GWAS to prioritize causal genes in maize. Plant Cell 30(12):2922–2942
Marshall-Colón A, Kliebenstein DJ (2019) Plant networks as traits and hypotheses: moving beyond description. Trends Plant Sci 24(9):840–852
Shahan R, Zawora C, Wight H, Sittmann J, Wang W, Mount SM et al (2018) Consensus coexpression network analysis identifies key regulators of flower and fruit development in wild strawberry. Plant Physiol 178(1):202–216
Togninalli M, Seren Ü, Freudenthal JA, Monroe JG, Meng D, Nordborg M et al (2020) AraPheno and the AraGWAS catalog 2020: a major database update including RNA-Seq and knockout mutation data for Arabidopsis thaliana. Nucleic Acids Res 48(D1):D1063–D1068
Pearce S, Vazquez-Gross H, Herin SY, Hane D, Wang Y, Gu YQ et al (2015) WheatExp: an RNA-seq expression database for polyploid wheat. BMC Plant Biol 15(1):299
Xia L, Zou D, Sang J, Xu X, Yin H, Li M et al (2017) Rice expression database (RED): an integrated RNA-Seq-derived gene expression database for rice. J Genet Genomics 44(5):235–241
Chao H, Li T, Luo C, Huang H, Ruan Y, Li X et al (2020) BrassicaEDB: a gene expression database for brassica crops. Int J Mol Sci 21(16):5831
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL (2019) Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37(8):907–915
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36
Boratyn GM, Thierry-Mieg J, Thierry-Mieg D, Busby B, Madden TL (2019) Magic-BLAST, an accurate RNA-seq aligner for long and short reads. BMC Bioinformatics 20(1):405
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9(4):357–359
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14):1754–1760
Anders S, Pyl PT, Huber W (2015) HTSeq—a python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR et al (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and cufflinks. Nat Protoc 7(3):562–578
Kovaka S, Zimin AV, Pertea GM, Razaghi R, Salzberg SL, Pertea M (2019) Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol 20(1):278
Pertea M, Pertea GM, Antonescu CM, Chang T-C, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33(3):290–295
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550
Bodt SD, Carvajal D, Hollunder J, den Cruyce JV, Movahedi S, Inzé D (2010) CORNET: a user-friendly tool for data mining and integration. Plant Physiol 152(3):1167–1179
Song L, Langfelder P, Horvath S (2012) Comparison of co-expression measures: mutual information, correlation, and model based indices. BMC Bioinformatics 13:328
Toubiana D, Puzis R, Sadka A, Blumwald E (2019) A genetic algorithm to optimize weighted gene co-expression network analysis. J Comput Biol 26(12):1349–1366
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol 4:Article17
Barabasi A-L, Bonabeau E (2003) Scale-free networks. Sci Am 288(5):60–69
Chen Q, Shi D (2004) The modeling of scale-free networks. Phys A Stat Mech Appl 335(1):240–248
Horvath S, Dong J (2008) Geometric interpretation of gene coexpression network analysis. PLoS Comput Biol 4(8):e1000117
Langfelder P, Horvath S (2007) Eigengene networks for studying the relationships between co-expression modules. BMC Syst Biol 1(1):54
Pardo-Diaz J, Bozhilova LV, Beguerisse-Díaz M, Poole PS, Deane CM, Reinert G (2021) Robust gene coexpression networks using signed distance correlation. Bioinformatics 37(14):1982–1989. https://doi.org/10.1093/bioinformatics/btab041
Gysi D, Voigt A, Fragoso T et al (2018) wTO: an R package for computing weighted topological overlap and a consensus network with integrated visualization tool. BMC Bioinformatics 19:392. https://doi.org/10.1186/s12859-018-2351-7
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Montenegro, J.D. (2022). Gene Co-expression Network Analysis. In: Edwards, D. (eds) Plant Bioinformatics. Methods in Molecular Biology, vol 2443. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2067-0_19
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2067-0_19
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2066-3
Online ISBN: 978-1-0716-2067-0
eBook Packages: Springer Protocols