Abstract
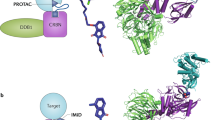
Assessing the specificity of PROTACs and confirming their proposed mechanism of action are critical for a robust targeted protein degradation program. Owing to their novel mechanism, new assays are needed to meet these goals. We and others have shown that a common explanation of PROTAC efficacy is the ability of the PROTAC to form a ternary complex between the E3 ubiquitin ligase and the target protein. In this chapter, we provide a simple in vitro method to quickly and inexpensively assess this property of PROTAC molecules. We provide detailed instructions for the purification of the specific E3 ubiquitin ligase VHL and then a generic protocol which can be adapted to any E3 ligase and substrate protein combination. This accessible method to study the ternary complex can strengthen any PROTAC-focused medicinal chemistry effort.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sakamoto KM, Kim KB, Kumagai A et al (2001) Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc Natl Acad Sci U S A 98:8554–8559
Bondeson DP, Mares A, Smith IED et al (2015) Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat Chem Biol 11:611–617
Winter GE, Buckley DL, Paulk J et al (2015) Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348:1376–1381
Zengerle M, Chan K-H, Ciulli A (2015) Selective small molecule induced degradation of the BET Bromodomain protein BRD4. ACS Chem Biol 10:1770–1777
Lu J, Qian Y, Altieri M et al (2015) Hijacking the E3 ubiquitin ligase Cereblon to efficiently target BRD4. Chem Biol 22:755–763
Gandhi AK, Kang J, Havens CG et al (2014) Immunomodulatory agents lenalidomide and pomalidomide co-stimulate T cells by inducing degradation of T cell repressors Ikaros and Aiolos via modulation of the E3 ubiquitin ligase complex CRL4CRBN. Br J Haematol 164:811–821
Lu G, Middleton RE, Sun H et al (2014) The myeloma drug Lenalidomide promotes the Cereblon-dependent destruction of Ikaros proteins. Science 343:305–309
Krönke J, Udeshi ND, Narla A et al (2014) Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343:301–305
Uehara T, Minoshima Y, Sagane K et al (2017) Selective degradation of splicing factor CAPERα by anticancer sulfonamides. Nat Chem Biol 13:675–680
Han T, Goralski M, Gaskill N et al (2017) Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science 356:eaal3755
Isobe Y, Okumura M, McGregor L et al (2020) Manumycin polyketides act as molecular glues between UBR7 and P53. Nat Chem Biol 16:1189–1198
Davis MI, Hunt JP, Herrgard S et al (2011) Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol 29:1046–1051
Klaeger S, Heinzlmeir S, Wilhelm M et al (2017) The target landscape of clinical kinase drugs. Science 358:eaan4368
Douglass EF, Miller CJ, Sparer G et al (2013) A comprehensive mathematical model for three-body binding equilibria. J Am Chem Soc 135:6092–6099
Wang L, Guillen VS, Sharma N et al (2018) New class of selective estrogen receptor degraders (SERDs): expanding the toolbox of PROTAC Degrons. ACS Med Chem Lett 9:803–808
Riching KM, Mahan S, Corona CR et al (2018) Quantitative live-cell kinetic degradation and mechanistic profiling of PROTAC mode of action. ACS Chem Biol 13:2758–2770
Wittmann BM, Sherk A, McDonnell DP (2007) Definition of functionally important mechanistic differences among selective estrogen receptor down-regulators. Cancer Res 67:9549–9560
Jones LH (2018) Small-molecule kinase Downregulators. Cell Chem Biol 25:30–35
Yang J, Li Y, Aguilar A et al (2019) Simple structural modifications converting a Bona fide MDM2 PROTAC degrader into a molecular glue molecule: a cautionary tale in the design of PROTAC degraders. J Med Chem 62:9471–9487
Bondeson DP, Smith BE, Burslem GM et al (2018) Lessons in PROTAC design from selective degradation with a promiscuous warhead. Cell Chem Biol 25:78–87.e5
Huang H-T, Dobrovolsky D, Paulk J et al (2018) A Chemoproteomic approach to query the degradable Kinome using a multi-kinase degrader. Cell Chem Biol 25:88–99.e6
Smith BE, Wang SL, Jaime-Figueroa S et al (2019) Differential PROTAC substrate specificity dictated by orientation of recruited E3 ligase. Nat Commun 10:131
Gadd MS, Testa A, Lucas X et al (2017) Structural basis of PROTAC cooperative recognition for selective protein degradation. Nat Chem Biol 13:514–521
Nowak RP, Deangelo SL, Buckley D et al (2018) Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat Chem Biol 14:706–714
Remillard D, Buckley DL, Paulk J et al (2017) Degradation of the BAF complex factor BRD9 by heterobifunctional ligands. Angew Chem Int Ed Engl 56:5738–5743
Acknowledgements
We thank Craig M. Crews for mentorship and supervision of this assay design. DPB is supported by the National Cancer Institute fellowship F99/K00 CA212229. BES (T32 GM007753) and ADB (T32 GM008152) are supported by Medical Scientist Training Program grants from the National Institute of General Medical Sciences at their respective institutions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Bondeson, D.P., Smith, B.E., Buhimschi, A.D. (2021). An In Vitro Pull-down Assay of the E3 Ligase:PROTAC:Substrate Ternary Complex to Identify Effective PROTACs . In: Cacace, A.M., Hickey, C.M., Békés, M. (eds) Targeted Protein Degradation. Methods in Molecular Biology, vol 2365. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1665-9_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1665-9_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1664-2
Online ISBN: 978-1-0716-1665-9
eBook Packages: Springer Protocols