Abstract
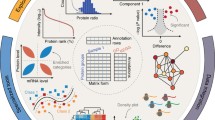
“Omics” techniques (e.g., proteomics, genomics, metabolomics), from which huge datasets can nowadays be obtained, require a different way of thinking about data analysis that can be summarized with the idea that, when data are enough, they can speak for themselves. Indeed, managing huge amounts of data imposes the replacement of the classical deductive approach (hypothesis-driven) with a data-driven hypothesis-generating inductive approach, so to generate mechanistical hypotheses from data.
Data reduction is a crucial step in proteomics data analysis, because of the sparsity of significant features in big datasets. Thus, feature selection/extraction methods are applied to obtain a set of features based on which a proteomics signature can be drawn, with a functional significance (e.g., classification, diagnosis, prognosis). Despite big data generated almost daily by proteomics studies, a well-established statistical workflow for data analysis in proteomics is still lacking, opening up to misleading and incorrect data analysis and interpretation. This chapter will give an overview of the methods available for feature selection/extraction in proteomics datasets and how to choose the most appropriate one based on the type of dataset.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ma’ayan A (2017) Complex systems biology. J R Soc Interface 14(134):20170391
Broad CD (1925) Mind and its place in nature. Harcourt, Brace & Company, Inc., New York
Hayes BK, Heit E, Swendsen H (2010) Inductive reasoning. Wiley Interdiscip Rev Cogn Sci 1:278–292
Hayes BK, Heit E (2018) Inductive reasoning 2.0. Wiley Interdiscip Rev Cogn Sci 9:e1459
He Q-Y, Chiu J-F (2003) Proteomics in biomarker discovery and drug development. J Cell Biochem 89:868–886
Kohn EC, Azad N, Annunziata C et al (2007) Proteomics as a tool for biomarker discovery. Dis Markers 23:411–417
Suppers A, van Gool AJ, Wessels HJCT (2018) Integrated chemometrics and statistics to drive successful proteomics biomarker discovery. Proteomes 6(2):20
Bittner L (1962) R. Bellman, adaptive control processes. A guided tour. XVI + 255 S. Princeton, N. J., 1961. Princeton University Press. Preis geb. $ 6.50. ZAMM J Appl Math Mech Z Für Angew Math Mech 42:364–365
Guyon I, Elisseeff A (2003) An introduction to variable and feature selection. J Mach Learn Res 3:1157–1182
Hira ZM, Gillies DF A review of feature selection and feature extraction methods applied on microarray data. Adv Bioinforma 2015:198363. https://www.hindawi.com/journals/abi/2015/198363/
Hoque N, Bhattacharyya DK, Kalita JK (2014) MIFS-ND: a mutual information-based feature selection method. Expert Syst Appl 41:6371–6385
Yu L, Liu H (2003) Feature selection for high-dimensional data: a fast correlation-based filter solution. In: Proceedings, twentieth international conference on machine learning, pp 856–863
Robnik-Šikonja M, Kononenko I (2003) Theoretical and empirical analysis of ReliefF and RReliefF. Mach Learn 53:23–69
Radovic M, Ghalwash M, Filipovic N et al (2017) Minimum redundancy maximum relevance feature selection approach for temporal gene expression data. BMC Bioinformatics 18:9
Azuaje F (2006) Witten IH, Frank E: data mining: practical machine learning tools and techniques 2nd edition. Biomed Eng Online 5:51
Alelyani S, Tang J, Liu H (2013) Feature selection for clustering: a review. In: Data clustering: algorithms and applications
Kohavi R, John GH (1997) Wrappers for feature subset selection. Artif Intell 97:273–324
Jović A, Brkić K, Bogunović N (2015) A review of feature selection methods with applications. In: 2015 38th International convention on information and communication technology, electronics and microelectronics (MIPRO), pp 1200–1205
Bonferroni C (1936) Teoria statistica delle classi e calcolo delle probabilita. Pubblicazioni R Ist Super Sci Econ E Commericiali Firenze 8:3–62
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300
Aggarwal S, Yadav AK (2016) False discovery rate estimation in proteomics. Methods Mol Biol 1362:119–128
Diz AP, Carvajal-Rodríguez A, Skibinski DOF (2011) Multiple hypothesis testing in proteomics: a strategy for experimental work. Mol Cell Proteomics 10
Wold S, Sjöström M, Eriksson L (2001) PLS-regression: a basic tool of chemometrics. Chemom Intell Lab Syst 58:109–130
Brereton RG, Lloyd GR (2014) Partial least squares discriminant analysis: taking the magic away. J Chemom 28:213–225
Gromski PS, Muhamadali H, Ellis DI et al (2015) A tutorial review: metabolomics and partial least squares-discriminant analysis—a marriage of convenience or a shotgun wedding. Anal Chim Acta 879:10–23
Hotelling H (1933) Analysis of a complex of statistical variables into principal components. J Educ Psychol 24:417–441
Jolliffe IT, Cadima J (2016) Principal component analysis: a review and recent developments. Philos Transact A Math Phys Eng Sci 374
Dubitzky W, Granzow M, Berrar DP (2007) Fundamentals of data mining in genomics and proteomics. Springer, Berlin
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Kuligowski J, Pérez-Guaita D, Quintás G (2016) Application of discriminant analysis and cross-validation on proteomics data. Methods Mol Biol 1362:175–184
Wang W, Sue AC-H, Goh WWB (2017) Feature selection in clinical proteomics: with great power comes great reproducibility. Drug Discov Today 22:912–918
Goh WWB, Wong L (2016) Evaluating feature-selection stability in next-generation proteomics. J Bioinforma Comput Biol 14:1650029
Lim K, Wong L (2014) Finding consistent disease subnetworks using PFSNet. Bioinformatics 30:189–196
Goh WWB, Wong L (2016) Advancing clinical proteomics via analysis based on biological complexes: a tale of five paradigms. J Proteome Res 15:3167–3179
Goh WWB (2016) Fuzzy-FishNET: a highly reproducible protein complex-based approach for feature selection in comparative proteomics. BMC Med Genet 9:67
Christin C, Hoefsloot HCJ, Smilde AK et al (2013) A critical assessment of feature selection methods for biomarker discovery in clinical proteomics. Mol Cell Proteomics 12:263–276
Alterovitz G, Liu J, Afkhami E et al (2007) Bayesian methods for proteomics. Proteomics 7:2843–2855
Hernández B, Pennington SR, Parnell AC (2015) Bayesian methods for proteomic biomarker development. EuPA Open Proteom 9:54–64
Dridi N, Giremus A, Giovannelli J-F et al (2017) Bayesian inference for biomarker discovery in proteomics: an analytic solution. EURASIP J Bioinforma Syst Biol 2017:9
Marchiori E, Heegaard NHH, West-Nielsen M et al (2005) Feature selection for classification with proteomic data of mixed quality. In: 2005 IEEE symposium on computational intelligence in bioinformatics and computational biology, pp 1–7
Guyon I, Weston J, Barnhill S et al (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46:389–422
Kira K, Rendell LA (1992) The feature selection problem: traditional methods and a new algorithm. In: Proceedings of the tenth national conference on artificial intelligence. AAAI Press, San Jose, CA, pp 129–134
Conrad TOF, Genzel M, Cvetkovic N et al (2017) Sparse proteomics analysis—a compressed sensing-based approach for feature selection and classification of high-dimensional proteomics mass spectrometry data. BMC Bioinformatics 18:160
Lualdi M, Fasano M (2019) Statistical analysis of proteomics data: a review on feature selection. J Proteome 198:18–26
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Lualdi, M., Fasano, M. (2021). Features Selection and Extraction in Statistical Analysis of Proteomics Datasets. In: Cecconi, D. (eds) Proteomics Data Analysis. Methods in Molecular Biology, vol 2361. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1641-3_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1641-3_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1640-6
Online ISBN: 978-1-0716-1641-3
eBook Packages: Springer Protocols