Abstract
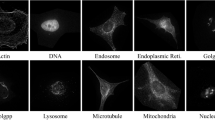
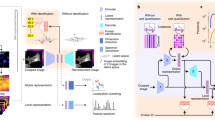
Protein subcellular localization prediction (PSLP), which plays an important role in the field of computational biology, identifies the position and function of proteins in cells without expensive cost and laborious effort. In the past few decades, various methods with different algorithms have been proposed in solving the problem of subcellular localization prediction; machine learning and deep learning constitute a large portion among those proposed methods. In order to provide an overview about those methods, the first part of this article will be a brief review of several state-of-the-art machine learning methods on subcellular localization prediction; then the materials used by subcellular localization prediction is described and a simple prediction method, that takes protein sequences as input and utilizes a convolutional neural network as the classifier, is introduced. At last, a list of notes is provided to indicate the major problems that may occur with this method.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Gardy JL, Brinkman FS (2006) Methods for predicting bacterial protein subcellular localization. Nat Rev Microbiol 4(10):741–751
Karp G (2009) Cell and molecular biology: concepts and experiments. Wiley, Hoboken, NJ
Tsien RY (1998) The green fluorescent protein. Annu Rev Biochem 67(1):509–544
Rey S, Gardy JL, Brinkman FS (2005) Assessing the precision of high-throughput computational and laboratory approaches for the genome-wide identification of protein subcellular localization in bacteria. BMC Genomics 6(1):162
Shen Y, Ding Y, Tang J, Zou Q, Guo F (2020) Critical evaluation of web-based prediction tools for human protein subcellular localization. Brief Bioinform 21(5):1628–1640
Gudenas BL, Wang L (2018) Prediction of LncRNA subcellular localization with deep learning from sequence features. Sci Rep 8(1):1–10
Javed F, Hayat M (2019) Predicting subcellular localization of multi-label proteins by incorporating the sequence features into Chou’s PseAAC. Genomics 111(6):1325–1332
Kumar KR, Cowley MJ, Davis RL (2019) Next-generation sequencing and emerging technologies. Semin Thromb Hemost 45(7):661–673
Zhang S, Duan X (2018) Prediction of protein subcellular localization with oversampling approach and Chou’s general PseAAC. J Theor Biol 437:239–250
Chou KC, Cai YD (2004) Prediction of protein subcellular locations by GO–FunD–PseAA predictor. Biochem Biophys Res Commun 320(4):1236–1239
Guo X, Liu F, Ju Y, Wang Z, Wang C (2016) Human protein subcellular localization with integrated source and multi-label ensemble classifier. Sci Rep 6:28087
Hasan MAM, Ahmad S, Molla MKI (2017) Protein subcellular localization prediction using multiple kernel learning based support vector machine. Mol BioSyst 13(4):785–795
Almagro Armenteros JJ, Sønderby CK, Sønderby SK, Nielsen H, Winther O (2017) DeepLoc: prediction of protein subcellular localization using deep learning. Bioinformatics 33(21):3387–3395
Wei L, Ding Y, Su R, Tang J, Zou Q (2018) Prediction of human protein subcellular localization using deep learning. J Parallel Distr Com 117:212–217
Cooper GM, Hausman RE (2004) The cell: molecular approach. Medicinska naklada
UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47(D1):D506–D515
Sastry A, Monk J, Tegel H, Uhlen M, Palsson BO, Rockberg J, Brunk E (2017) Machine learning in computational biology to accelerate high-throughput protein expression. Bioinformatics 33(16):2487–2495
Li H, Tian S, Li Y, Fang Q, Tan R, Pan Y, Huang C, Xu Y, Gao X (2020) Modern deep learning in bioinformatics. J Mol Cell Biol 12(11):823–827
Chen HU, Huang NI, Sun Z (2006) SubLoc: a server/client suite for protein subcellular location based on SOAP. Bioinformatics 22(3):376–377
Shen Y, Tang J, Guo F (2019) Identification of protein subcellular localization via integrating evolutionary and physicochemical information into Chou’s general PseAAC. J Theor Biol 462:230–239
Ding Y, Tang J, Guo F (2020) Human protein subcellular localization identification via fuzzy model on kernelized neighborhood representation. Appl Soft Comput 96:106596
He J, Gu H, Liu W (2012) Imbalanced multi-modal multi-label learning for subcellular localization prediction of human proteins with both single and multiple sites. PLoS One 7(6):e37155
Wei L, Liao M, Gao X, Wang J, Lin W (2016) mGOF-loc: a novel ensemble learning method for human protein subcellular localization prediction. Neurocomputing 217:73–82
Wu CH, Huang H, Arminski L, Castro-Alvear J, Chen Y, Hu ZZ, Ledley RS, Lewis KC, Mewes H-W, Orcutt BC, Suzek BE (2002) The protein information resource: an integrated public resource of functional annotation of proteins. Nucleic Acids Res 30(1):35–37
Gene Ontology Consortium (2019) The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res 47(D1):D330–D338
Wan S, Mak MW, Kung SY (2012) mGOASVM: multi-label protein subcellular localization based on gene ontology and support vector machines. BMC Bioinformatics 13(1):290
Wan S, Mak MW, Kung SY (2015) mLASSO-Hum: a LASSO-based interpretable human-protein subcellular localization predictor. J Theor Biol 382:223–234
Shen HB, Chou KC (2009) A top-down approach to enhance the power of predicting human protein subcellular localization: Hum-mPLoc 2.0. Anal Biochem 394(2):269–274
Zhang ML, Zhou ZH (2013) A review on multi-label learning algorithms. IEEE Trans Knowl Data Eng 26(8):1819–1837
Stormo GD, Schneider TD, Gold L, Ehrenfeucht A (1982) Use of the “Perceptron” algorithm to distinguish translational initiation sites in E. coli. Nucleic Acids Res 10(9):2997–3011
Stormo GD (2000) DNA binding sites: representation and discovery. Bioinformatics 16(1):16–23
Bhagwat M, Aravind L (2007) Comparative genomics. In: Psi-blast tutorial. Humana Press, Totowa, NJ, pp 177–186
Chou KC (2009) Pseudo amino acid composition and its applications in bioinformatics, proteomics and system biology. Curr Proteomics 6(4):262–274
Jeong JC, Lin X, Chen XW (2010) On position-specific scoring matrix for protein function prediction. IEEE/ACM Trans Comput Biol Bioinform 8(2):308–315
Nanni L, Brahnam S, Lumini A (2012) Wavelet images and Chou’s pseudo amino acid composition for protein classification. Amino Acids 43(2):657–665
Nanni L, Lumini A, Brahnam S (2014) An empirical study of different approaches for protein classification. Sci World J 2014:236717
Pan G, Wang J, Zhao L, Hoskins W, Tang J (2020) Computational methods for predicting DNA binding proteins. Curr Proteomics 17(4):258–270
Pan G, Jiang L, Tang J, Guo F (2018) A novel computational method for detecting DNA methylation sites with DNA sequence information and physicochemical properties. Int J Mol Sci 19(2):511
Guo F, Zou Q, Yang G, Wang D, Tang J, Xu J (2019) Identifying protein-protein interface via a novel multi-scale local sequence and structural representation. BMC Bioinformatics 20(15):1–11
Ding Y, Tang J, Guo F (2019) Protein crystallization identification via fuzzy model on linear neighborhood representation. IEEE/ACM Trans Comput Biol Bioinform. https://doi.org/10.1109/TCBB.2019.2954826
Collobert R, Weston J (2008) A unified architecture for natural language processing: deep neural networks with multitask learning. In: Proceedings of the 25th international conference on machine learning, pp 160–167
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. In: Advances in neural information processing systems, pp 1097–1105
Xu L, Ren JS, Liu C, Jia J (2014) Deep convolutional neural network for image deconvolution. In: Advances in neural information processing systems, pp 1790–1798
Lin X, Zhao C, Pan W (2017) Towards accurate binary convolutional neural network. In: Advances in neural information processing systems, pp 345–353
Zhou J, Troyanskaya OG (2015) Predicting effects of noncoding variants with deep learning–based sequence model. Nat Methods 12(10):931–934
Angermueller C, Lee HJ, Reik W, Stegle O (2017) DeepCpG: accurate prediction of single-cell DNA methylation states using deep learning. Genome Biol 18(1):1–13
Zhang Y, An L, Xu J, Zhang B, Zheng WJ, Hu M, Tang J, Yue F (2018) Enhancing Hi-C data resolution with deep convolutional neural network HiCPlus. Nat Commun 9(1):1–9
Zhang H, Weng TW, Chen PY, Hsieh CJ, Daniel L (2018) Efficient neural network robustness certification with general activation functions. In: Advances in neural information processing systems, pp 4939–4948
Han J, Moraga C (1995) The influence of the sigmoid function parameters on the speed of backpropagation learning. In: International workshop on artificial neural networks. Springer, Berlin, pp 195–201
De Boer PT, Kroese DP, Mannor S, Rubinstein RY (2005) A tutorial on the cross-entropy method. Ann Oper Res 134(1):19–67
Okada S, Ohzeki M, Taguchi S (2019) Efficient partition of integer optimization problems with one-hot encoding. Sci Rep 9(1):1–12
Li J, Si Y, Xu T, Jiang S (2018, 2018) Deep convolutional neural network based ECG classification system using information fusion and one-hot encoding techniques. Math Probl Eng:7354081
Pan G, Tang J, Guo F (2017) Analysis of co-associated transcription factors via ordered adjacency differences on motif distribution. Sci Rep 7(1):1–9
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Pan, G., Sun, C., Liao, Z., Tang, J. (2021). Machine and Deep Learning for Prediction of Subcellular Localization. In: Cecconi, D. (eds) Proteomics Data Analysis. Methods in Molecular Biology, vol 2361. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1641-3_15
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1641-3_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1640-6
Online ISBN: 978-1-0716-1641-3
eBook Packages: Springer Protocols